Abstract
Currently, the majority of antibiotics in clinical use have broad activity spectra, killing pathogenic and beneficial microorganisms indiscriminately. The disruption of the ecological balance of normal flora often results in secondary infections or other antibiotic-associated complications. Therefore, targeted antimicrobial therapies capable of specifically eliminating pathogenic bacteria while retaining the protective benefits of a normal microflora would be advantageous. In this study, we successfully constructed a series of Enterococcus faecalis-targeted antimicrobial peptides from wide-spectrum antimicrobial peptide precursors. These peptides are designed based on fusion of the species-specific peptide pheromone cCF10 and modification of the active region of the antimicrobial peptide. The results showed that cCF10-C4 possessed specific antimicrobial activity against E. faecalis and was not active against other types of bacteria tested. The specificity of this hybrid peptide was shown by the absence of antimicrobial effects in the pheromone-substituted derivative. Further studies indicated that cCF10-C4 and its parent peptide C4 exert their activities by damaging cytoplasmic membrane integrity. The present study reveals the application potential of these molecules as “probiotic” antimicrobials for the control of specific bacterial infections, and it also helps to elucidate the design and construction of species-specific antimicrobials with precise targeting specificity.
Similar content being viewed by others
Introduction
The mucosal surfaces and skins of animals are colonized by plenty of microorganisms. Most of the bacteria within the multispecies microbial community are beneficial1. This indigenous flora plays a very important role in nutrient acquisition and protective colonization2,3,4. Additionally, the normal flora represents an important ecological system, and alterations to the microflora may result in bacterial infections5,6,7. Unfortunately, the majority of conventional antibiotics have wide activity spectra, killing pathogens and normal microflora indiscriminately and disrupting the micro-ecological balance. The unavoidable loss of microflora and ecological disruption resulting from antibiotic treatment may lead to severe and recurrent complications from persistent pathogens or opportunistic microorganisms that recolonize the vacated niche easily8. The imbalance of normal microflora and the increasing threat of multidrug-resistant microbes highlight the urgent need for novel “targeted” antimicrobial therapies that can selectively eliminate pathogens without significantly disrupting resident normal flora.
Antimicrobial peptides (AMPs) found in host immune systems have attracted considerable attention due to their excellent antimicrobial properties and unique action mechanism. These peptides are broad-spectrum with potent activities against microbes, viruses, parasites and even tumor cells9,10,11. Furthermore, unlike traditional antimicrobial agents that inhibit specific metabolic pathways, the majority of antimicrobial peptides exert bactericidal effects via the irreversible disruption of bacterial membranes, inducing the leakage of vital cell components and resulting in eventual cell death12,13. The physical nature of membrane disruption is expected to prevent or delay the development of bacterial resistance to AMPs14. Due to these inherent advantages, AMPs can be used as effective weapons against pathogenic microorganisms such as multidrug-resistant microbes and are amongst the most promising candidates for a new generation of antimicrobial agents.
The idea of targeted antimicrobial therapy is supported by several naturally occurring AMPs, including plantaricin, lantibiotic and nisin. These AMPs were found to contain both targeting and antimicrobial domains within their peptide sequences15,16,17. The presence of targeting domains promoted the accumulation of peptides on the bacterial membrane, increasing local peptide concentrations and enhancing bactericidal activities against the targeted microbes18,19. In recent years, several attempts have been made to construct pathogen-selective antimicrobials by conjugating a bacterial recognition domain to antimicrobial peptides or bactericidal proteins20,21,22,23,24,25. These findings provided an intriguing starting point for the development of multidomain AMPs with selective bactericidal effects against specific bacteria. Although all of these synthetic molecules exhibited enhanced antimicrobial potency, selectivity, and kinetics against specific bacteria, inevitable bactericidal effects on unrelated bacteria were also observed with these peptides. The main reason for this problem may lay in the fact that the killing moiety of these fusion peptides had not been redesigned or modified, and thus electrostatic attractions between cationic peptide molecules and anionic lipids of the bacterial membrane were retained, thereby causing indiscriminant bactericidal effect via a non-receptor-mediated pathway. Thus, these approaches have yet to generate target-specific antimicrobials.
In this study, a rational approach was adopted to construct a series of Enterococcus faecalis-targeted AMPs using the enterococcal pheromone cCF10 as the targeting domain and optimizing the active centre of AMP for enhanced target specificity. Bacterial peptide pheromones are species-specific signalling molecules that mediate intercellular communication in some Gram-positive bacteria and fungi26. These small peptides can traverse the cell wall and bind to the cognate membrane receptors with high affinities27,28. The targeting specificity and remarkable affinity of pheromones to their membrane receptors make them ideal candidates for species-specific targeting peptide domains that provide specific binding to a selected organism. The antimicrobial activities and targeting specificities of the designed peptides were evaluated against a panel of microbial species including both Gram-negative and Gram-positive bacteria. The haemolytic properties these peptides were measured, and flow cytometric, fluorescent spectrographic, and electron microscopy assays were performed to further evaluate peptide-membrane interaction mechanisms. The aim of the present study is to provide a rational approach to convert normally wide-spectrum antimicrobial peptides into “smart” targeted antimicrobials with precise specificity against target organisms without damaging benign bystanders.
