Abstract
Peptidoglycan-binding protein-modified magnetic nanobeads (PGBP-MNBs) were prepared for efficient magnetic capturing of Staphylococcus aureus (S. aureus), which is associated with sepsis, using the binding affinity of PGBP for the peptidoglycan (PG) layer on S. aureus. These PGBP-MNBs can simply capture S. aureus in plasma within 1 hr or even 15 min. Importantly, they also can capture various types of Gram-positive bacteria, such as Bacillus cereus and methicillin-resistant and methicillin-susceptible S. aureus (MRSA and MSSA). We believe that PGBP-based systems will be used to develop diagnostic systems for Gram-positive bacteria-related diseases.
Similar content being viewed by others
Introduction
Staphylococcus aureus (S. aureus) is a widely distributed Gram-positive pathogen that causes many serious diseases in humans, such as local purulent infections, pneumonia and sepsis. Some of these diseases are life-threatening clinical syndromes associated with significant patient morbidity and mortality1,2,3,4,5,6,7,8,9,10,11. Therefore, rapid and sensitive detection of S. aureus has become crucial for improving patient survival rates12,13,14,15,16,17. Because S. aureus is hard to detect at low concentrations (e.g., ≤100 CFU/mL), long reaction times are usually needed before analysis. The most commonly used methods are culture-based assays, which are often composed of a series of steps, including selective culture enrichment, differential plating, and biochemical/serological testing. However, these assays are slow, expensive, time consuming and labor intensive because the assay procedure is complicated and requires amplification or enrichment of S. aureus in the sample18,19,20,21. In this study, we developed a novel concentration platform functionalized with peptidoglycan-binding proteins (PGBPs) for expedient capturing and enrichment of Gram-positive S. aureus.
Results and Discussion
The surface of the MNBs are modified with PGBP, enabling them to specifically recognize and strongly bind to the peptidoglycan (PG) layer on S. aureus (Fig. 1). PG is an essential component of the cell wall of all bacteria and is especially abundant in Gram-positive bacteria, in which it accounts for approximately half of the cell wall mass. It is well-known target for not only antibiotics but also the host immune response through recognition by pattern recognition receptors, including PGBPs22,23,24,25,26,27,28,29,30,31. Thus, PGBPs play a role in the recognition of PG in bacteria with nanomolar affinity. Thus, we developed MNBs modified with PGBP (PGBP-MNBs), and its PGBP is able to recognize S. aureus by binding to its PG layer like an antibody32,33,34. Furthermore, S. aureus captured by PGBP-MNBs was enriched by a magnetic field, allowing further analysis of whether a patient is infected with pathogens (S. aureus)35,36,37,38,39,40,41,42. We purified PGBP with a molecular weight of approximately 25 kDa, corresponding to approximately 421 amino acid. We also tagged PGBP with green fluorescent protein (GFP) to impart green fluorescence (Fig. 2a). The affinity of PGBP for the PG layer of Gram-positive S. aureus bacteria, which was measured as the binding affinity (KD), was determined using a BLItz system. The result showed that the KD was 6.49 nM (Fig. S2). In addition, we visually confirmed the specific binding affinity of PGBP for PG layer from S. aureus by fluorescence microscopy analysis. S. aureus bacteria stained with red fluorescent reagents were incubated with PGBP (green fluorescence). After 1 hr, the bacteria were purified by centrifugation to eliminate unbound PGBP and then were re-suspended in buffer containing 10% plasma, similar to physiological blood conditions in humans (Fig. 2b). As shown in Fig. 2c, the red and green fluorescence signals were colocalized, indicating that PGBP was bound to S. aureus. It was confirmed in fluorescence spectra as well that PGBP/S. aureus showed both green and red fluorescence intensities (Fig. S3). PGBP-MNBs were prepared for magnetic capturing of S. aureus at room temperature. The bacteria were then