Abstract
Despite significant progress by genome-wide association studies, the ability of genetic variants to conduce to the prediction or prognosis of type-2 diabetes (T2D) is weak. Expression analysis of the corresponding genes may suggest possible links between single-nucleotide polymorphisms and T2D phenotype and/or risk. Herein, we investigated the expression patterns of 24 T2D-susceptibility genes, and their individual transcript variants (tv), in peripheral blood of T2D patients and controls (CTs), applying RNA-seq and real-time qPCR methodologies, and explore possible associations with disease features. Our data revealed the deregulation of certain transcripts in T2D patients. Among them, the down-regulation of CAPN10 tv3 was confirmed as an independent predictor for T2D. In patients, increased expression of CDK5 tv2, CDKN2A tv3 or THADA tv5 correlated positively with serum insulin levels, of CDK5 tv1 positively with % HbA1c levels, while in controls, elevated levels of TSPAN8 were associated positively with the presence of T2D family history. Herein, a T2D-specific expression profile of specific transcripts of disease-susceptibility genes is for the first time described in human peripheral blood. Large-scale studies are needed to evaluate the potential of these molecules to serve as disease biomarkers.
Similar content being viewed by others
Introduction
Type-2 diabetes mellitus (T2D), a chronic metabolic disorder with increased cardiovascular morbidity and mortality, accounts currently for one of the global epidemics with ever growing prevalence1. Despite recent advances in T2D diagnosis and management, challenges in its prevention and treatment still remain2.
T2D epidemic is mainly ascribed to the continuous increase in obesity globally, favored nowadays by the adoption of a sedentary lifestyle2, while the risk for T2D development depends also on genetic components. During the last decade, over 60 genome-wide association studies (GWAS) revealed more than 250 single nucleotide polymorphisms (SNPs) related to T2D or glycemic traits3. However, each of them individually increases disease risk with rather modest effect sizes (25–40% increase in the homozygous state for the genes conveying the greatest risk)4, which are further weakened when introduced in multivariate analysis models5.
The implication of the genome in the development of human disorders can be elucidated through the study of the transcriptome, given that the last reflects functionality6,7,8,9. Recent advances in transcriptome analysis provide key-data for (i) the link between genotype and phenotype, (ii) molecular networks underlying pathophysiological processes, and (iii) molecular fingerprints, paving the way for the identification of possible therapeutic targets and/or disease biomarkers7,10. Next-generation RNA-sequencing (RNA-Seq) has pivotally fashioned the mode of transcriptome profiling, giving the chance for gene-transcription levels and splicing isoforms to be detected and quantitated, in a high-throughput manner7,11,12.
The gene-expression signature of T2D, including the expression patterns of T2D-susceptibility genes, has been hardly investigated. Previous studies were confined to pancreatic islets or beta-cell lines from animal models or deceased human donors13, mainly due to difficulties in obtaining biopsy specimens from the T2D-target tissue(s) of living donors. However, recent evidence support that the gene-expression profile of peripheral blood cells reflects significantly (>80%) the gene-expression profile of other tissues, including disease-affected tissues, and that changes in the former mirror changes in the micro- and macro-environment of the latter14. Thus, peripheral blood is considered as a reliable alternative for the investigation of transcriptome dynamics of organ-specific and systemic diseases, as it is easily accessible, and provides data for pathophysiological processes taking place in various sites throughout the human body15.
Herein, we investigated the expression patterns of highly-related T2D-susceptibility genes in peripheral blood samples of patients and controls and explored possible associations with disease parameters and risk factors.
Materials and Methods
Study design
First, we developed a panel of highly-associated T2D-susceptibility genes. For the quantification of their expression, appropriate reverse transcription (RT) - real-time PCR (qPCR) protocols were developed and applied on RNA extracted from whole peripheral blood samples of T2D patients and controls (CT). RNA-Seq and specific qPCR protocols were utilized to identify specific transcript variants of these genes that are differentially expressed between the two groups. To examine specific distribution patterns in individuals at high risk of developing the disease, a distinct group of controls bearing T2D-risk factors was included in the total group of controls. The two subgroups were analyzed both together and separately. Finally, possible associations between the gene or transcript-variant expression levels and various disease parameters were explored.
Development of the T2D-susceptibility gene panel
The 24 highly-associated T2D-susceptibility gene panel was developed upon in-depth search in the NHGRI-EBI Catalog of published GWAS and SNPedia online databases3,16 (Table 1). CAPN10 is not included in GWAS-significant genes, however, in SNPedia it presents as carrying variants related to T2D in different populations, and thus it was included in the panel. CDK5 was also included, since it is highly regulated by the T2D-susceptibility gene CDKAL117.
Patients and samples
Peripheral blood samples were collected from 48 consecutive T2D patients and 40 control (CT) individuals (with normal glucose metabolism), upon informed written consent. The study was approved by the Ethics Committee of the Attikon Hospital (Athens, Greece). Both groups were characterized according to the current criteria for T2D diagnosis18. The medical records of the participants were evaluated for various clinical, laboratory, and therapeutic variables. The group of controls consisted of two distinct subgroups: controls without risk factors for the development of T2D (CTRF−; n = 17) and controls bearing risk factors for the disease (CTRF+; n = 23), as these were previously described by Nathan2 (Table 2).
RNA extraction, RT-qPCR and RNA-seq
All the methods used for gene-expression analysis, were applied on RNA extracted from whole peripheral blood using direct-blood lysis. For materials and protocols applied for RNA extraction, reverse transcription, and qPCR, as well as cDNA library construction and RNA-seq, see Supplemental Fig. 1. Specific primers designed for the amplification of the genes-of-interest, or certain transcript variants of them are reported in Supplemental Table 1. Relative quantification (RQ) of gene expression was performed by the 2−ΔΔCt method, using the hypoxanthine phosphoribosyltransferase 1 (HPRT1) gene, as endogenous reference gene for normalization purposes, and the immortalized 1.2B4 pancreatic beta-cell line (ECACC, Salisbury, UK), as our assay calibrator, for the calculation of the fold-changes. Representative amplification, melting and standard curves for certain genes/transcript variants are indicatively presented in Supplemental Fig. 2.
Bioinformatics analysis
For analysis of RNA-seq raw data see Supplemental Fig. 1. Differential expression between the two groups was considered significant if fold-change of their average RPKMs (CT:T2D ratio) was <0.5 or >2. Area-proportional Euler diagrams were generated using the BioVenn tool (http://www.biovenn.nl/). Analysis of tissue-specific expression patterns of genes and transcript variants and expression quantitative trait loci (eQTLs) was performed utilizing the portal of the Genotype-Tissue Expression (GTEx) project19 and the Blood eQTL browser20.