Results
Peptide design and characterization
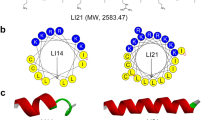
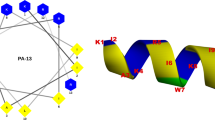
Initially, targeted antimicrobial peptide molecules were constructed by fusing the E. faecalis-specific peptide pheromone cCF10 with the broad-spectrum antimicrobial peptide C6, derived from the designed tryptophan-zipper antimicrobial peptide WK3 at either the N-terminus or the C-terminus29. To ensure that the targeting domain and the killing domain remain functionally and structurally independent, a flexible peptide linker (GGG) was incorporated between the two functional domains. Biological testing of these peptides showed that the addition of cCF10 at the N terminus of C6 significantly increased antimicrobial activity against E. faecalis relative to C6 alone; however, the hybrid peptides still possessed antimicrobial activities against the other bacteria tested. We reasoned that these undesired bactericidal effects against unrelated bacteria might arise from a high density of positive charges, which contributes to AMPs attraction toward and attachment to anionic bacteria cell membranes. In nature, such interactions occur via nonspecific electrostatic interactions. Given the known high affinity and specificity of a pheromone to its membrane receptor, we theorized that the cCF10 pheromone is adequate for targeted delivery of the antibacterial peptide to E. faecalis. Weakening the electrostatic interactions between AMPs and microbes may therefore prevent the molecules from entering the microbial membrane, decreasing or abolishing antimicrobial activity against untargeted bacterial species, while maintaining potent antimicrobial activity toward pheromone-sensing bacteria. To test this hypothesis, peptides with decreased positive charges were designed by replacing cationic K residues with anionic E or neutral Q residues at defined positions. The net charge of the peptide was decreased from +6 (cCF10-C6) to +4 (cCF10-C4) and +3 (cCF10-C3). Additionally, to evaluate the pheromone’s role in the specificity of the targeted antimicrobial peptide, the cCF10 targeting domain in cCF10-C4 was replaced with a random peptide sequence of the same length as cCF10.
The key physicochemical parameters of the designed peptides are summarized in Table 1. MALDI-TOF MS analysis showed that the measured molecular weights of the synthetic peptides were in close agreement with their theoretical values, suggesting that the products correspond to the designed compositions.
Antimicrobial activity
To evaluate the antibacterial property and general specificity of the designed peptides, MIC tests were performed against a broad selection of model bacteria, including Gram-negative bacteria and Gram-positive bacteria. As shown in Table 2, cCF10 had no effect on the growth of the bacterial species tested. C6 exhibited broad-spectrum antimicrobial activity against the panel of microbes, with MICs ranging from 2 to 16 µM. With the fusion of cCF10, the C6-cCF10 peptide showed similar antimicrobial activity against E. faecalis compared to the parent antimicrobial peptide C6, while cCF10-C6 displayed four-fold increased antimicrobial activity relative to that of C6 alone. However, the abilities of these two hybrid peptides to inhibit the growth other bacteria were much weaker than that of C6 alone. These findings showed the first indications that the presence of the cCF10 targeting domain increased antimicrobial activity and selectivity against E. faecalis but no other microorganisms. More importantly, the reduction of net positive charges within the killing domain resulted in the loss of bactericidal effects toward all the untargeted bacteria while maintaining robust antimicrobial activity against E. faecalis: cCF10-C4 exhibited an MIC against E. faecalis eight times lower than C4 alone, indicating a unique specificity for the targeted bacteria. However, it should be noted that further decreasing the positive charge to +3 abolished antimicrobial activity completely. Furthermore, the substitution of the cCF10 targeting domain with a random peptide sequence led to the loss of antimicrobial activity to E. faecalis (Table 2). These results indicate that the specificity of cCF10-C4 against the targeted bacteria is dependent on pheromone-receptor interactions.
Hemolytic activity
The haemolytic activities of the fusion peptides against human erythrocytes were evaluated to provide an indication of their cytotoxicity to mammalian cells (Fig. 1). The parent peptides C4 and C6 induced no haemolysis in human red blood cells at all concentrations tested. In contrast, the hybrid peptides cCF10-C4 and cCF10-C6 demonstrated little detectable haemolysis, with MHCs (the concentration that induces 10% or more haemolysis) of 32 µM and 128 µM, respectively. These MHCs indicate that the haemolytic activity of the peptides increases when the cCF10 pheromone is fused to the peptide.
Haemolytic effect of the peptides against hRBCs. Results were presented as the mean ± standard deviation (SD) of three independent assays. Differences between groups exposed to the same peptide concentration were analyzed by one-way ANOVA followed by Turkey’s post-hoc analysis (n = 3). The values with different superscripts (a, b, c) indicate a significant difference (P < 0.05).