enriched by applying a magnet, as Ni-NTA was immobilized on the MNBs along with PGBP43,44,45,46. Histidine (His)-tags on PGBP act as a chelating agent and form chelate complexes with nickel (Ni) ions from Ni-NTA, which offer vacant electron orbitals to form coordinate bonds. The size of the MNBs before PGBP binding was approximately 900 nm, and their size after PGBP binding increased to approximately 1 μm (Fig. 3a). After reaction of the PGBP-MNBs with S. aureus in buffer containing 10% plasma for 1 hr, the resulting PGBP-MNBs/S. aureus complex was dropped on a glass slide, and then a magnet was placed under this glass slide. The complex was magnetically manipulated by the external magnetic field, showing a yellow fluorescence signal overlapped with both the red and green fluorescence signals (Fig. 3b,c). In addition, we recorded fluorescence spectra of the PGBP-MNBs/S. aureus complex at excitation wavelengths of 488 and 588 nm using a multimode-microplate reader to confirm the binding capacity of PGBP-MNB for S. aureus. Free PGBP, MNBs and S. aureus were also measured in the same manner as controls (Fig. S3). The PGBP-MNB/S. aureus complex showed fluorescence intensities corresponding GFP, which is similar to free PGBP (Figs 3d and S3). In addition, at the excitation wavelength of 588 nm, PGBP-MNBs/S. aureus exhibited fluorescence signals corresponding to S. aureus. As expected, as the reaction time increased, the fluorescence intensity under 588 nm excitation increased, indicating that the amount of S. aureus captured by the PGBP-MNBs was increased (Figs 3e and S3). Therefore, the ability of PGBP-MNBs to capture S. aureus could increase with the reaction time. Furthermore, for quick acquisition of S. aureus by magnetic concentration using these PGBP-MNBs, we evaluated the capturing abilities of S. aureus by the PGBP-MNBs with different reaction time and S. aureus concentrations by real-time PCR. First, S. aureus bacteria at concentrations of 105.7, 103.7 and 101.7 CFU/mL (according to the optical density at 600 nm) were incubated with PGBP-MNBs for various reaction times (0 min, 15 min, 30 min and 1 hr). Subsequently, the PGBP-MNBs magnetically captured S. aureus by forming PGBP-MNBs/S. aureus complexes, and Ct values and concentrations of the captured S. aureus were measured using real-time PCR46,47,48,49,50. As mentioned above, the longer the reaction time was, the higher the capturing ability (Fig. 4). After the PGBP-MNBs were mixed with S. aureus for 1 hr, a point magnet easily attracted S. aureus bacteria because their cell surfaces were bound to the PGBP-MNBs. Even at a S. aureus input concentration of 101.7 CFU/mL, approximately 101.38 CFU/mL S. aureus bacteria were captured by the PGBP-MNBs (Fig. 4b). Importantly, the PGBP-MNBs still showed sufficient ability as capturing probes at a short reaction time of 15 min. We further evaluated whether the PGBP-MNBs were capable of detecting Gram-positive bacteria regardless of species (Fig. 5). We chose S. aureus, Bacillus cereus, methicillin-susceptible Staphylococcus aureus (MSSA) and methicillin-resistant Staphylococcus aureus (MRSA) as Gram-positive bacteria with a thick PG layer as Gram-positive bacteria18,51,52. Notably, MRSA is one of the most common resistant S. aureus strains in hospitals. S. aureus and B. cereus were obtained from the Korean Collection for Type Cultures (KCTC), and clinically isolated MRSA (#77, #78, #79 and #80) and MSSA (#85, #86, #87 and #88) were obtained from BioNano Health Guard Research Center (H-GUARD). Bacteria (103.7 CFU/mL) were separately mixed with PGBP-MNBs for 1 hr at room temperature. After incubation, the unbound bacteria were removed by magnetically assisted washing, and then the captured bacterial concentrations were measured by real-time PCR assay. The data revealed the great efficacy of the PGBP-MNBs in magnetic capturing of approximately 102.8~104 CFU/mL Gram-positive bacteria with a capture efficiency of about 81.67%53. These results confirmed that PGBP-MNBs could be used universally to efficiently detect most Gram-positive bacteria.