Statistical analysis
Differential expression patterns between CT and T2D, or among CTRF−, CTRF+, and T2D individuals, were explored using the non-parametric Mann–Whitney U or Jonckheere-Terpstra tests, respectively. Benjamini-Hochberg procedures for adjusting the false discovery rate (FDR = 0.25) in multiple comparisons were also applied. Possible associations with binary, ordinal or continuous values of various clinicopathological and laboratory parameters were investigated by Mann-Whitney U, Jonckheere-Terpstra, or Spearman’s rank correlation coefficient tests, respectively. Binomial multivariate logistic regression analysis was performed (enter model) using the occurrence of T2D as the dependent variable and the expression levels of genes and transcript variants, age and sex as independent variables. Analyses were performed using the softwares: GraphPad Prism 5 (GraphPad Software Inc., San Diego, CA, USA) or IBM SPSS Statistics 21 (IBM Corp., Armonk, NY, USA). P-values < 0.05 were considered significant.
Statement of Ethical Approval and Informed Consent
The study was approved by the Ethics Committee of the Attikon Hospital (Athens, Greece). All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2008 (5). Informed consent was obtained from all patients for being included in the study.
Results
Differential expression of certain T2D-susceptibility genes in patients versus controls
Firstly, specifically designed qPCR protocols applied on RNA extracted from whole peripheral blood samples, detected quantifiable expression levels in the cases of the following 20 genes: CAMK1D, CAPN10, CDC123, CDK5, CDKAL1, CDKN2A, CDKN2B, FTO, HHEX, IGF2BP2, JAZF1, KCNJ11, KCNQ1, NOTCH2, PPARG, SLC30A8, TCF7L2, THADA, TSPAN8 and WFS1. On the contrary, mRNA expression was not detected in samples of either patients or controls, in the cases of ADAMTS9, HNF1B, HNF4A and MTNR1B genes.
Relative quantification (RQ) values (median; range) in the groups of T2D patients (n = 48) and controls (n = 40) are summarized in Table 3. Mann-Whitney U test revealed that compared to controls, T2D patients expressed significantly higher levels of the genes CDK5 [p = 0.0056, RQ values (median; range) for T2D = 1.151 (0.600–8.103) and for CT = 0.945 (0.512–2.473), fold-change T2D vs. CT = 1.22], CDKN2A [p = 0.0411, RQ values (median; range) for T2D = 0.910 (0.320–4.030) and for CT = 0.655 (0.150–2.420), fold-change T2D vs. CT = 1.39] and TSPAN8 [p = 0.0055, RQ values (median; range) for T2D = 0.234 (0.0398–2.124) vs. CT = 0.159 (0.0247–1.132), fold-change T2D vs. CT = 1.47] (Table 3 and Fig. 1A; upper row). Further analysis within the group of CTs, revealed that CTRF+ individuals (n = 23) were characterized by elevated levels of the three abovementioned genes compared to CTRF− (n = 17) ones (Table 3 and Fig. 1A; lower row). Jonckheere-Terpstra test further confirmed this increase, revealing a gradual up-regulation in the mRNA levels of these genes among the groups of CTRF−, CTRF+ and T2D subjects [as for the CDK5 gene: p = 0.009, RQ values (median; range) for CTRF− = 0.919 (0.625–2.473), for CTRF+ = 1.005 (0.512–1.998) and for T2D = 1.151 (0.600–8.103), fold-change for CTRF+ vs. CTRF− = 1.09 and for T2D vs. CTRF+ = 1.15; as for the CDKN2A gene: p = 0.010, RQ values (median; range) for CTRF− = 0.426 (0.200–2.220), for CTRF+ = 0.920 (0.150–2.420) and for T2D = 0.910 (0.320–4.030), fold-change for CTRF+ vs. CTRF− = 2.16 and for T2D vs. CTRF+ = 0.99; as for the TSPAN8 gene: p = 0.001, RQ values (median; range) for CTRF− = 0.1071 (0.0247–1.132), for CTRF+ = 0.1894 (0.0741–0.832) and for T2D = 0.2340 (0.0398–2.124), fold-change for CTRF+ vs. CTRF− = 1.77 and for T2D vs. CTRF+ = 1.24) (Table 3 and Fig. 1A; lower row). Statistics for all comparisons are reported in Table 3. After applying correction for multiple comparisons, the differential expression patterns that remained significant were those of: CDK5 and TSPAN8 between CT and T2D groups, and of CDK5, CDKN2A and TSPAN8 among the CTRF−, CTRF+ and T2D groups.