Cytoplasmic membrane electrical potential
Due to the peptide’s greatly enhanced antimicrobial and unique specificity against E. faecalis, the mechanism of action of the targeted peptide cCF10-C4 was studied. The ability of the peptide to depolarize bacterial cytoplasmic membranes was determined using the membrane potential-dependent probe diSC3-5, which is quenched in the cytoplasmic membrane. The cytoplasmic membrane potential dissipates when the membrane is disrupted and permeabilized, releasing diSC3-5 into the medium and resulting in a conspicuous increase in fluorescence. The results showed that, the depolarization of the cell membrane of E. faecalis by the peptides was time- and dose-dependent (Fig. 2). The peptide cCF10-C4 caused a rapid increase in fluorescence at low micromolar concentrations. This increase was obviously higher than in parallel samples treated with the same concentration of C4 parent peptide, indicating that the addition of pheromone to C4 selectively enhanced the capacity to dissipate the transmembrane potential of the target bacteria.
Flow cytometry
The DNA-intercalating dye PI was used to evaluate the integrity of the microbial cell membranes. PI stains the nucleic acids in cells when their membranes are disrupted, causing an increase in fluorescence. As shown in Fig. 3, only 0.1% of E. faecalis cells were stained with PI in the absence of peptide. After treatment with cCF10-C4 and C4 at their minimum inhibitory concentrations, 98.2% and 82.7% of cells presented PI fluorescent signals, respectively. It is clear from the data that both peptides possessed the capability to destruct the cell membrane integrity of E. faecalis. However, cCF10-C4 caused a greater accumulation of PI than its parent peptide C4.
Flow cytometric analysis. The membrane damage of E. faecalis ATCC 29212 treated by peptides was measured by an increase in the fluorescence intensity of propidium iodide (PI) at 4 °C for 30 min. (a) control; (b) cCF10-C4 treated; (c) C4 treated. Bacteria were treated with peptides at 1 × MICs for 30 min. The control was processed without peptides. Data are representative of three independent experiments.
Scanning electron microscopy (SEM) observations
SEM was employed to directly visualize bacterial surface morphology and membrane damage after treatment with peptides at 1 × MICs for 2 h. In comparison to the untreated cells, which exhibited smooth and intact surfaces (Fig. 4A), treatment with cCF10-C4 and C4 induced significant morphological alterations. The membrane surfaces of peptide-exposed E. faecalis cells became roughened, deformed, and covered by numerous blebs (Fig. 4B,C).
Transmission electron microscopy (TEM) observations
To investigate the membrane integrity and intracellular alterations of E. faecalis cells treated with peptides, samples were analysed by TEM. As shown in Fig. 5, intact membrane envelopes and dense internal structures were observed in the untreated bacterial cells (Fig. 5A). Following treatment with peptides, the cytoplasmic membranes became blurred and began to collapse (Fig. 5B,C). In addition, dispersion of intracellular contents was also observed in peptide-treated E. faecalis cells.
Discussion
The overwhelming majority of antimicrobials in clinical use possess wide-spectrum bactericidal activities. These antibiotics have an advantage when the specific pathogen causing an infection is unknown. However, the use of such antimicrobials may indiscriminately kill benign and and pathogenic microbes, inducing an imbalance in the nomal flora and resulting in severe posttreatment complications17. Therefore, pathogen-specific antimicrobial therapies capable of precisely killing pathogens without affecting normal flora would be an ideal solution and could help to re-establish ecological balance and provide long-term protection. Unlike conventional antimicrobial agents, which may lose effect when their molecular structures are changed, AMPs can be optimized to obtain desirable properties through modification of their primary sequences30,31, making them a potential treasure trove of starting points for the development of targeted antimicrobial biomaterials. Here, we successfully designed a targeted antimicrobial peptide with unique specificity against a target organism without damaging other types of bacteria. The peptide was constructed by incorporating a natural pheromone as the targeting domain and reducing the net positive charge within the killing domain for enhanced target specificity. Given the universality and scalability of this approach, we believe that the methods presented here would be an effective strategy to make target-specific therapies a reality.
Our results showed that the fusion of the targeting region to the N-terminus of the antimicrobial peptides provided a clear increase in antimicrobial activity against the targeted bacteria. Given the high affinity of pheromones for their membrane receptors, it is likely that the improved activities of the hybrid peptides arise from selective accumulation of the peptide molecules on E. faecalis membranes, mediated by the pheromone cCF10. This accumulation generates higher local peptide concentrations, thereby enhancing antibacterial activity23. In addition, it is possible that the hydrophobicity of the fused cCF10 (containing 6 hydrophobic residues) may enhance the intrinsic ability of the molecules to insert into the cytoplasmic membrane bilayer independent of receptor binding. However, we found that all the fusion peptides exhibited decreased antimicrobial activities compared to their parent peptides against the other untargeted microorganisms tested. These findings suggest that the pheromone-receptor interaction is the primary reason for the increased antimicrobial activities. Further studies demonstrated that substitution of cCF10 with a random sequence resulted in a complete loss of bactericidal effects. The results further validate the notion that the specific antimicrobial activity of cCF10-C4 against the targeted bacteria is receptor-dependent. Both cCF10 and C4 components were necessary; however, neither component alone was sufficient. It has been reported that the cCF10 pheromone binds to the PrgZ receptor at the N-terminus28. As observed in this study, cCF10-C6 with a free pheromone N-terminus exhibited robust antimicrobial activity against E. faecalis; the activity of C6-cCF10 was much weaker in comparison, indicating that the pheromone-receptor interaction may be affected by where C6 was fused.