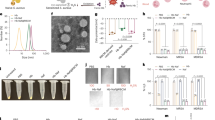
(a) Size measurement of MNBs and PGBP-MNBs by DLS analysis. (b) Illustration of magnetic separation of S. aureus using PGBP-MNBs. After binding with S. aureus and PGBP-MNBs for 1 hr, and (c) its fluorescence microscopic image (S. aureus: red and PGBP: green). Fluorescence spectra of PGBP-MNBs/S. aureus (d) at 488 nm (excitation) and (e) at 588 nm (excitation) under different reaction time (Circles: 1 hr and triangles: 2 hr), respectively.
Real-time PCR analysis of S. aureus magnetically captured using PGBP-MNBs at various bacterial concentrations (Input: 105.7, 103.7 and 101.7 CFU/mL) and reaction times (0 min, 15 min, 30 min and 1 hr). (a) Average threshold cycle value (Ct) of the captured S. aureus and (b) their corresponding output concentrations.
The concentrations of various types of bacteria (103.7 CFU/mL) magnetically captured using PGBP-MNBs after 1 hr of incubation, as determined by real-time PCR (S. aureus: Staphylococcus aureus, B. cereus: Bacillus cereus, MSSA: methicillin-susceptible Staphylococcus aureus and MRSA: methicillin-resistant Staphylococcus aureus). Peptidoglycan-binding protein-modified magnetic nanobeads (PGBP-MNBs) were prepared for efficient magnetic capturing of Staphylococcus aureus (S. aureus), which is associated with sepsis, using the binding affinity of PGBP for the peptidoglycan (PG) layer on S. aureus.
Conclusions
We developed PGBP-MNB as a magnetic capturing probe of Gram-positive S. aureus, which is associated with sepsis, and confirmed its ability to capture these bacteria within just 15 min at room temperature. In particular, since there is a PG layer on Gram-positive bacteria, the PGBP-MNBs can capture not only S. aureus but also B. cereus, MRSA and MSSA. Notably, because it uses a protein (PGBP) that binds universally to Gram-positive bacteria, this probe (PGBP-MNBs) has increased practicality compared to probes that increase selectivity by using an antibody that binds to a specific bacterium. Furthermore, we expect this PGBP-based capture platform to be applicable to diagnostic systems for bacteria-related diseases.
Experimental Section
Chemicals
We purchased HisPur™ Ni-NTA Magnetic Beads and BacLight Red Bacterial Stain from THERMO FISHER SCIENTIFIC (USA). SYBR Green PCR master mix was purchased from QIAGEN (Germany). Dulbecco’s phosphate buffered saline (DPBS, 1X) was purchased from GIBCO (Life Technologies). Human plasma (pooled normal, K2 EDTA) was purchased from INNOVATIVE RESEARCH INC. (USA). A Mini-BEST bacterial genomic DNA extraction kit was purchased from TAKARA (Japan). Primer sets for bacterial DNA amplification by real-time PCR were purchased from BIONEER INC. (Korea), and their sequences are summarized in Table S1.
Cloning, expression and purification of PGBP
We constructed a green fluorescence protein (GFP)-tagged peptidoglycan binding protein (PGBP) vector to express the fusion protein of PGBP and GFP (Fig. S1). The prepared expression vector was transformed into BL21 (DE3) component Escherichia coli expressing the recombinant protein, and the obtained transformant was inoculated into LB liquid medium supplemented with 50 μg/mL of ampicillin and then cultured at 37 °C until the optical density (OD) at 600 nm reached 0.6. After adding 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG), the cells were further cultured with shaking for 4 hr to obtain the recombinant protein (PGBP). To extract the expressed recombinant protein, 20 mM Tris-Cl (pH 8.0) and 0.2 M NaCl buffer solution was added to the E. coli cells, which were then recovered by centrifugation to suspend the cells and lysed using an ultrasonicator. The lysate was dissolved in an 8 M urea solution for efficient refolding of the insoluble protein, was subjected to metal affinity chromatography using 6X histidine as a metal affinity tag and was dialyzed against refolding solution (50 mM Tris-HCl (pH 8.5), 1 M arginine, 2 mM EDTA, 5 mM cysteamine, and 0.5 mM cystamine) at 4 °C for 48 hr under stirring. After sufficient refolding, the buffer of the recombinant protein was exchanged with PBS (pH 7.4) using ultrafiltration (MWCO: 10 KDa) and was concentrated to 1 mg/mL.
