Abstract
The TMPRSS2:ERG gene fusion is the most prevalent early driver gene activation in prostate cancers of European ancestry, while the fusion frequency is much lower in Africans and Asians. The genomic characteristics and mechanisms for patients lacking ERG fusion are still unclear. In this study, we systematically compared the characteristics of gene fusions, somatic mutations, copy number alterations and gene expression signatures between 201 ERG fusion positive and 296 ERG fusion negative prostate cancer samples. Both common and group-specific genomic alterations were observed, suggesting shared and different mechanisms of carcinogenesis in prostate cancer samples with or without ERG fusion. The genomic alteration patterns detected in ERG-negative group showed similarities with 77.5% of tumor samples of African American patients. These results emphasize that genomic and gene expression features of the ERG-negative group may provide a reference for populations with lower ERG fusion frequency. While the overall expression patterns were comparable between ERG-negative and ERG-positive tumors, we found that genomic alterations could affect the same pathway through distinct genes in the same pathway in both groups of tumor types. Altogether, the genomic and molecular characteristics revealed in our study may provide new opportunities for molecular stratification of ERG-negative prostate cancers.
Similar content being viewed by others
Introduction
Prostate cancer is the second most commonly diagnosed cancer type in men globally and the fifth leading cause of cancer death, accounting for 6.6% of death among men1. Significant efforts have been made to characterize recurrent genomic alterations in prostate cancers, which may be potential driver events2,3,4,5. The overall mutation burden in prostate cancer is relatively low (0.3–2 non-synonymous somatic mutations per megabase) compared to other cancer types2,6,7. The most common genomic alteration is the fusion of 5′-UTR of TMPRSS2 (21q22) with 3′-end of ETS family members, such as ERG (21q22), ETV1 (7p21), ETV4 (17q21), or ETV5 (3q27)8,9,10,11. Significantly mutated genes include SPOP, FOXA1, TP53, MED12, and CDKN1B2,5,12. In addition to somatic mutations, somatic copy number alterations (SCNA) are recurrently seen in prostate cancer, including the amplification of chromosome 7 and 8q (affecting the MYC locus), and the focal deletion of chromosome 1q42, 3p13 (FOXP1), 4p15, 6q12–22 (MAP3K7), 8p, 13q, 16q, 17p (TP53), 18q12, and 21q22.3 (TMPRSS2-ERG fusion)5,7,12,13. However, there is still a large proportion of prostate cancer genomes that remains to be evaluated5,14,15.
Further studies confirmed that the TMPRSS2-ERG fusion is caused by an interstitial deletion on chromosome 21 or by a chromosomal translocation. These genomic rearrangements results in the overexpression of the ERG oncogene and ERG oncoprotein16,17. A variety of biological processes and pathways including cell invasion, Androgen receptor (AR) signaling, Transforming growth factor beta 1 (TGF-β) signaling have been implicated in ERG dysregulation18,19,20,21,22. ERG oncogenic activation is an early causal event in prostate cancer23,24,25. In some reports TMPRSS2-ERG fusion is positively correlated with advanced tumor stage, high Gleason score, and worse survival17,26,27,28,29,30. While some studies did not found significant association between ERG fusion and disease progression26,31,32,33,34, numerous studies reported positive correlation of ERG-negative prostate tumor type with disease progression35,36,37.
Since TMPRSS2-ERG fusion is a dominant molecular subtype in prostate cancer in European descents, it provides opportunities for targeted cancer therapy. Along these lines, direct and indirect ERG targeted therapeutic approaches are being developed38,39,40. Patients harboring ERG oncoprotein positive tumors are more likely to benefit from ERG targeted therapy. However, the frequency of TMPRSS2-ERG fusion significantly varies in different ethnic groups41. African American (20%~30%) and Asian (less than 20%) has much lower fusion frequency compared to Caucasian (~50%)42,43,44. In contrast to ERG fusion positive tumors, the genomic characteristics are not yet clear for the ERG fusion negative tumor type45. Therefore, identification of driver events in ERG-negative prostate cancer is important for understanding the mechanism of tumorigenesis.