(A) Dot-plots depicting the differential distribution of mRNA levels (RQ units) of CDK5, CDKN2A and TSPAN8 in controls (CT) and T2D patients (T2D) (upper row of panels), as well as among controls without T2D risk factors (CTRF−), controls with T2D risk factors (CTRF+) and T2D patients (lower row of panels), as attested by appropriate non-parametric tests. Mann-Whitney analysis revealed that T2D patients are characterized by higher mRNA levels of CDK5, CDKN2A and TSPAN8, compared to controls (upper row). Jonckheere-Terpstra test showed a stepwise increase in the total mRNA levels of the abovementioned genes among the CTRF−, CTRF+ and T2D groups (lower row). P-values are designated by asterisks (*p < 0.05, **p < 0.01, ***p < 0.001), whereas horizontal bars represent the median value of the group (B). Area-proportional Euler diagrams showing the differentially expressed genes and transcripts variants between the samples of controls (CT) and T2D patients (T2D), as analysed by RNA-seq. a. Diagrams representing the differentially expressed genes between T2D patients and controls, within the total 24 genes-of-interest (T2D-susceptibility genes; grey circle). Based on our data, T2D patients expressed exclusively higher levels of total mRNA of 3 genes (namely CAPN10, KCNQ1, TCF7L2) and lower levels of total mRNA of 2 genes (namely CDKAL1 and IGF2BP2), compared to controls, while mRNA expression of one gene (THADA) was found to be either up- or- downregulated in T2D versus CT groups. Further analysis of the individual gene-transcript variants revealed that the levels of 6 out of the 77 transcript variants of interest [NM_023085 (CAPN10), NM_000218 (KCNQ1), NM_001198530, NM_001146284, NM_001198527 (all of TCF7L2), NR_073394 (THADA)] were increased, while 8 [NM_017774 (CDKAL1), NM_001291873, NM_001291872, NM_001291875 (all of IGF2BP2), NM_001271643, NM_001271644, NM_001083953, NM_022065 (all of THADA)] decreased in patients versus controls. Differential expression was considered as fold-change of the relative expression levels (mean of reads per kilobase million, RPKM) between the two groups (CT:T2D ratio) <0.5 or >2. (C) Dot-plots depicting the differential distribution of mRNA levels (RQ units) of CAPN10 tv3, CDK5 tv1, CDKN2A tv3, CDKN2A tv4, IGF2BP2 tv7, KCNQ1 tv1 and TSPAN8 in controls (CT) and T2D patients (T2D) (upper row of panels), as well as among controls without T2D risk factors (CTRF−), controls with T2D risk factors (CTRF+) and T2D patients (lower row of panels), as attested by appropriate non-parametric tests. Mann-Whitney analysis revealed that T2D patients are characterized by higher mRNA levels of CDK5 tv1, CDKN2A tv3, CDKN2A tv4, IGF2BP2 tv7, and TSPAN8, while lower levels of CAPN10 tv3 and KCNQ1 tv1, compared to controls (upper row). Jonckheere-Terpstra test showed that the mRNA levels’ distribution of the abovementioned transcripts followed a linear trend among the CTRF−, CTRF+ and T2D groups: increase in the cases of CDK5 tv1, CDKN2A tv3, CDKN2A tv4, IGF2BP2 tv7, and TSPAN8 and decrease in the cases of CAPN10 tv3 and KCNQ1 tv1 (lower row). P-values are designated by asterisks (*p < 0.05, **p < 0.01, ***p < 0.001), whereas horizontal bars represent the median value of the group; D. Dot-plots depicting the differential distribution of mRNA levels (RQ units) of CDK5 tv2 and THADA tv5 in controls (CT) and T2D patients (T2D) (upper row of panels), as well as among controls without T2D risk factors (CTRF−), controls with T2D risk factors (CTRF+) and T2D patients (lower row of panels), as attested by appropriate non-parametric tests. Mann-Whitney analysis revealed that T2D patients are characterized by higher mRNA levels of CDK5 tv2 and THADA tv5 (upper row). More specifically, there is a significant difference of these tv levels between T2D individuals and the group of controls with T2D risk factors (CTRF+), while not with the controls without such factors (lower row). P-values are designated by asterisks (*p < 0.05), whereas horizontal bars represent the median value of the group.
RNA-Seq analysis
Following the quantification of the expression levels of the abovementioned 20 genes, we studied also the expression patterns of their individual transcript variants in order to: (i) detect the specific transcript variant(s) responsible for the aforesaid differences and (ii) reveal any possible “hidden” differences in the levels of specific variants of the rest 17 genes, in patients versus controls. For that reason, data from RNA-Seq performed on peripheral blood samples of representative T2D patients (n = 4) and controls (n = 2) were analyzed appropriately.
Focusing on the 24 genes-of-interest, T2D patients were found to express: (i) higher levels of the genes CAPN10, KCNQ1 and TCF7L2 and of the transcripts NM_023085 (CAPN10), NM_000218 (KCNQ1), NM_001198530, NM_001146284, NM_001198527 (all of TCF7L2), NR_073394 (THADA) (Fig. 1B and Supplemental Table 2) and (ii) and lower levels of the genes CDKAL1 and IGF2BP2 and of the transcripts NM_017774 (CDKAL1), NM_001291873, NM_001291872, NM_001291875 (all of IGF2BP2), NM_001271643, NM_001271644, NM_001083953, NM_022065 (all of THADA) (Fig. 1B and Supplemental Table 2), compared to controls.
Differential expression of certain transcript variants in T2D patients versus CT controls
After collecting together data from both the qPCR and RNA-Seq assays, we further evaluated in the total cohort of the study (48 T2D patients and 40 controls): (i) the levels of individual transcript variants of the genes found to be differentially expressed by qPCR (CDK5, CDKN2A, TSPAN8; Table 3 and Fig. 1A) and (ii) the levels of certain transcript variants found to be differentially expressed in RNA-Seq experiments (Fig. 1B and Supplemental Table 2). These transcript variants (tv) were the: NM_023085 (CAPN10 tv3), NM_004935.3 and NM_001164410.2 (CDK5 tv1 and 2, respectively), NM_000077.4; p16INK4A, NM_058197.4, NM_058195.3; p14ARF, and NM_001195132.1 (CDKN2A tv1, 3, 4 & 5, respectively), NM_001291872, NM_001291873 and NM_001291875 (IGF2BP2 tv4, 5 and 7, respectively), NM_000218 (KCNQ1 tv1), TCF7L2 tv4, 9 and 12 (NM_001146284, NM_001198527 and NM_001198530, respectively), NM_022065, NM_001083953, NM_001271643 and NM_001271644 (THADA tv1, 3, 4 and 5, respectively). The CDKAL1 and TSPAN8 genes, which have only one tv each (NM_017774 and NM_004616.2, respectively) were studied in the series of qPCR experiments described above. The NR_073394 non-coding tv of the THADA gene, was not selected to be further studied.