It is generally agreed that the first step in AMP activity is binding of cationic residues in the peptides to the anionic lipids of the microbial membrane via electrostatic interactions32. Previous literature has demonstrated that within a threshold (usually +6), increasing the positive charge of an antibacterial peptide facilitates its anchoring to the membrane, resulting in improved antibacterial properties33,34. Both of the initially designed peptides (cCF10-C6 and C6-cCF10) contained high net charges (+6), indicating sufficient driving of the electrostatic attachment to the bacterial membrane. Thus it was not surprising to find that these fused peptides still retained antimicrobial activities against unrelated bacterial strains. As a targeting domain, the pheromone cCF10 provides specific binding to its target organism with high affinity independent of electrostatic attraction. Therefore, to further narrow the antibacterial spectrum of the peptide, we decreased the net positive charge within the killing domain. Our data showed that the peptide cCF10-C4 is capable of precisely killing E. faecalis without affecting any of the other microorganisms tested. Therefore, we believe that our goal to construct a specifically targeted antimicrobial peptide was achieved. Furthermore, the specificity of targeted antibacterial peptides can be increased by reducing positive charges within a certain range. Further decreasing the net charge to +3 (cCF10-C3) resulted in the loss of antimicrobial effects, implying that the antimicrobial peptide domain was not sufficient to disrupt bacterial membranes. This finding provides evidence that the cationic amino acid residues within AMPs play important roles not only in the electrostatic interactions of peptides with anionic lipids but also in the physical disruption of microbial cell membranes. Further studies are underway to elucidate the implications of this new finding.
Hemolysis is often thought to be one of the obstacles preventing the clinical applications of AMPs as antibacterial agents. Previous literatures showed that peptide hydrophobicity is positively correlated with haemolytic activity29,35. Consistent with this interpretation, the addition of the cCF10 pheromone to C6 or C4 caused a slight increase in haemolytic rates. Fortunately, the hybrid peptides induced minimal or no haemolysis against human red blood cells at antimicrobial levels, indicating that these peptides could be developed as potential antibacterial agents for clinical use.
According to previous studies, the majority of cationic AMPs exert antimicrobial activities by damaging cytoplasmic membrane integrity13,36. Therefore, in this work, the bactericidal mechanism was evaluated, placing a special focus on the influences of peptides on the cytoplasmic membrane. Generally, membrane depolarization is an obvious feature of peptide-membrane interactions that plays an important role in determining the antibacterial potency of AMPs37. Our results indicated that the cCF10-C4 peptide results in a significant increase in the degree of cytoplasmic membrane depolarization in E. faecalis compared to C4 alone (Fig. 2), which correlates with their antibacterial activities against the targeted bacteria. This property was not observed in tests of untargeted bacterial species, implying that the enhanced killing activity and target specificity of the hybrid peptide may result from increased depolarization of the bacterial membrane. Furthermore, the results from TEM and SEM studies demonstrated that the designed peptides exert their bactericidal activities by damaging cytoplasmic membranes, causing the leakage of cytoplasmic components into the extracellular medium (Fig. 4 and Fig. 5). In addition, flow cytometry data further demonstrated that the designed peptides killed bacteria by disrupting cell membrane integrity (Fig. 3). This mechanism of action, based on physical destruction of the membrane, make it difficult for microbes to acquire resistance because this would require bacteria to mutate or repair their entire membrane lipid composition.
Conclusions
In this work, a series of targeted antimicrobial peptides were constructed and evaluated for their biological activities and antimicrobial mechanisms. The peptide was designed by fusing a natural pheromone as the targeting domain and reducing positive charge for enhanced target specificity. The designed peptides, in particular cCF10-C4, exhibited robust specific activity against targeted E. faecalis at micromolar concentrations, while pheromone-insensitive organisms were not affected. This specificity, combined with the low hemolytic activity of the designed peptides, suggests that these peptides can be used as promising antimicrobial agents for clinical applications against specific bacterial infections. Additionally, cCF10-C4 exerted its bactericidal effects via the destruction of cytoplasmic membrane integrity, allowing the efflux of internal components and resulting in cell death. This physical membrane disruption mechanism may decrease the occurrence of microbial resistance. Taken together, our findings presented here provides a feasible strategy for the construction of targeted AMPs with precise specificity against target organism without damaging other bacterial species. Furthermore, peptide pheromones widely existed in microorganisms, implying a large pool which provide growing candidates for the development of novel targeting peptides. This indicates that targeted antimicrobial agent construction in the future will be a “tunable” process whereby a wide variety of combinations of targeting and killing regions may be combined.