In this study, we systematically explored the genomic and molecular differences of gene fusions, somatic mutations, SCNAs, gene expression signatures and dysregulation of pathways in prostate tumors with or without ERG fusion using publicly available data. Our results provide new insights into the molecular landscape highlighting specific mechanisms of prostate tumorigenesis.
Results
Data sources and the relationship between ERG fusion, deletion, and expression
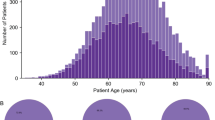
We collected the ERG fusion status information from two prostate cancer genome studies and compared the relationship among ERG fusion, deletion and expression5,46. The two datasets were highly consistent, except for 13 samples where the fusion status was unclear in the genomic data (Fig. 1a). We checked the ERG expression in these 13 samples and found significantly higher expression compared with ERG fusion negative samples (t.test, p-value = 0.001), indicating ERG may be activated in these 13 samples. Therefore, we assigned a sample into the ERG-positive group if its ERG fusion was detected in either study. As a result, we identified 201 ERG-positive samples and 296 ERG-negative samples for subsequent analysis (Supplementary Table 1). Also, we used ERG gene expression to verify the genomic classification of samples. Since, ERG fusion could result from either translocation or deletion at 21q22.316, we found that 40.8% ERG-positive samples harbored ERG deletion.
Classification of prostate cancer samples based on ERG fusion. (a) The relationship among ERG fusion, copy number variation, ERG mRNA expression and the final sample group. The top two rows show ERG fusion status based on the evidences from TCGA research article (333 samples) and TFGDP database respectively (See Method). (b) The proportion of ERG-positive and ERG-negative samples in different ethnic groups. AA: African American; CA: Caucasian American. Asian is not shown because of the small sample size.
Clinical characteristics of the ERG-positive and ERG-negative groups are summarized in Table 1. Although, patients with higher Gleason score (4 + 3 or 8–10) were more frequently found in the ERG-negative group, biochemical recurrence-free survival of patients showed no difference between the two groups (Supplementary Fig. 1, p-value = 0.29, Log-rank test). The TCGA prostate cancer cohort contained 279 Caucasian American (CA), 40 African American (AA), 5 Asian men and 173 without known ancestry. The proportion of ERG-positive samples in CA was higher than that in AA (47% vs. 35%, Fig. 1b), which is in accordance with previous studies (Supplementary Table 2). Like TCGA, most of the previous studies focused on patients of European ancestry. Indeed, more studies are needed for African and Asian patients that harbor mostly ERG-negative prostate cancers.
Common and specific genomic alterations in ERG-positive and ERG-negative prostate cancers
Gene fusions
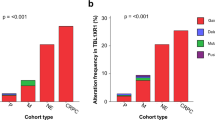
Consistent with previous studies47,48, in ERG-positive group, the most frequent fusion partner of ERG in our study was TMPRSS2 (94.1%), and the second was the SLC45A3 gene (6.4%, located at 1q32.1, Fig. 2a). These two genes both have AR responsive promoter and share similar mechanisms in ERG overexpression48. As expected, significantly higher ERG expression was detected in samples harboring SLC45A3:ERG fusion compared with samples with non-detectable ERG fusion (pvalue = 5e-5, one-tailed t.test). Other two ETS-family members, ETV1 and ETV4, show relatively high genomic rearrangement frequencies in ERG-negative group (4.7% and 2.7%, respectively). We found that the LSAMP gene that is frequently deleted in ERG-negative prostate tumors of African American men49, was often rearranged including fusion with ZBTB20 specifically in the ERG-negative group. Moreover, tumor suppressor gene MIPOL1 and TTC6 fusion were also specifically detected in the ERG-negative group at notable frequency (3.7%, Fig. 2a). Recent study of 65 Chinese prostate cancer whole genomes also reported TTC6:MIPOL1 fusion detected at 6.2% frequency44. Indeed, detection of TTC6:MIPOL1 fusion may have potential implication for prostate cancers of non-European ancestry. In addition, ten of eleven recurrent gene fusions (detected at least in three samples) have been reported in other literatures. Thus prostate cancer genomic fusions detected in our study, as well as in other reports are more likely real than false positives (Supplementary Table 3).