The levels (median; range) of the transcript variants in patient and control groups are reported in Table 3. Αs attested by Mann-Whitney U test, compared to controls, T2D patients expressed lower levels of CAPN10 tv3 [p = 0.0004, RQ levels (median; range) for T2D = 2.208 (0.313–12.43) and for CT = 3.924 (0.313–12.30), fold-change T2D vs. CT = 0.56] and of KCNQ1 tv1 [p = 0.0761, RQ levels (median; range) for T2D = 8.456 (2.880–16.93) and for CT = 9.408 (2.445–18.34), fold-change T2D vs. CT = 0.89] (Table 3 and Fig. 1C; upper row). Jonckheere-Terpstra test strongly supported these findings by revealing a significant gradual decrease among the CTRF−, CTRF+ and T2D groups in the levels of CAPN10 tv3 [p < 0.0005, RQ levels (median; range) for CTRF− = 5.098 (1.016–12.30), for CTRF+ = 3.405 (0.313–8.056), for T2D = 2.208 (0.313–12.43), fold-change for CTRF+ vs. CTRF− = 0.67 and for T2D vs. CTRF+ = 0.65] and KCNQ1 tv1 [p = 0.013, RQ levels (median; range) for CTRF− = 10.520 (7.504–18.34), for CTRF+ = 8.951 (2.445–16.34) and for T2D = 8.456 (2.880–16.93), fold-change for CTRF+ vs. CTRF− = 0.85 and for T2D vs. CTRF+ = 0.95] (Table 3 and Fig. 1C; lower row). On the other hand, compared to controls, patients exhibited higher levels of CDK5 tv1 [p = 0.0034, RQ levels (median; range) for T2D = 0.943 (0.494–8.233) and for CT = 0.745 (0.285–2.266), fold-change T2D vs. CT = 1.27], of CDKN2A tv3 [p = 0.0035, RQ levels (median; range) for T2D = 0.0722 (0.0062–3.481) and for CT = 0.0300 (0.0062–2.083), fold-change T2D vs. CT = 2.41), of CDKN2A tv4 (p = 0.0125, RQ levels (median; range) for T2D = 0.928 (0.292–3.300) and for CT = 0.669 (0.321–2.600), fold-change T2D vs. CT = 1.39) and of IGF2BP2 tv7 (p = 0.22, RQ levels (median; range) for T2D = 4.485 (1.032–14.14) and for CT = 3.903 (0.516–12.14), fold-change T2D vs. CT = 1.15) (Table 3 and Fig. 1C; upper row). Also, a significant gradual increase in the levels of these transcripts was observed among the groups of CTRF−, CTRF+ and T2D patients [for CDK5 tv1: p = 0.006, RQ levels (median; range) for CTRF− = 0.754 (0.509–2.266), for CTRF+ = 0.735 (0.285–1.441) and for T2D = 0.943 (0.494–8.233), fold-change CTRF+ vs. CTRF− = 0.97 and for T2D vs. CTRF+ = 1.28; for CDKN2A tv3: p = 0.002, RQ levels (median; range) for CTRF− = 0.0145 (0.0062–2.083), for CTRF+ = 0.0390 (0.0094–0.820) and for T2D = 0.0722 (0.0062–3.481), fold-change for CTRF+ vs. CTRF− = 2.67 and for T2D vs. CTRF+ = 1.85; for CDKN2A tv4: p = 0.001, RQ levels (median; range) for CTRF− = 0.570 (0.321–2.600), for CTRF+ = 0.850 (0.360–1.489) and for T2D = 0.928 (0.292–3.300), fold-change for CTRF+ vs. CTRF− = 1.49 and for T2D vs. CTRF+ = 1.09; for IGF2BP2 tv7: p = 0.050, RQ levels (median; range) for CTRF− = 3.670 (1.4040–12.14), for CTRF+ = 3.989 (0.5160–11.91) and for T2D = 4.485 (1.032–14.14), fold-change for CTRF+ vs. CTRF− = 1.09 and for T2D vs. CTRF+ = 1.12) (Table 3 and Fig. 1C; lower row).
A different distribution pattern was detected in the case of CDK5 tv2 and THADA tv5: T2D patients expressed elevated levels compared to controls [for CDK5 tv2: p = 0.0367, RQ levels (median; range) for T2D = 1.308 (0.631–3.843) and for CT = 1.117 (0.453–2.938), fold-change T2D vs. CT = 1.17; for THADA tv5: p = 0.0479, RQ levels (median; range) for T2D = 3.452 (0.718–120.7) and for CT = 2.582 (0.596–35.83), fold-change = 1.34) (Table 3 and Fig. 1D; upper row), though, the lowest levels were detected in CTRF+ individuals and intermediate values in CTRF− subjects [for CDK5 tv2: RQ levels (median; range) for CTRF− = 1.190 (0.788–2.938), for CTRF+ = 1.071 (0.453–2.139) and for T2D = 1.308 (0.631–3.843), fold-change for CTRF+ vs. CTRF− = 0.90 and for T2D vs. CTRF+ = 1.22; for THADA tv5: RQ levels (median; range) for CTRF− = 2.933 (1.128–35.83), for CTRF+ = 2.304 (0.596–16.01) and for T2D = 3.452 (0.718–120.7), fold-change CTRF+ vs. CTRF− = 0.79 and for T2D vs. CTRF+ = 1.50] (Table 3 and Fig. 1D; lower row).
Moreover, correction for multiple comparisons revealed statistically significant differences in the levels of CAPN10 tv3, CDK5 tv1, CDK5 tv2, CDKN2A tv3, CDKN2A tv4, and THADA tv5 between controls and T2D patients, and of CAPN10 tv3, CDK5 tv1, CDKN2A tv3, CDKN2A tv4, IGF2BP2 tv7, KCNQ1 tv1 among CTRF−, CTRF+ and T2D subjects.
Based on the above findings, the panel of the T2D-specific transcript variants finally included the: CAPN10 tv3, CDK5 tv1, CDK5 tv2, CDKN2A tv3, CDKN2A tv4, IGF2BP2 tv7, KCNQ1 tv1, THADA tv5 and TSPAN8. Among them, binomial multivariate analysis corrected for age and sex revealed that CAPN10 tv3 can predict T2D among participants of the current study (p = 0.022, OR = 0.726). A schematic representation of the T2D-specific transcript variants, also in comparison with the canonical transcript for each gene, is shown in Fig. 2.
Detailed gene-structure of the transcript (splice) variants of CDK5, CDKN2A, CAPN10, IGF2BP2, KCNQ1, THADA and TSPAN8 that were found to be differentially expressed in T2D. Exons are presented as boxes and introns as lines. Grey and white boxes represent coding and non-coding exons, respectively. The numbers within the boxes and above the lines indicate exon’s or intron’s length in nucleotides (nt). Arrows (↓) and asterisks (*) indicate the positions of the ATG starting codon and the stop codon (TGA or TAA or TAG), respectively. In each gene, the canonical (classic) and the differentially expressed transcript variants in T2D patients, indicated by arrow (→) are depicted. For each transcript variant, the GenBank® accession number, as well as the protein isoform and length in amino acids (aa) are shown.
Associations of mRNA levels with clinicopathological data
The levels of the T2D-specific transcripts were found to associate with various clinicopathological parameters (Supplemental Table 3). Notably, the associations revealed were different in patient versus control groups.
In detail, CDK5 tv1 levels correlated positively with serum insulin (μU/ml) and glycated haemoglobin (HbA1c; % or mmol/mol) levels and negatively with hyperlipidemia in T2D patients, while positively with serum triglycerides levels (mg/dl) (p < 0.05) in CTRF+ subjects. As for the CDK5 tv2 levels, these also correlated strongly with serum insulin levels, and moreover with the presence of central obesity in the T2D group (p < 0.05).
The levels of CDKN2A tv3 were as well significantly associated with serum insulin levels in T2D patients (p < 0.01), while these of CDKN2A tv4 correlated positively with BMI and waist-to-hip ratio and negatively with serum HDL levels (mg/dl) in the CTRF+ subgroup (p < 0.05).