Bacterial culture and harvest conditions
The bacterial strains of Staphylococcus aureus (KCTC: 1621), Bacillus cereus (KCTC: 3624), methicillin-resistant Staphylococcus aureus (MRSA) (#77, #78, #79, and #80) and methicillin-susceptible Staphylococcus aureus (MSSA) (#85, #86, #87, and #88) were supplied by the Korean Collection for Type Cultures (KCTC), KRIBB (Korea) and BioNano Health Guard Research Center (H-GUARD) (Korea). All bacterial stains were cultured with LB broth (USA) at 37 °C. The bacterial concentration of S. aureus was determined by measuring the optical density (OD) at 600 nm (an OD600 of 1.0 Au = 8.8 × 108 CFU/mL).
Binding capacity of PGBP for S. aureus (PGBP/S. aureus)
We measured the binding affinity between PGBP and S. aureus using a biolayer interferometry-based biosensor BLItz system (FORTEBIO). PG from S. aureus was purchased from SIGMA-ALDRICH. PGBP loaded on Ni-NTA biosensors to bind with 6X histidine of PGBP and Ni-NTA, equilibrated in PBS for 1 min to establish a stable baseline, and then dipped into 4uL of PG from S. aureus (0 ~ 50 nM) to obtain the association curve for 300 s. The dissociation curve was obtained for 300 s using dipping holder. Afterwards, Binding affinities were calculated by fitting the curves using BLItz software (Fig. S2). We also performed binding capacity of PGBP against S. aureus in a same manner. Moreover, to visually confirm the binding of PGBP for S. aureus (107 CFU/mL), we used a red fluorescent dye (BACLIGHT RED BACTERIAL STAIN) for bacterial staining (Ex: 581–596 nm/Em: 644 nm) (THERMO FISHER SCIENTIFIC). Then, bacteria were co-incubated with PGBP (1 mg/mL, 20 µL) for 1 or 2 hr at 37 °C. After incubation, unbound PGBPs were washed with PBS three times by centrifugation at 10,000 rpm for 10 min. The PGBP/S. aureus complex was observed by Jaewoo Lim using a fluorescent Microscope (EVOS Cell Imaging Systems, THERMO FISHER SCIENTIFIC).
Preparation of PGBP-magnetic nanobeads (PGBP-MNBs)
Ni-NTA magnetic beads (8 µL, 12.5 mg/mL) and 10 µL of PGBP (1 mg/mL) (the ratio of magnetic beads to PGBP was 1:0.1) were mixed in PBS (1 mL) and incubated overnight at 4 °C. Then, unbound PGBP was removed from the PGBP-MNBs by washing three times with PBS using a magnetic tube rack, and the PGBP-MNBs were redispersed in PBS. The PGBP-MNBs were stored at −20 °C before use.
Ability of PGBP-MNBs to capture S. aureus (PGBP-MNBs/S. aureus)
We prepared 101.7~105.7 CFU/mL S. aureus in human plasma solution (10% human plasma in PBS). Then, 1 mL of PGBP-MNBs (0.1 mg/mL) were injected into the bacterial sample, and then they (PGBP-MNBs/S. aureus) mixed for 15 min, 30 min and 1 hr, respectively. To remove unbound bacteria, the samples were washed three times with PBS using magnetic separation methods. Additionally, we confirmed the capturing ability using various Gram-positive bacteria, including S. aureus, B. cereus, MRSA and MSSA. In these experiments, each type of bacteria (103.7 CFU/mL) was separately mixed with the PGBP-MNBs (1 mL, 0.1 mg/mL) for 1 hr, and then unbound bacteria were removed by using magnetic separation methods.