Comparision of the genomic characteristics between ERG-positive and ERG-negative groups. (a) The number of gene fusions in the two groups. Fusions present in more than two patients are shown. Red stars: fusions with significantly different frequency (p < 0.05, fisher test). (b) The frequency of significantly mutated genes in the two groups. Genes with significantly different mutation frequency in the two groups are marked with red stars. (p < 0.01, chi-squared test). The frequency of copy number amplification (c) and deletion (d) in two groups. Significantly altered cytobands separately detected in each group are annotated in black. Recurrently altered cytobands with significantly different frequency between the two groups are indicated in red. (chi-squared test, p < 0.01; frequency > 30%). Common (e) and group-specific (f) SCNA genes.
Somatic mutations
We used MutSigCV to identify significantly mutated genes in the ERG-positive and ERG-negative groups respectively50. Only two genes, TP53 and PTEN, were significantly mutated in ERG-positive group. By contrast, eight genes were significantly mutated in the ERG-negative group (Fig. 2b). In addition to known recurrently mutated genes SPOP and FOXA1 which were reported to be mutually exclusive with ERG rearrangements2,5, we found that the mutation frequency of CDK12 and KDM6A were significantly higher in the ERG-negative group (Fig. 2b, p-value = 1.18e-3 and 3.26e-4, respectively. Fisher.test).
Somatic copy number alterations
We applied the GISTIC algorithm to discern significant copy number alterations in the ERG-positive and ERG-negative groups51. First, we assessed the overall distribution of copy number alterations of all prostate cancer genomes in our study (Fig. 2c,d). Overall, deletions were more commonly than amplifications showing similar distribution in both ERG-positive and ERG-negative groups. Copy number alterations affected similar regions within the two groups, while deletion and amplification frequencies showed variations.
Twenty one amplified regions including chromosome 8q, 11q13, 14q21, 16q11, 1q22, 3q26 and 17q23, were recurrently altered in the ERG-negative group (Supplementary Table 4, residual q value < 0.05). The ERG-positive group harbored similar amplified regions, but did not reach statistical significance due to lower frequencies. Among the regions of copy number gains, chromosome 8q that includes the MYC oncogene exhibited a relatively high frequency (~40%). In another complex CNV region at 14q21.1 spanning MIPOL1/FOXA1/TTC6 locus, the MIPOL1:TTC6 gene fusion was detected. Moreover, we found several chromosome arm-level amplifications with significantly higher frequency in the ERG-negative tumors than in ERG-positives, including chromosome 8 (38.5% vs. 19%) and chromosome 7 (26.1% vs. 11.5%) (Fig. 2c).
Ten regions were commonly deleted in both ERG-positive and ERG-negative groups, including 6q14.3, 13q14.13, 10q23.31, 12p13.1, 5q11.2, 5q13.2, 17p13.1, and 16q22.3 (residual q < 0.05), which is consistent with previous reports5,13. Twenty two and twenty five copy number losses were detected only in the ERG-positive or in the ERG-negative group, respectively. Among these focal deleted regions, some showed significantly different frequency between the two groups. Similar to previous studies we also detected frequent deletions of 21q22 (ERG, TMPRSS2), 17p13.1 (TP53), and 10q23.31 (PTEN) in ERG-positive tumors, while 6q14.3 and 13q14.13 deletions were more frequent in ERG-negatives (Fig. 2d). Additionally, two novel regions, 6q16.3 (HACE1) and 6q22 (FRK) were deleted more frequently in the ERG-negative group.