Serum insulin levels in T2D patients associated also with THADA tv5 (p < 0.01) and TSPAN8 levels (p < 0.05), tended to correlate negatively with KCNQ1 tv1 levels (p = 0.06), while, in CTRF+ subjects, associated with IGF2BP2 tv7 levels (p < 0.05). Additionally, in T2D individuals, the levels of THADA tv5 correlated reversely with hyperlipidemia and those of IGF2BP2 tv7 positively with BMI (p < 0.05). In CTRF+ individuals, mRNA levels of TSPAN8 associated with T2D family history (p < 0.01), of IGF2BP2 tv7 with serum glucose levels, while in the total group of controls, KCNQ1 tv1 levels reversely with BMI, central obesity, glucose and LDL (mg/dl) levels (p < 0.05).
Furthermore, correction for multiple comparisons confirmed certain of the above correlations: a) In T2D subjects, serum insulin levels were associated with the levels of CDK5 tv1, CDK5 tv2, CDKN2A tv3, KCNQ1 tv1, THADA tv5 and TSPAN8, hyperlipidemia was associated with the levels of CDK5 tv1 and THADA tv5, and BMI with the levels of IGF2BP2 tv7; b) In control individuals, serum insulin levels were correlated with the levels of IGF2BP2 tv7, serum HDL levels, BMI, and waist-to-hip ratio with the levels of CDKN2A tv4, and family history of T2D with the levels of TSPAN8.
Various associations were also detected between clinicopathological data and the levels of transcripts which did not exhibit any differential distribution among the groups of patients and controls (data not shown).
Analysis of the tissue-specific expression pattern and eQTLs of the differentially expressed genes using public available datasets
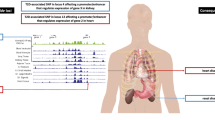
Based on public available data of the GTEx portal19, the CAPN10, CDK5, CDKN2A, IGF2BP2, KCNQ1, THADA and TSPAN8 genes, as well as their transcript variants CDK5 tv1, CDK5 tv2, IGF2BP2 tv7, KCNQ1 tv1 and THADA tv5, are expressed in a series of human tissues including blood and T2D-target tissues (adipose tissue, liver, skeletal muscle, pancreas) (Fig. 3). No data were available for CAPN10 tv3, CDKN2A tv3 and tv4.
Bar diagrams obtained from the GTEx portal (5/3/2018) depicting the expression pattern of the differentially expressed genes and transcript variants in various human tissues and cells. mRNA levels are expressed as transcripts per million (TPM). Boxes represent the median (range) of the normalized values. Boxes corresponding to whole blood or T2D-affected tissues are grey-colored highlighted.
Blood eQTL browser and GTEx portal were used for the analysis of eQTLs in the differentially expressed genes. For each one, a significant number of eQTLs appeared; however, we focused on eQTLs linked to T2D-related SNPs in blood or T2D-target tissues (Supplemental Table 4). Significant eQTLs related to T2D-SNPs were found on CAPN10, CDKN2A, IGF2BP2 and KCNQ1 in blood cells, and on CAPN10 and TSPAN8 in skeletal muscle. Affected genes are reported in Supplemental Table 4.
Also, CAPN10, the most significantly differentiated gene herein, is suggested to be an eGene (that is a gene having at least one cis-eQTL acting upon it), affected by several eQTLs on GPR35, RNPEPL1, and/or itself. Nevertheless, only the rs5030952-eQTL has a known (GWAS) association with T2D development (Supplemental Table 5).
Discussion
Despite numerous GWAS and other clinical studies revealing a large pool of SNPs associated with T2D, none of them has been yet proven promising for its diagnosis and/or prognosis5. Moreover, their causal relationship with T2D pathogenesis is not well-defined; epigenetic mechanisms can influence the expression of T2D-susceptibility genes, while DNA sequence itself alters the way the epigenome exerts its regulatory effect21,22,23,24,25. Furthermore, eQTLs associated with T2D-genetic variants may be also involved26. Herein, the expression profile of a panel of T2D-susceptibility genes, with special focus on their individual transcripts, was investigated in peripheral blood of T2D patients and controls, utilizing a combination of high-throughput and highly-sensitive molecular methodologies (RNA-Seq and qPCR).
Our data revealed a T2D-specific pattern of expression of nine transcript variants of the genes: CAPN10, CDK5, CDKN2A, IGF2BP2, KCNQ1, THADA and TSPAN8. Compared to controls, patients exhibited down-regulated levels of the tv3 of CAPN10 and tv1 of KCNQ1, while up-regulated levels of CDK5 tv1 and tv2, CDKN2A tv3 and tv4, IGF2BP2 tv7, THADA tv5 and TSPAN8. Publicly available datasets suggest that many human tissues including peripheral blood and T2D-target tissues express the abovementioned genes, allowing to postulate that the T2D-specific expression pattern found herein, may reflect pathogenetic mechanisms in disease-affected organs and/or peripheral blood14,15,19,20.
Among the down-regulated molecules is the tv3 of CAPN10. The gene encodes a protein implicated in glucose transporter 4 (GLUT4) translocation, insulin secretion, and apoptotic processes in pancreatic cells27. Compared to the canonical variant 1, tv3 lacks two consecutive exons, resulting in the loss of an in-frame segment in the 3’ coding region, and the encoded isoform (c) is shorter than isoform a (Fig. 2). The gene bears the rs3792267 and rs5030952 T2D-related SNPs3,16; however, tv3 does not bear any of them. Therefore, the decreased expression levels observed in patients, might be due to epigenetic and/or other transcriptional regulations24,25. It is worth mentioning that, herein, CAPN10 tv3 exhibited the highest association with the disease among all the deregulated molecules, as attested by both univariate and multivariate analysis. Yet, it is the only transcript which showed no association with any of the clinicopathological parameters tested, possibly indicating the highly complicated molecular networks underlying T2D.
The levels of the canonical transcript (tv1) of KCNQ1 (Fig. 2) were also decreased in the T2D group, while they were lower in the CTRF+ compared to the CTRF− group. ΚCNQ1 encodes the pore-forming subunit of a voltage-gated K+ channel (KvLQT1) that is essential for the repolarization phase of the action potential in cardiac muscle28. It is also expressed by pancreatic islets29, plays a key-role in the regulation of insulin secretion30 and its genetic variants have been associated with impaired insulin secretion in humans31,32. In these terms, the decreased expression of KCNQ1 tv1 in the CTRF+ and T2D groups observed in our study, and the reverse correlation with serum insulin levels, may indeed reflect the negative regulation on insulin secretory function exerted by KCNQ1 in patients and pre-disposed individuals.