Confirmation of the capture of S. aureus captured by PGBP-MNBs (PGBP-MNBs/S. aureus)
To confirm the capture ability of the PGBP-MNBs, we conducted flow cytometry analysis and real-time PCR. Flow cytometry analysis was performed using a FACSCalibur instrument (BECTON DICKINSON AND CO., USA) and software (WINMD) for data analysis. PGBP was tagged with GFP to impart green fluorescence, and S. aureus was stained with a red fluorescence dye. In addition, we carried out real-time PCR using a CFX96 Touch™ real-time PCR detection system (BIO-RAD LABORATORIES, USA) to measure the concentration of S. aureus captured by the PGBP-MNBs (PGBP-MNBs/S. aureus). The primer sets for amplification of S. aureus genomic DNA and control (16s rRNA) primer sequences were described in previous papers. PCR conditions were in accordance with the product manual of QuantiTect SYBR Green PCR kits (QIAGEN, Germany). We also measured the concentrations of the other types of captured bacteria (B. cereus, MRSA and MSSA) by real-time PCR in the same manner.
References
Abeyrathne, C. D. et al. Lab on a chip sensor for rapid detection and antibiotic resistance determination of Staphylococcus aureus. Analyst 141, 1922–1929, https://doi.org/10.1039/c5an02301g (2016).
Banada, P. P. et al. Highly sensitive detection of Staphylococcus aureus directly from patient blood. PLoS One 7, e31126, https://doi.org/10.1371/journal.pone.0031126 (2012).
Bhattacharya, D. et al. Detection of total count of Staphylococcus aureus using anti-toxin antibody labelled gold magnetite nanocomposites: a novel tool for capture, detection and bacterial separation. J. Mater. Chem. 21, 17273, https://doi.org/10.1039/c1jm12076j (2011).
Chang, Y. C. et al. Rapid single cell detection of Staphylococcus aureus by aptamer-conjugated gold nanoparticles. Sci Rep 3, 1863, https://doi.org/10.1038/srep01863 (2013).
Cho, J. H. et al. Human peptidoglycan recognition protein S is an effector of neutrophil-mediated innate immunity. Blood 106, 2551–2558, https://doi.org/10.1182/blood-2005-02-0530 (2005).
Corey, G. R. Staphylococcus aureus bloodstream infections: definitions and treatment. Clin. Infect. Dis. 48(Suppl 4), S254–259, https://doi.org/10.1086/598186 (2009).
David, M. Z., Dryden, M., Gottlieb, T., Tattevin, P. & Gould, I. M. Recently approved antibacterials for methicillin-resistant Staphylococcus aureus (MRSA) and other Gram-positive pathogens: the shock of the new. Int. J. Antimicrob. Agents 50, 303–307, https://doi.org/10.1016/j.ijantimicag.2017.05.006 (2017).
Dong, P., Zhou, Y., He, W. & Hua, D. A strategy for enhanced antibacterial activity against Staphylococcus aureus by the assembly of alamethicin with a thermo-sensitive polymeric carrier. Chem Commun (Camb) 52, 896–899, https://doi.org/10.1039/c5cc07054f (2016).
Dziarski, R. Peptidoglycan recognition proteins (PGRPs). Mol. Immunol. 40, 877–886, https://doi.org/10.1016/j.molimm.2003.10.011 (2004).
Dziarski, R. & Gupta, D. The peptidoglycan recognition proteins (PGRPs). Genome Biol 7, 232, https://doi.org/10.1186/gb-2006-7-8-232 (2006).
Fykse, E. M., Olsen, J. S. & Skogan, G. Application of sonication to release DNA from Bacillus cereus for quantitative detection by real-time PCR. J. Microbiol. Methods 55, 1–10, https://doi.org/10.1016/s0167-7012(03)00091-5 (2003).