Materials and Methods
Materials
The bacterial strains Enterococcus faecalis ATCC 29212, Staphylococcus epidermidis ATCC 12228, Staphylococcus aureus ATCC 29213, Salmonella typhimurium C7731, Escherichia coli ATCC 25922 and Salmonella Pullorum C7913 were obtained from the College of Veterinary Medicine, Northeast Agricultural University (Harbin, China). The human red blood cells (hRBCs) were obtained from the Northeast Agricultural University Hospital.
Phosphate-buffered saline (PBS) solution was purchased from Kermel (China). Mueller Hinton Broth (MHB) powder was purchased from AoBoX (China) and used to prepare the microbial broths according to the manufacturer’s instructions. 3,3′-dipropylthiadicarbocyanine (diSC3-5), dimethyl sulfoxide (DMSO), ethanol (analytical grade, 99%), tertiary butanol (analytical grade, 99%), acetone (analytical grade, 99%), glutaraldehyde (synthetic grade, 50% in H2O) and propidium iodide (PI) were all ordered from Sigma-Aldrich (China).
Peptides synthesis
The peptides used in this study were synthesized and purified by GL Biochem (Shanghai, China) through solid-phase methods using Fmoc chemistry. Matrix-assisted laser desorption/ionization time-of-flight mass spectroscopy (MALDI-TOF MS, Linear Scientific Inc., U.S.A.) was used to determine the true molecular masses of these peptides. Peptide purities were confirmed to be >95% using analytical reverse-phase high-performance liquid chromatography (HPLC). The peptide cCF10 was dissolved in 40% DMSO, and other peptides were dissolved in deionized water at a concentration of 2.56 mM. Peptides were stored at −20 °C for subsequent assessments.
Antimicrobial assays
The antimicrobial activities of the peptides were investigated against the following bacteria: E. faecalis ATCC 29212, S. epidermidis ATCC 12228, S. aureus ATCC 29213, E. coli ATCC 25922, S. typhimurium C7731 and S. Pullorum C7913. The MICs of the peptides were determined using a modified standard microtiter dilution method as described previously29,38. Briefly, the microbial strains were cultured in MHB at 37 °C to reach mid-log phase and then diluted to 0.5-1×106 CFU/ml. 50 μL volumes of bacteria suspension were incubated in sterile 96-well plates with 50 μl of 2-fold serially diluted peptides at different concentrations (0.25–128 μM). Cultures with or without bacterial cells was employed as positive and negative controls, respectively. MIC was defined as the minimal concentration of peptides that results in no visible turbidity after incubation at 37 °C for 16 h. Each measurement was reproduced at least four times using three replicates.
Hemolytic activity assay
The hemolytic activity of the peptides was evaluated as the amount of hemoglobin released by the disruption of human red blood cells (hRBCs)39,40. Fresh human erythrocytes were obtained from a healthy donor (Changxuan Shao, Harbin, China) after informed consent. Then, the collected hRBCs were washed three times with PBS (pH 7.2), centrifuged at 1,000 × g for 5 min at 4 °C, and re-suspended in PBS to attain a dilution of approximately 1% (v/v) of erythrocytes. A 50 μl portion of the erythrocyte suspension were incubated with 50 μl of serially diluted peptides (0.5–256 μM) dissolved in PBS for 1 h at 37 °C. After centrifugation (1,000 × g, 5 min, 4 °C), the supernatant was transferred to a new 96-well plate. Haemoglobin release was measured by monitoring optical density (OD) at 570 nm. The hRBCs in PBS and 0.1% (v/v) Triton X-100 were employed as negative and positive controls, respectively. Minimal haemolytic concentrations (MHCs) were defined as the peptide concentrations causing 10% haemolysis. Three independent experiments were performed in duplicate. The percent hemolysis was calculated using the following equation: Hemolysis (%) = [(OD570 of the treated sample − OD570 of the negative control)/(OD570 of the positive control − OD570 of the negative control)] × 100%. The experimental protocol was reviewed and approved by the ethics committee of the Northeast Agricultural University Hospital, and the experimental method was carried out in accordance with the approved guidelines and regulations.
Cytoplasmic membrane depolarization assay
The membrane depolarization activity of the peptides was evaluated using the membrane potential-sensitive fluorescent dye diSC3-5 as described previously36. Briefly, E. faecalis ATCC 29212 cells were grown to mid-log phase, harvested by centrifugation at 1,000 × g for 10 min, washed thrice, and re-suspended to an OD600 of 0.05 with 5 mM HEPES buffer (pH 7.4, containing 20 mM glucose) containing 0.1 M KCl to equilibrate cytoplasmic and external K+ concentrations. The cell suspensions were then incubated with 0.4 μM diSC3-5 for 90 min. Subsequently, 2 mL of cell suspension was added to a 1 cm quartz cuvette and mixed with peptides at final concentrations ranging from 4 to 64 µM. Changes in fluorescence were recorded (excitation λ = 622 nm, emission λ = 670 nm) with an F-4500 Fluorescence Spectrophotometer (Hitachi, Japan). The deionized water without peptides and natural βhairpin AMP PG-1 were employed as negative and positive controls, respectively.