Among the T2D-up-regulated molecules, there are both the two transcripts of CDK5 (Fig. 2). CDK5 is a serine/threonine protein kinase, involved in the degeneration of beta-cells and obstruction of insulin secretion through the generation of p35/CDK5 complexes33; its inhibition has been shown to protect these cells from glucotoxicity34. The overactivity of CDK5 and its activator p35 have been as well correlated with neuronal dysfunction in patients with Alzheimer’s disease (AD), and this could be one of the possible common mechanisms shared by these two degenerative disorders34. CDK5 is highly regulated by the T2D-susceptibility gene CDKAL117, through the rs7756992 SNP of the latter, which increases the risk for T2D35. Our data showed decreased levels of CDK5 tv1 and 2 in T2D patients compared to controls; though they exhibited different distribution patterns and correlated with different clinicopathological parameters in the CTRF− versus CTRF+ groups, but both with increased serum insulin levels. This might be probably attributed to different transcriptional regulations and distinct pathogenetic mechanisms leading, however, to the same “pre-disease” phenotype. Moreover, it is reported that as tv1, the tv2 of CDK5, in which an in-frame coding exon is skipped36 (Fig. 2), is also a negative regulator of Wnt/β-catenin signalling, a pathway involved in T2D development37.
SNPs in the CDKN2A/B locus were recently implicated in the negative regulation of beta-cell mass, proliferation and insulin secretory function, as well as in metabolic processes in adipose tissue, liver and muscles22. Also, in human islets, this locus is affected by epigenetic factors38, however, no effect on gene expression is known22. CDKN2A/B variants affect also the risk for cardiovascular disease39 and cancer40, and this could be a link for the common pathogenetic mechanisms shared with T2D41. Additionally, in blood, T2D-associated SNPs on CDKN2A are eQTLs which affect the expression of PSEN119 involved in AD and cancer42,43, connecting possibly these three morbidities. In this study, T2D patients expressed elevated levels of CDKN2A, and specifically of CDKN2A tv3 and 4: the first one highly associated with serum insulin levels in patients, and the second one with certain T2D-risk factors in controls. This may suggest their differential implication in disease development and/or progress. However, tv3 contains an alternative open reading frame (Fig. 2) and it is specifically expressed in the pancreas44. CDKN2A tv4 has a distinct first, but shares a common second exon with the canonical tv1, translated in different reading frames (Fig. 2): the encoded protein (p14ARF) lacks sequence similarity to the classic isoform (p16INK4a), and it is known to be nucleoplasmic but also recruited to mitochondria45. These characteristics may suggest tv-specific functions, possibly implicated in disease’s pathogenesis.
IGF2BP2 binds the 5′ UTR of the insulin-like growth factor 2 mRNA and regulates its translation16. Moreover, T2D-related SNPs on IGF2BP2 are eQTLs affecting SENP2, a gene crucially involved in adipogenesis and T2D development46. Herein, the levels of the tv7 of IGF2BP2 (which lacks exons 1 and 2 compared to the canonical tv1) exhibited a significant stepwise up-regulation from CTRF− to CTRF+ and to T2D individuals. Their correlation with BMI in patients, and serum glucose and insulin levels in CTRF+ cases, indicates its functional involvement in T2D pathogenesis.
Patients exhibited also elevated levels of TSPAN8 (only one known tv) and of THADA tv5 (with alternative 3′ coding region and 3′ UTR, encoding a shorter isoform (c) with a distinct C-terminus) (Fig. 2). The first gene is regarded as a prognostic factor and potential therapeutic target for certain human carcinomas47,48,49,50, while chromosomal aberrations of the second are observed in benign thyroid adenomas51. They both bear SNPs associated with T2D, though there is no knowledge regarding their involvement in its development3,16. Herein, the correlation of their levels with T2D, certain parameters and/or risk factors provides the first evidence for their possible implication in T2D pathogenesis.
The levels of TCF7L2, the most highly-related T2D-susceptibility gene, as well as of other T2D-susceptibility genes, were comparable between patients and controls; however, they correlated with certain disease characteristics or risk factors, supporting their implication in T2D development.
However, certain limitations of this study need to be considered: (a) the fact that not all the known T2D-susceptibilty genes are examined, (b) the relatively small number of participants and RNA-seq samples tested; the latter was overcome by the subsequent qPCR validation of the proposed deregulated transcript variants, discriminating between the true- and false-positive results, though, RNA-seq false-negative results could not been ruled out, (c) no paired-analysis of both the transcript levels and the presence of T2D-related SNPs was conducted; search throughout the Blood eQTL browser and the GTEx portal served only as a guide for their association and does not adequately explore the genetic determinants of the gene-expression variation, (d) the significant difference in the median age of patients and controls tested (Table 2); age associates with epigenetic changes52, thus it cannot be excluded as possible factor influencing the gene-expression variations observed in our cohort. However, after binomial multivariate analysis corrected for age and sex, CAPN10 tv3 still remain capable to predict T2D.
Nevertheless, by analyzing the expression patterns of a panel of the most highly-associated T2D-susceptibility genes, the current study offers suggestive data on the deregulated levels of certain transcript variants. Future research is required to elucidate their involvement in principal molecular and biochemical networks underlying T2D pathogenesis. Also, large-scale perspective clinical studies are needed to evaluate their potential to serve as possible biomarkers for its diagnosis, prognosis and/or monitoring.
Data Availability
The RNA-seq and qPCR raw data used to support the findings of this study are available from the corresponding author upon request.
References
International Diabetes Federation. IDF Diabetes Atlas (8th edn) Ch.2, 32–39, (International Diabetes Federation, 2017).
Nathan, D. M. Diabetes: Advances in Diagnosis and Treatment. Jama 314, 1052–1062, https://doi.org/10.1001/jama.2015.9536 (2015).
Burdett, T. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Research 45, D896–D901 (2017).
Bonnefond, A. & Froguel, P. Rare and common genetic events in type 2 diabetes: what should biologists know? Cell metabolism 21, 357–368, https://doi.org/10.1016/j.cmet.2014.12.020 (2015).
Lyssenko, V. & Laakso, M. Genetic screening for the risk of type 2 diabetes: worthless or valuable. Diabetes care 36(Suppl 2), S120–126, https://doi.org/10.2337/dcS13-2009 (2013).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature methods 5, 621–628, https://doi.org/10.1038/nmeth.1226 (2008).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nature reviews. Genetics 10, 57–63, https://doi.org/10.1038/nrg2484 (2009).