Goldmann, O. & Medina, E. Staphylococcus aureus strategies to evade the host acquired immune response. Int. J. Med. Microbiol., https://doi.org/10.1016/j.ijmm.2017.09.013 (2017).
Bicart-See, A. et al. Rapid Isolation of Staphylococcus aureus Pathogens from Infected Clinical Samples Using Magnetic Beads Coated with Fc-Mannose Binding Lectin. PLoS One 11, e0156287, https://doi.org/10.1371/journal.pone.0156287 (2016).
Cooper, R. M. et al. A microdevice for rapid optical detection of magnetically captured rare blood pathogens. Lab Chip 14, 182–188, https://doi.org/10.1039/c3lc50935d (2014).
Didar, T. F. et al. Improved treatment of systemic blood infections using antibiotics with extracorporeal opsonin hemoadsorption. Biomaterials 67, 382–392, https://doi.org/10.1016/j.biomaterials.2015.07.046 (2015).
Kang, J. H. et al. An Engineered Human Fc-Mannose-Binding-Lectin Captures Circulating Tumor Cells. Advanced Biosystems 1, https://doi.org/10.1002/adbi.201700094 (2017).
Kang, J. H. et al. An extracorporeal blood-cleansing device for sepsis therapy. Nat. Med. 20, 1211–1216, https://doi.org/10.1038/nm.3640 (2014).
Graber, H. U. et al. Mastitis-related subtypes of bovine Staphylococcus aureus are characterized by different clinical properties. J. Dairy Sci. 92, 1442–1451 (2009).
Gu, H., Xu, K., Xu, C. & Xu, B. Biofunctional magnetic nanoparticles for protein separation and pathogen detection. Chem Commun (Camb), 941–949, https://doi.org/10.1039/b514130c (2006).
Guan, R., Malchiodi, E. L., Wang, Q., Schuck, P. & Mariuzza, R. A. Crystal structure of the C-terminal peptidoglycan-binding domain of human peptidoglycan recognition protein Ialpha. J. Biol. Chem. 279, 31873–31882, https://doi.org/10.1074/jbc.M404920200 (2004).
Guan, R. et al. Structural basis for peptidoglycan binding by peptidoglycan recognition proteins. Proc Natl Acad Sci USA 101, 17168–17173, https://doi.org/10.1073/pnas.0407856101 (2004).
Guk, K. et al. A facile, rapid and sensitive detection of MRSA using a CRISPR-mediated DNA FISH method, antibody-like dCas9/sgRNA complex. Biosens. Bioelectron. 95, 67–71, https://doi.org/10.1016/j.bios.2017.04.016 (2017).
Hassan, M. M., Ranzoni, A. & Cooper, M. A. A nanoparticle-based method for culture-free bacterial DNA enrichment from whole blood. Biosens. Bioelectron. 99, 150–155, https://doi.org/10.1016/j.bios.2017.07.057 (2018).
Heijenoort, J. V. Formation of the glycan chains in the synthesis of bacterial peptidoglycan. Glycobiology 11, 25R–36R (2001).
Kell, A. J. & Simard, B. Vancomycin architecture dependence on the capture efficiency of antibody-modified microbeads by magnetic nanoparticles. Chem Commun (Camb), 1227–1229, https://doi.org/10.1039/b617427b (2007).
Kell, A. J. et al. Vancomycin-Modified Nanoparticles for Efficient Targeting and Preconcentration of Gram-Positive and Gram-Negative Bacteria. ACS Nano 2, 1777–1788 (2008).
Kong, W. et al. Magnetic microspheres-based cytometric bead array assay for highly sensitive detection of ochratoxin A. Biosens. Bioelectron. 94, 420–428, https://doi.org/10.1016/j.bios.2017.03.025 (2017).
Lee, C., Kim, J., Shin, S. G. & Hwang, S. Absolute and relative QPCR quantification of plasmid copy number in Escherichia coli. J. Biotechnol. 123, 273–280, https://doi.org/10.1016/j.jbiotec.2005.11.014 (2006).