To gain more insight into the functional effects of SCNA regions, we assessed the genomic defects of tumor suppressor genes (TSGs) and oncogenes (Supplementary Table 5). Thirty-two TSGs were recurrently altered with frequencies higher than 20% in both groups (Fig. 2e). Twenty-one (65.6%) of these genes were previously shown to play roles in the progression of prostate cancer. Other genes with high alteration frequencies need to be further defined. Thirteen TSGs and one oncogene showed significantly higher alteration frequency in the ERG-negative group, and another thirteen tumor suppressor genes and one oncogene showed significantly higher alteration frequency in ERG-positives (Fig. 2f). The candidate CNV genes found in TCGA dataset show comparable alteration frequency in an independent whole genome sequencing dataset, which includes 7 ERG-positive and 7 ERG-negative prostate tumors (Supplementary table 5, CPDR dataset). Among these group-specific SCNA genes, we found that ten genes were significantly associated with biochemical recurrence. In addition to previously reported disease progression related genes TP53, PTEN and FOXP1, we also found that an additional seven genes were associated with biochemical recurrence (Supplementary Fig. 2). Although ERG rearrangement status alone might not be a definitive marker for disease progression, our findings highlight a subset of genes associated with higher risk of disease progression (Overall prevalence: 44.87%). Furthermore, we found a group of tumor suppressor genes including FRK, WISP3, PRDM1, and LRP1B whose CNV and expression may indicate interactions with known drugs and therefore, are potentially actionable (Supplementary Fig. 3).
Candidate genes associate with genomic alteration patterns in ERG-negative prostate tumors
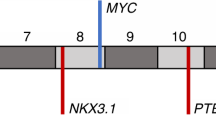
Since we have characterized both common and group-specific genomic alterations with high frequency in ERG-positive and ERG-negative prostate tumors, we next examined the molecular portrait of the ERG-negative group based on the associated candidate genes. First, we combined the genomic alterations of gene fusions, somatic mutations and copy number alterations which occur recurrently in the ERG-negative group. Next, we removed the redundant alterations to find a subset of genes highly represented in the genomic alteration pattern of ERG-negative tumors. Nine representative genes have emerged from the analysis (Fig. 3a). Genomic alteration of one or more of these nine genes were detected in 67.7% of the ERG-negative group.
Genomic portraits of ERG-negative group and similar patterns in African Americans. (a) Alteration pattern of the most representative genes in ERG-negative group, covering 67.7% samples. Red: genes with significantly higher alteration frequency in ERG-negative group. (b) Similar patterns were found in 40 AA samples with the coverage of 77.5%. (c) MYC amplification is significantly correlated with expression (p = 0.002, t-test) (d) Kaplan–Meier plot of biochemical relapse-free survival probabilities of patients with and without MYC amplification (p = 4.7e-05, Log-rank test). (e) The expression of ZNF292 is significantly lower in samples with deletion (p < 2.2e-16, t-test).
Since ERG-rearrangement are less frequent in prostate cancers of African descents, we explored whether candidate gene defects found in the ERG-negative group are present or absent in prostate cancers of AA men. As ERG is less frequent in prostate cancers of AA patients, we evaluated the alteration patterns of the nine genes characteristic to ERG-negative tumors in available datasets of 40 AA prostate tumor samples. We found that 77.5% AA tumors harbor at least one of the nine gene signatures associated with ERG-negative tumors indicating similar patterns between prostate cancers of AA patients and the overall genomic alteration pattern of ERG-negative tumors (Fig. 3b).