Streelman, J. T. & Kocher, T. D. From phenotype to genotype. Evolution & development 2, 166–173 (2000).
Faherty, S. L., Campbell, C. R., Larsen, P. A. & Yoder, A. D. Evaluating whole transcriptome amplification for gene profiling experiments using RNA-Seq. BMC biotechnology 15, 65, https://doi.org/10.1186/s12896-015-0155-7 (2015).
Jiang, Z. et al. Whole transcriptome analysis with sequencing: methods, challenges and potential solutions. Cellular and molecular life sciences: CMLS 72, 3425–3439, https://doi.org/10.1007/s00018-015-1934-y (2015).
Costa, V., Angelini, C., De Feis, I. & Ciccodicola, A. Uncovering the complexity of transcriptomes with RNA-Seq. Journal of biomedicine & biotechnology 2010, 853916, https://doi.org/10.1155/2010/853916 (2010).
Wilhelm, B. T. & Landry, J. R. RNA-Seq-quantitative measurement of expression through massively parallel RNA-sequencing. Methods 48, 249–257, https://doi.org/10.1016/j.ymeth.2009.03.016 (2009).
Segerstolpe, A. et al. Single-Cell Transcriptome Profiling of Human Pancreatic Islets in Health and Type 2 Diabetes. Cell metabolism 24, 593–607, https://doi.org/10.1016/j.cmet.2016.08.020 (2016).
Liew, C. C., Ma, J., Tang, H. C., Zheng, R. & Dempsey, A. A. The peripheral blood transcriptome dynamically reflects system wide biology: a potential diagnostic tool. The Journal of laboratory and clinical medicine 147, 126–132, https://doi.org/10.1016/j.lab.2005.10.005 (2006).
Mohr, S. & Liew, C. C. The peripheral-blood transcriptome: new insights into disease and risk assessment. Trends in molecular medicine 13, 422–432, https://doi.org/10.1016/j.molmed.2007.08.003 (2007).
Cariaso, M. & Lennon, G. SNPedia: a wiki supporting personal genome annotation, interpretation and analysis. Nucleic Acids Res 40, D1308–D1312 (2012).
Wei, F. Y. et al. Cdk5-dependent regulation of glucose-stimulated insulin secretion. Nature medicine 11, 1104–1108, https://doi.org/10.1038/nm1299 (2005).
Standards of Medical Care in Diabetes-2017. Summary of Revisions. Diabetes care 40, S18–S31 (2017).
The Genotype-Tissue Expression (GTEx) Project (2018). National Institutes of Health.
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nature genetics 45, 1238, https://doi.org/10.1038/ng.2756 (2013).
Kong, A. et al. Parental origin of sequence variants associated with complex diseases. Nature 462, 868–874, https://doi.org/10.1038/nature08625 (2009).
Kong, Y., Sharma, R. B., Nwosu, B. U. & Alonso, L. C. Islet biology, the CDKN2A/B locus and type 2 diabetes risk. Diabetologia 59, 1579–1593, https://doi.org/10.1007/s00125-016-3967-7 (2016).
Prasad, R. B. & Groop, L. Genetics of type 2 diabetes-pitfalls and possibilities. Genes 6, 87–123, https://doi.org/10.3390/genes6010087 (2015).
Hariharan, M., Scaria, V. & Brahmachari, S. K. dbSMR: a novel resource of genome-wide SNPs affecting microRNA mediated regulation. BMC bioinformatics 10, 108, https://doi.org/10.1186/1471-2105-10-108 (2009).
Fernandez-Valverde, S. L., Taft, R. J. & Mattick, J. S. MicroRNAs in beta-cell biology, insulin resistance, diabetes and its complications. Diabetes 60, 1825–1831, https://doi.org/10.2337/db11-0171 (2011).
Liang, X. et al. Integrating Genome-Wide Association and eQTLs Studies Identifies the Genes and Gene Sets Associated withDiabetes. BioMed research international 2017, 1758636, https://doi.org/10.1155/2017/1758636 (2017).
Ridderstrale, M. & Nilsson, E. Type 2 diabetes candidate gene CAPN10: first, but not last. Current hypertension reports 10, 19–24 (2008).
Barhanin, J. et al. K(V)LQT1 and lsK (minK) proteins associate to form the I(Ks) cardiac potassium current. Nature 384, 78–80, https://doi.org/10.1038/384078a0 (1996).
Demolombe, S. et al. Transgenic mice overexpressing human KvLQT1 dominant-negative isoform. Part I: Phenotypic characterisation. Cardiovascular research 50, 314–327 (2001).
Ullrich, S. et al. Effects of I(Ks) channel inhibitors in insulin-secreting INS-1cells. Pflugers Archiv: European journal of physiology 451, 428–436, https://doi.org/10.1007/s00424-005-1479-2 (2005).
Yasuda, K. et al. Variants in KCNQ1 are associated with susceptibility to type 2 diabetes mellitus. Nature genetics 40, 1092–1097, https://doi.org/10.1038/ng.207 (2008).
Mussig, K. et al. Association of type 2 diabetes candidate polymorphisms in KCNQ1 with incretin and insulin secretion. Diabetes 58, 1715–1720, https://doi.org/10.2337/db08-1589 (2009).
Ching, Y. P., Leong, V. Y., Wong, C. M. & Kung, H. F. Identification of an autoinhibitory domain of p21-activated protein kinase 5. The Journal of biological chemistry 278, 33621–33624, https://doi.org/10.1074/jbc.C300234200 (2003).
Ubeda, M., Rukstalis, J. M. & Habener, J. F. Inhibition of cyclin-dependent kinase 5 activity protects pancreatic beta cells from glucotoxicity. The Journal of biological chemistry 281, 28858–28864, https://doi.org/10.1074/jbc.M604690200 (2006).
Ohara-Imaizumi, M. et al. Deletion of CDKAL1 affects mitochondrial ATP generation and first-phase insulin exocytosis. PloS one 5, e15553, https://doi.org/10.1371/journal.pone.0015553 (2010).
Li, Q. et al. Characterization of a novel human CDK5 splicing variant that inhibits Wnt/beta-catenin signaling. Molecular biology reports 37, 2415–2421, https://doi.org/10.1007/s11033-009-9752-7 (2010).
Elghazi, L. et al. Importance of beta-Catenin in glucose and energy homeostasis. Scientific reports 2, 693, https://doi.org/10.1038/srep00693 (2012).