Lee, I. S. et al. Ni/NiO Core-Shell Nanoparticles for Selective Binding and Magnetic Separation of Histidine-Tagged Proteins. J. Am. Chem. Soc. 128, 10658–10659 (2006).
Lee, K. J. et al. Simple and rapid detection of bacteria using a nuclease-responsive DNA probe. Analyst 143, 332–338, https://doi.org/10.1039/c7an01384a (2017).
Lin, Y.-S., Tsai, P.-J., Weng, M.-F. & Chen, Y.-C. Affinity Capture Using Vancomycin-Bound Magnetic Nanoparticles for the MALDI-MS Analysis of Bacteria. Anal. Chem. 77, 1753–1760 (2005).
Liu, C., Xu, Z., Gupta, D. & Dziarski, R. Peptidoglycan recognition proteins: a novel family of four human innate immunity pattern recognition molecules. J. Biol. Chem. 276, 34686–34694, https://doi.org/10.1074/jbc.M105566200 (2001).
Liu, C. L. et al. The impact of mgrA on progression of Staphylococcus aureus sepsis. Microb. Pathog. 71-72, 56–61, https://doi.org/10.1016/j.micpath.2014.03.012 (2014).
Lu, X. et al. Peptidoglycan recognition proteins are a new class of human bactericidal proteins. J. Biol. Chem. 281, 5895–5907, https://doi.org/10.1074/jbc.M511631200 (2006).
McAdow, M., DeDent, H. K. K. A. C., Hendrickx, A. P. A., Schneewind, O. & Missiakas, D. M. Preventing Staphylococcus aureus Sepsis through the Inhibition of Its Agglutination inBlood. PLoS Path. 7, e1002307–e1002319, https://doi.org/10.1371/journal.ppat.1002307.t001 (2011).
Meng, X. et al. Sensitive Detection of Staphylococcus aureus with Vancomycin-Conjugated Magnetic Beads as Enrichment Carriers Combined with Flow Cytometry. ACS Appl Mater Interfaces 9, 21464–21472, https://doi.org/10.1021/acsami.7b05479 (2017).
Ocsoy, I. et al. DNA aptamer functionalized gold nanostructures for molecular recognition and photothermal inactivation of methicillin-Resistant Staphylococcus aureus. Colloids Surf. B. Biointerfaces 159, 16–22, https://doi.org/10.1016/j.colsurfb.2017.07.056 (2017).
Royet, J. & Dziarski, R. Peptidoglycan recognition proteins: pleiotropic sensors and effectors of antimicrobial defences. Nat. Rev. Microbiol. 5, 264–277, https://doi.org/10.1038/nrmicro1620 (2007).
Royet, J., Gupta, D. & Dziarski, R. Peptidoglycan recognition proteins: modulators of the microbiome and inflammation. Nat. Rev. Immunol. 11, 837–851, https://doi.org/10.1038/nri3089 (2011).
Santos, S. R. D. et al. Scintigraphic imaging of Staphylococcus aureus infection using (99m)Tc radiolabeled aptamers. Appl. Radiat. Isot. 128, 22–27, https://doi.org/10.1016/j.apradiso.2017.06.043 (2017).
Shangguan, J. et al. A combination of positive dielectrophoresis driven on-line enrichment and aptamer-fluorescent silica nanoparticle label for rapid and sensitive detection of Staphylococcus aureus. Analyst 140, 4489–4497, https://doi.org/10.1039/c5an00535c (2015).
Shen, H. et al. Rapid and Selective Detection of Pathogenic Bacteria in Bloodstream Infections with Aptamer-Based Recognition. ACS Appl Mater Interfaces 8, 19371–19378, https://doi.org/10.1021/acsami.6b06671 (2016).
Shin, M. K. et al. Synthesis of Fe3O4@nickel-silicate core-shell nanoparticles for His-tagged enzyme immobilizing agents. Nanotechnology 27, 495705, https://doi.org/10.1088/0957-4484/27/49/495705 (2016).