FACScan analysis
Bacterial cell membrane integrity was evaluated by flow cytometry38. Briefly, E. faecalis ATCC 29212 was grown to mid-log phase, harvested, washed thrice with 10 mM PBS and diluted to 105 CFU/ml. Bacterial suspensions were incubated with peptides at their minimum inhibitory concentrations for 30 min at 28 °C. Propidium iodide (PI, Sigma) was added at a final concentration of 10 µg/ml and incubated for an additional 30 min at 4 °C. Unbound dye was removed by washing with excess PBS. Cells incubated with PI in the absence of peptides were served as the negative control. Data were recorded using a FACScan instrument (Becton-Dickinson, San Jose, CA) with a laser excitation wavelength of 488 nm.
Scanning electron microscopy (SEM) characterization
Bacteria samples were prepared as described previously38. Briefly, E. faecalis ATCC 29212 cells were cultured in MHB at 37 °C to mid-log phase, harvested by centrifugation at 1,000 × g for 10 min, washed twice with 10 mM PBS, and re-suspended to an OD600 of 0.2. The cell suspensions were incubated at 37 °C for 120 min with different peptides at their 1× MICs. Controls were run without peptides. Following the incubation, the cells were centrifuged and washed thrice with PBS at 5,000 × g for 5 min. Subsequently, the bacterial cell pellets were fixed overnight with 2.5% (v/v) glutaraldehyde at 4 °C and washed twice with PBS followed by dehydration in a graded ethanol series (50%, 70%, 90%, and 100%) for 15 min each. The dried specimens were transferred to a mixture of ethanol and tertiary butanol (1:1, v/v) for 20 min followed by pure tertiary butanol for 30 min. Finally, the samples were dried using a critical point dryer, coated with gold, and then observed by SEM (Hitachi S-4800, Japan).
Transmission electron microscopy (TEM) characterization
TEM was performed to visualize intracellular alterations as described previously40,41. Bacteria samples were initially prepared as for SEM. After pre-fixation with 2.5% glutaraldehyde overnight, the cell pellets were washed twice and were post-fixed with 2% osmium tetroxide for 90 min. After being washed twice with PBS, the samples were dehydrated for 10 min in a graded ethanol series (50%, 70%, 90%, and 100%) followed by 10 min in a mixture of absolute ethanol and acetone (1:1, v/v) and 10 min in absolute acetone. Subsequently, the specimens were transferred to a mixture of absolute acetone and epoxy resin (1:1, v/v) for 30 min and then to pure epoxy resin overnight at a constant temperature. Finally, the samples were sectioned using an ultramicrotome, stained with uranyl acetate and lead citrate, and then observed by TEM (Hitachi H-7650, Japan).
Statistical analysis
Data were analysed by one-way ANOVA using SPSS 16.0 software. Quantitative data are presented as the means ± standard deviation. Differences were defined as significant at a P-value of less than 0.05.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Eckert, R., Sullivan, R. & Shi, W. Targeted antimicrobial treatment to re-establish a healthy microbial flora for long-term protection. Adv. Dent. Res. 24, 94–97, https://doi.org/10.1177/0022034512453725 (2012).
Boman, H. G. Innate immunity and the normal microflora. Immunol. Rev. 173, 5–16, https://doi.org/10.1034/j.1600-065x.2000.917301.x (2000).
Pickard, K. M., Bremner, A. R., Gordon, J. N. & MacDonald, T. T. Microbial-gut interactions in health and disease. Immune responses. Best. Pract. Res. Clin. Gastroenterol. 18, 271–285, https://doi.org/10.1016/j.bpg.2003.10.009 (2004).
Rastall, R. A. Bacteria in the gut: Friends and foes and how to alter the balance. J. Nutr. 134, 2022s–2026s, https://doi.org/10.1093/jn/134.8.2022S (2004).
Pultz, N. J., Hoyen, C. K. & Donskey, C. J. Inhibition of methicillin-resistant Staphylococcus aureus by an in vitro continuous-flow culture containing human stool microflora. FEMS Microbiol. Lett. 241, 201–205, https://doi.org/10.1016/j.femsle.2004.10.021 (2004).
Mitchell, T. J. The pathogenesis of streptococcal infections: from tooth decay to meningitis. Nat. Rev. Microbiol. 1, 219–230, https://doi.org/10.1038/nrmicro771 (2003).
Sansonetti, P. J. War and peace at mucosal surfaces. Nat. Rev. Immunol. 4, 953–964, https://doi.org/10.1038/nri1499 (2004).