Among the nine representative genes, NKX3-1, RB1, and CDH13 were commonly deleted in both ERG-positive and ERG-negative tumors. Other genes had significantly more alterations in ERG-negative samples. The oncogene MYC mRNA is up-regulated in tumor compared to normal. Tumors with MYC amplification show significantly higher expression of MYC gene and higher probability of disease progression than other patients (Fig. 3c,d). The Zinc finger transcription factor, ZNF292 was shown to function as a tumor suppressor in gastric cancer, colorectal cancer, and chronic lymphocytic leukemia52,53. Deletion of ZNF292 in prostate cancer results in decreased expression (Fig. 3e), which may promote tumor development.
Comparison of methylation and expression between ERG-positive and ERG-negative tumors
Since promoter hypermethylation is widely observed in multiple cancers, we investigated the hypermethylated sites in promoter regions (TSS200, TSS1500, 5’UTR and 1stExon) of genes with low mRNA expression (See Method). Compared to normal samples, 2191 CpG sites (694 genes) and 1871 CpG sites (645 genes) were hyper-methylated in ERG-positive and ERG-negative groups, respectively. Approximately 70% of them were overlapped between the two groups (Supplementary Fig. 4a). Direct comparison between two tumor groups indicated 51 hyper-methylated sites (31 genes) in ERG-negative and 14 hyper-methylated sites (8 genes) in ERG-positive tumors (Supplementary Fig. 4b). Therefore, the overall methylation profiles showed similarities between the two groups.
We compared the expression profiles of ERG-positive, ERG-negative tumor and prostate tissue samples with morphologically normal appearance to identify differentially expressed genes among these three groups. A large proportion (>70%) of differentially expressed (DE) genes were common in the ERG-positive and negative groups (Fig. 4a). As expected, common DEs were significantly enriched in essential pathways like calcium signaling and cAMP signaling pathways (Fig. 4b). Common up-regulated genes were significantly enriched in cell cycle which is recurrently altered in cancer. However, no significant functional GO term was enriched for group-specific genes indicating comparable expression profiles between ERG-positive and ERG-negative prostate tumor types, despite in their differences in their dominant driver genomic alterations. These findings indicate that different genomic alternations may have similar effects on gene expression, resulting in similar phenotype.
Gene expression in ERG-positive and ERG-negative groups. (a) Venn diagram for differentially expressed genes (DEs) detected in ERG-positive and ERG-negative groups respectively compared to normal samples. 2020 DEs (>70%) are common. (b) The enriched KEGG pathways for the common DEs (FDR < 0.05). No significantly terms were enriched for group-specific DEs.
The impact of genomic alterations on pathway dysregulation in ERG-positive and ERG-negative prostate tumors
We selected eleven pathways either cancer-related or reported to be important in prostate cancer from Misgdb54,55,56. Next, we compared the frequency of CNV, somatic mutation and gene fusion of the ERG-positive and ERG-negative groups based on publicly available TCGA data. The male hormone axis (AR pathway) was the only node that altered significantly more frequently in ERG-positive group that is consistent with the AR regulation of ERG in the context of TMPRSS2:ERG fusion (Fig. 5a). However, there were still 65.3% of ERG-negative samples with AR pathway disruption, which were apparently affected by other genes in the AR pathway (Fig. 5b). For example, CDK6 (10.7% ERG-negative vs. 4% ERG-positive), NCOA2 (23.7% ERG-negative vs. 10.5% ERG-positive) and PRKDC (20.0% ERG-negative vs. 11.5% ERG-positive). Similarly, some component of NOTCH signaling pathway signatures had higher alteration frequency in the ERG-positive group (e.g., DVL2,11.3% ERG-negative vs. 25.0% ERG-positive) while HDAC2 (24.4% ERG-negative vs. 7.0% ERG-positive) had higher alteration frequency in ERG-negative group (Fig. 5c). They both inhibit NOTCH signaling pathway but function at different contexts. Therefore, the observed prostate cancer genomic and expression alterations of different genes may affect the same pathway resulting in comparable expression profiles between ERG-positive and ERG-negative prostate tumor types.