Stegmiller, N. P. et al. Intranasal vaccination with adjuvant-free S. aureus antigens effectively protects mice against experimental sepsis. Vaccine 34, 3493–3499, https://doi.org/10.1016/j.vaccine.2016.04.018 (2016).
Suaifan, G. A., Alhogail, S. & Zourob, M. Rapid and low-cost biosensor for the detection of Staphylococcus aureus. Biosens. Bioelectron. 90, 230–237, https://doi.org/10.1016/j.bios.2016.11.047 (2017).
Suzuki, M. T. & Giovannoni, S. J. Bias Caused by Template Annealing in the Amplification of Mixtures of 16sRNA genes by PCR. Appl. Environ. Microbiol. 62, 625–630 (1996).
Wang, C. & Irudayaraj, J. Multifunctional magnetic-optical nanoparticle probes for simultaneous detection, separation, and thermal ablation of multiple pathogens. Small 6, 283–289, https://doi.org/10.1002/smll.200901596 (2010).
Wang, Y. et al. Ultrasensitive Rapid Detection of Human Serum Antibody Biomarkers by Biomarker-Capturing Viral Nanofibers. ACS Nano 9, 4475–4483 (2015).
Xu, H. et al. Gold-nanoparticle-decorated silica nanorods for sensitive visual detection of proteins. Anal. Chem. 86, 7351–7359, https://doi.org/10.1021/ac502249f (2014).
Yang, S. et al. Dual-recognition detection of Staphylococcus aureus using vancomycin-functionalized magnetic beads as concentration carriers. Biosens. Bioelectron. 78, 174–180, https://doi.org/10.1016/j.bios.2015.11.041 (2016).
Zhang, X., Hu, X. & Rao, X. Apoptosis induced by Staphylococcus aureus toxins. Microbiol. Res. 205, 19–24, https://doi.org/10.1016/j.micres.2017.08.006 (2017).
Zhu, M. et al. Construction of Fe3O4/Vancomycin/PEG Magnetic Nanocarrier for Highly Efficient Pathogen Enrichment and Gene Sensing. ACS Appl Mater Interfaces 7, 12873–12881, https://doi.org/10.1021/acsami.5b02374 (2015).
Chen, J. et al. Bacteriophage-based nanoprobes for rapid bacteria separation. Nanoscale 7, 16230–16236, https://doi.org/10.1039/c5nr03779d (2015).
Acknowledgements
This work was supported by the Center for BioNano Health-Guard funded by the Ministry of Science and ICT (MSIT) as Global Frontier Project (H-GUARD_2014M3A6B2060507 and H-GUARD_2013M3A6B2078950), the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by MSIT (NRF-2018M3A9E2022821), the Basic Science Research Program of the NRF funded by MSIT (NRF-2018R1C1B6005424), and KRIBB Research Initiative Program.
Author information
Authors and Affiliations
Contributions
Dr. E.-K.L. and Dr. J.J. conceived and designed the experiments. J.L., J.C. and D.K.L. performed all the experiments. Dr. S.-J.Y. provided PCR primers for bacterial detection and S.U.S., K.G. and Dr. T.K. analyzed the data for this manuscript. Figures 1, 2 and 3 were drawn directly by J.L. Dr. E.-K.L. and J.L. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lim, J., Choi, J., Guk, K. et al. Peptidoglycan binding protein (PGBP)-modified magnetic nanobeads for efficient magnetic capturing of Staphylococcus aureus associated with sepsis in blood. Sci Rep 9, 129 (2019). https://doi.org/10.1038/s41598-018-37194-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37194-2
This article is cited by
-
Sorghum Water Extract Application Mediates Antioxidant Defense and Confers Drought Stress Tolerance in Wheat
Journal of Plant Growth Regulation (2022)
-
Antibiotic-Based Magnetic Nanoprobes Combined with mPCR for Simultaneous Detection of Staphylococcus aureus and Bacillus cereus
Food Analytical Methods (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.