Eckert, R. Road to clinical efficacy: challenges and novel strategies for antimicrobial peptide development. Future Microbiol. 6, 635–651, https://doi.org/10.2217/fmb.11.27 (2011).
Shao, C. et al. Central beta-turn increases the cell selectivity of imperfectly amphipathic alpha-helical peptides. Acta Biomater. 69, 243–255, https://doi.org/10.1016/j.actbio.2018.01.009 (2018).
Takiguchi, T. et al. Cathelicidin antimicrobial peptide LL-37 augments interferon-beta expression and antiviral activity induced by double-stranded RNA in keratinocytes. Br. J. Dermatol. 171, 492–498, https://doi.org/10.1111/bjd.12942 (2014).
Torrent, M., Pulido, D., Rivas, L. & Andreu, D. Antimicrobial peptide action on parasites. Curr. Drug. Targets 13, 1138–1147, https://doi.org/10.2174/138945012802002393 (2012).
Teixeira, V., Feio, M. J. & Bastos, M. Role of lipids in the interaction of antimicrobial peptides with membranes. Prog. Lipid Res. 51, 149–177, https://doi.org/10.1016/j.plipres.2011.12.005 (2012).
Ong, Z. Y., Gao, S. J. & Yang, Y. Y. Short Syntheticβ-Sheet Forming Peptide Amphiphiles as Broad Spectrum Antimicrobials with Antibiofilm and Endotoxin Neutralizing Capabilities. Adv. Funct. Mater. 23, 3682–3692, https://doi.org/10.1002/adfm.201202850 (2013).
Gopal, R., Seo, C. H., Song, P. I. & Park, Y. Effect of repetitive lysine-tryptophan motifs on the bactericidal activity of antimicrobial peptides. Amino acids 44, 645–660, https://doi.org/10.1007/s00726-012-1388-6 (2013).
Kristiansen, P. E., Fimland, G., Mantzilas, D. & Nissen-Meyer, J. Structure and mode of action of the membrane-permeabilizing antimicrobial peptide pheromone plantaricin A. J. Biol. Chem. 280, 22945–22950, https://doi.org/10.1074/jbc.M501620200 (2005).
Asaduzzaman, S. M. & Sonomoto, K. Lantibiotics: diverse activities and unique modes of action. J. Biosci. Bioeng. 107, 475–487, https://doi.org/10.1016/j.jbiosc.2009.01.003 (2009).
Breukink, E. et al. Use of the Cell Wall Precursor Lipid II by a Pore-Forming Peptide Antibiotic. Sci. 286, 2361, https://doi.org/10.1126/science.286.5448.2361 (1999).
Shai, Y. Mode of action of membrane active antimicrobial peptides. Biopolym. 66, 236–248, https://doi.org/10.1002/bip.10260 (2002).
Brogden, K. A. Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Nat. Rev. Microbiol. 3, 238–250, https://doi.org/10.1038/nrmicro1098 (2005).
Xi, D. et al. Design, expression and characterization of the hybrid antimicrobial peptide LHP7, connected by a flexible linker, against Staphylococcus and Streptococcus. Process. Biochem. 48, 453–461, https://doi.org/10.1016/j.procbio.2013.01.008 (2013).
He, J. et al. Systematic Approach to Optimizing Specifically Targeted Antimicrobial Peptides against Streptococcus mutans. Antimicrob. Agents Ch. 54, 2143–2151, https://doi.org/10.1128/Aac.01391-09 (2010).
He, J., Anderson, M. H., Shi, W. & Eckert, R. Design and activity of a ‘dual-targeted’ antimicrobial peptide. Int. J. Antimicrob. Ag. 33, 532–537, https://doi.org/10.1016/j.ijantimicag.2008.11.013 (2009).
Eckert, R. et al. Adding selectivity to antimicrobial peptides: Rational design of a multidomain peptide against Pseudomonas spp. Antimicrob. Agents Ch. 50, 1480–1488, https://doi.org/10.1128/Aac.50.4.1480-1488.2006 (2006).
Eckert, R. et al. Targeted Killing of Streptococcus mutans by a Pheromone-Guided “Smart” Antimicrobial Peptide. Antimicrob. Agents Ch. 50, 3651–3657, https://doi.org/10.1128/Aac.00622-06 (2006).
Qiu, X. Q. et al. An engineered multidomain bactericidal peptide as a model for targeted antibiotics against specific bacteria. Nat. Biotechnol. 21, 1480–1485, https://doi.org/10.1038/nbt913 (2003).
Yajima, A. Recent progress in the chemistry and chemical biology of microbial signaling molecules: quorum-sensing pheromones and microbial hormones. Tetrahedron Lett. 55, 2773–2780, https://doi.org/10.1016/j.tetlet.2014.03.051 (2014).
Lyon, G. J. & Novick, R. P. Peptide signaling in Staphylococcus aureus and other Gram-positive bacteria. Peptides 25, 1389–1403, https://doi.org/10.1016/j.peptides.2003.11.026 (2004).