Popov, N. & Gil, J. Epigenetic regulation of the INK4b-ARF-INK4a locus: in sickness and in health. Epigenetics 5, 685–690 (2010).
Hannou, S. A., Wouters, K., Paumelle, R. & Staels, B. Functional genomics of the CDKN2A/B locus in cardiovascular and metabolic disease: what have we learned from GWASs? Trends in endocrinology and metabolism: TEM 26, 176–184, https://doi.org/10.1016/j.tem.2015.01.008 (2015).
Li, W. Q. et al. Genetic polymorphisms in the 9p21 region associated with risk of multiple cancers. Carcinogenesis 35, 2698–2705, https://doi.org/10.1093/carcin/bgu203 (2014).
Carling, D. AMPK signalling in health and disease. Current opinion in cell biology 45, 31–37, https://doi.org/10.1016/j.ceb.2017.01.005 (2017).
Rosenberg, R. N., Lambracht-Washington, D., Yu, G. & Xia, W. Genomics of Alzheimer Disease: A Review. JAMA neurology 73, 867–874, https://doi.org/10.1001/jamaneurol.2016.0301 (2016).
Telerman, A. & Amson, R. The molecular programme of tumour reversion: the steps beyond malignant transformation. Nature reviews. Cancer 9, 206–216, https://doi.org/10.1038/nrc2589 (2009).
Robertson, K. D. & Jones, P. A. Tissue-specific alternative splicing in the human INK4a/ARF cell cycle regulatory locus. Oncogene 18, 3810–3820, https://doi.org/10.1038/sj.onc.1202737 (1999).
Irvine, M. et al. Amino terminal hydrophobic import signals target thep14(ARF) tumor suppressor to the mitochondria. Cell cycle 9, 829–839, https://doi.org/10.4161/cc.9.4.10785 (2010).
Chung, S. S. et al. Control of adipogenesis by the SUMO-specific protease SENP2. Molecular and cellular biology 30, 2135–2146, https://doi.org/10.1128/MCB.00852-09 (2010).
Fekete, T. et al. Meta-analysis of gene expression profiles associated with histological classification and survival in 829 ovarian cancer samples. International journal of cancer 131, 95–105, https://doi.org/10.1002/ijc.26364 (2012).
Maisonial-Besset, A. et al. Tetraspanin 8 (TSPAN 8) as a potential target for radio-immunotherapy of colorectal cancer. Oncotarget 8, 22034–22047, https://doi.org/10.18632/oncotarget.15787 (2017).
El Kharbili, M. et al. Tetraspanin 8 is a novel regulator of ILK-driven beta1 integrin adhesion and signaling in invasive melanoma cells. Oncotarget 8, 17140–17155, https://doi.org/10.18632/oncotarget.15084 (2017).
Fang, T. et al. Tetraspanin-8 promotes hepatocellular carcinoma metastasis by increasing ADAM12m expression. Oncotarget 7, 40630–40643, https://doi.org/10.18632/oncotarget.9769 (2016).
Rippe, V. et al. Identification of a gene rearranged by 2p21 aberrations in thyroid adenomas. Oncogene 22, 6111–6114, https://doi.org/10.1038/sj.onc.1206867 (2003).
Fraga, M. F. et al. Epigenetic differences arise during the lifetime of monozygotic twins. Proceedings of the National Academy of Sciences of the United States of America 102, 10604–10609, https://doi.org/10.1073/pnas.0500398102 (2005).
Cheng, A. Y. et al. Implications of recent clinical trials for the National Cholesterol Education Program Adult Treatment Panel III guidelines. Current opinion in cardiology 21, 400–404, https://doi.org/10.1097/01.hco.0000231412.15049.fb (2006).
Expert Panel on Detection, Evaluation and Treatment of High Blood Cholesterol in Adults. Executive Summary of The Third Report of The National Cholesterol Education Program (NCEP) Expert Panel on Detection, Evaluation, And Treatment of High Blood Cholesterol In Adults (Adult Treatment Panel III). Jama 285, 2486–2497 (2001).
Acknowledgements
The authors would like to acknowledge Professor Sotirios A. Raptis, who passed away in 2017, for his invaluable contribution in the implementation of this study. Professor Raptis was pivotally involved, in terms of both scientific supervision and managerial support of the study. This research was co-funded by the European Union (European Regional Development Fund–ERDF) & Greek national funds through the Operational Program “Competitiveness & Entrepreneurship” of the National Strategic Reference Framework (NSFR) - Research Funding Program: SYNERGASIA (COOPERATION) 2011: “UtilizAtion of bioinfoRmatics and microchip Emerging Technologies towArds a nExt generation architectUre health system for diabetes mellitus type 2 management, prevention and perSonalized medical care - ARETAEUS” (#11SYN-10–1821).
Author information
Authors and Affiliations
Contributions
Study conception and design: E.G.F., M.I.C., A.S. Development of methodology: M.I.C., M.A., I.K. Acquisition of data: M.I.C., M.A., I.K., E.M., P.M. Analysis and interpretation of data: M.I.C., M.A., I.K., C.K.K., A.S., E.G.F. Statistical analysis: M.I.C. Enrollment of patients and controls: E.B., P.M. Drafting of the manuscript: M.I.C. Critical revision of the manuscript: E.G.F., A.S., M.A., M.I.C. Technical support: C.K.K., E.P. Study supervision: E.G.F., A.S.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Christodoulou, MI., Avgeris, M., Kokkinopoulou, I. et al. Blood-based analysis of type-2 diabetes mellitus susceptibility genes identifies specific transcript variants with deregulated expression and association with disease risk. Sci Rep 9, 1512 (2019). https://doi.org/10.1038/s41598-018-37856-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37856-1
This article is cited by
-
THADA inhibition in mice protects against type 2 diabetes mellitus by improving pancreatic β-cell function and preserving β-cell mass
Nature Communications (2023)
-
Implications of genetic variations, differential gene expression, and allele-specific expression on metformin response in drug-naïve type 2 diabetes
Journal of Endocrinological Investigation (2022)
-
Tissue, age, sex, and disease patterns of matrisome expression in GTEx transcriptome data
Scientific Reports (2021)
-
rs11927381 Polymorphism and Type 2 Diabetes Mellitus: Contribution of Smoking to the Realization of Susceptibility to the Disease
Bulletin of Experimental Biology and Medicine (2020)
-
Decreased expression of microRNAs targeting type-2 diabetes susceptibility genes in peripheral blood of patients and predisposed individuals
Endocrine (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.