Chandler, J. R. & Dunny, G. M. Enterococcal peptide sex pheromones: synthesis and control of biological activity. Peptides 25, 1377–1388, https://doi.org/10.1016/j.peptides.2003.10.020 (2004).
Xu, L. et al. Antimicrobial activity and membrane-active mechanism of tryptophan zipper-like beta-hairpin antimicrobial peptides. Amino acids 47, 2385–2397, https://doi.org/10.1007/s00726-015-2029-7 (2015).
Mai, J. et al. A Novel Target-Specific, Salt-Resistant Antimicrobial Peptide against the Cariogenic Pathogen Streptococcus mutans. Antimicrob. Agents Ch. 55, 5205–5213, https://doi.org/10.1128/Aac.05175-11 (2011).
Wiesner, J. & Vilcinskas, A. Antimicrobial peptides The ancient arm of the human immune system. Virulence 1, 440–464, https://doi.org/10.4161/viru.1.5.12983 (2010).
Fjell, C. D., Hiss, J. A., Hancock, R. E. W. & Schneider, G. Designing antimicrobial peptides: form follows function. Nat. Rev. Drug. Discov. 11, 37–51, https://doi.org/10.1038/nrd3591 (2011).
Tossi, A., Sandri, L. & Giangaspero, A. Amphipathic, α-helical antimicrobial peptides. Biopolymers 55, 4–30, 10.1002/1097-0282(2000)55:1<4::aid-bip30>3.0.co;2-m (2000).
Lai, Z. H. et al. Highly Stabilized alpha-Helical Coiled Coils Kill Gram-Negative Bacteria by Multicomplementary Mechanisms under Acidic Condition. Acs Appl. Mater. Inter. 11, 22113–22128, https://doi.org/10.1021/acsami.9b04654 (2019).
Chou, H. T. et al. Design and synthesis of cationic antimicrobial peptides with improved activity and selectivity against Vibrio spp. Int. J. Antimicrob. Ag. 32, 130–138, https://doi.org/10.1016/j.ijantimicag.2008.04.003 (2008).
Chou, S. et al. Short, multiple-stranded beta-hairpin peptides have antimicrobial potency with high selectivity and salt resistance. Acta Biomater. 30, 78–93, https://doi.org/10.1016/j.actbio.2015.11.002 (2016).
Wang, J. et al. Antimicrobial peptides: Promising alternatives in the post feeding antibiotic era. Med. Res. Rev. 39, 831–859, https://doi.org/10.1002/med.21542 (2019).
Lyu, Y., Yang, Y., Lyu, X., Dong, N. & Shan, A. Antimicrobial activity, improved cell selectivity and mode of action of short PMAP-36-derived peptides against bacteria and Candida. Sci. Rep. 6, 27258, https://doi.org/10.1038/srep27258 (2016).
Zhu, X. et al. Design of imperfectly amphipathic alpha-helical antimicrobial peptides with enhanced cell selectivity. Acta Biomater. 10, 244–257, https://doi.org/10.1016/j.actbio.2013.08.043 (2014).
Dong, N. et al. Antimicrobial potency and selectivity of simplified symmetric-end peptides. Biomater. 35, 8028–8039, https://doi.org/10.1016/j.biomaterials.2014.06.005 (2014).
Ma, Z. et al. Characterization of cell selectivity, physiological stability and endotoxin neutralization capabilities of alpha-helix-based peptide amphiphiles. Biomater. 52, 517–530, https://doi.org/10.1016/j.biomaterials.2015.02.063 (2015).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (31272453, 31472104, 31672434, 31872368), the China Agriculture Research System of China (CARS-35) and. and the Natural Science Foundation of Heilongjiang Province (TD2019C001).
Author information
Authors and Affiliations
Contributions
A.S. and L.X. conceived and designed the experiments; L.X., C.S. and G.L. conducted the main experiments assays; S.C., C.S. and J.W. conducted flow cytometry assay and Cytoplasmic membrane depolarization assay; Q.M. and N.D. provided professional advices; C.S. analysed the data; L.X. and A.S. wrote the main manuscript text; All authors discussed the results and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, L., Shao, C., Li, G. et al. Conversion of Broad-Spectrum Antimicrobial Peptides into Species-Specific Antimicrobials Capable of Precisely Targeting Pathogenic Bacteria. Sci Rep 10, 944 (2020). https://doi.org/10.1038/s41598-020-58014-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-58014-6
This article is cited by
-
In Silico and In Vitro Analyses Reveal Promising Antimicrobial Peptides from Myxobacteria
Probiotics and Antimicrobial Proteins (2023)
-
The future of recombinant host defense peptides
Microbial Cell Factories (2022)
-
Combining genetic algorithm with machine learning strategies for designing potent antimicrobial peptides
BMC Bioinformatics (2021)
-
Antibiofilm activity of host defence peptides: complexity provides opportunities
Nature Reviews Microbiology (2021)
-
Helicobacter pylori treatment in the post-antibiotics era—searching for new drug targets
Applied Microbiology and Biotechnology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.