Genomic alterations show different preference on pathway dysregulation in ERG-positive and ERG-negative groups. (a) The overall alteration frequency on prostate cancer-related pathways. AR pathway show significant difference in two groups (p < 0.01). Genomic alterations in AR pathway (b) and NOTCH signaling pathway (c). Genes with significantly different alteration frequency in the two groups are shown. Red: higher frequency in ERG-positive group. Blue: higher frequency in ERG-negative group. Green: high expression in both groups.
Discussion
Our study provides new insights into the molecular landscape of ERG-negative prostate cancers. Except for known alterations mutually exclusive with ERG rearrangements, such as mutation in SPOP and FOXA1, we found that gene fusion of TTC6:MIPOL1 and somatic mutation on CDK12 and KDM6A occurred more frequently in the ERG-negative group. Recurrent gene fusions and somatic mutations could explain only a subset of ERG-negative tumors, noting that more of these genes harbor somatic copy number alterations. Some of them are shared between the two groups of tumors, others occurred more frequently in one group over the other. In addition to confirm several previous studies, we found novel recurrent SCNA for ERG-negative prostate cancers, such as ZNF292 deletion. In summary, the ERG-negative group was found more heterogeneous in our study.
When validated, the recurrently altered genes in specific patient groups may contribute to better tumor stratification and prognosis. Among these genes, MYC is a well-known oncogene that plays an important role in tumor progression. The amplification of MYC is frequently observed in numerous human cancers57. In this study, we found that MYC amplification frequency was significantly higher in the ERG-negative group. As expected, patients with tumors harboring MYC amplification show a strong association with poor outcome. Previous studies have reported that intact CHD1 is required for ERG rearrangements in the process of tumor initiation and deletion of CHD1 is mutually exclusive with ETS fusions58, that was consistently observed in our study. In addition to confirming known gene defects, we also identified several novel prostate cancer associated genes which may play important roles in the tumorigenesis of ERG-negative cancer type.
Our study highlights potentially actionable genes which may provide opportunities for target therapy of ERG-negative prostate tumors. These findings include the frequent deletion of the tumor suppressor gene FRK (6q22.1), a tyrosine-protein kinase that negatively regulates cell proliferation59, in ERG-negative group (22.3% vs. 8.0%). Decreased expression of FRK gene strongly correlated with its deletion. Moreover, FRK protein could interact with known drugs and may have potential application in clinical practice60. Other potentially druggable genes including WISP3 (6q21), LRP1B (2q22.1), and PRDM1 (6q21)60,61. In total, 21 (34.4%) genes in our candidate gene list have potential clinical relevance, covering 66.7% of ERG negative tumors.
Interestingly, we found that different gene alterations may result in similar expression change or pathway alteration. NOTCH signaling pathway is a typical example. Similar phenomenon has been observed in other cancer types. Taken Wnt signaling pathway as an example, TP53, CTNNB1 and AXIN1 are important elements in Wnt signaling network; CTNNB1 is more frequently mutated in HCV-infected hepatocellular carcinoma (HCC)62, while the mutations of TP53 and AXIN1 are more frequent in HBV-infected HCC63,64, which indicated different viral etiologies might activate Wnt signaling in distinct ways.
Increasing number of studies reports race/ethnicity differences in cancer research. Due to the lack of large-scale omics study of African and Asian prostate cancer patients, directly comparisons among multiple races are challenging. Our focuse on the ERG-negative group could provide a reference for populations with low frequency of ERG positive tumor types. Nine representative genes were sufficient to classify into sub-categories 67.7% ERG-negative tumors that was consistently seen in 77.5% of prostate cancers of African American men. Our previous studies found that approximately 20% Chinese patients harbor ERG-positive tumors41. Therefore we are particularly interested in the frequently altered and targetable genes in the ERG-negative tumor type. The validation of the genomic alteration and expression of these genes in Chinese patients is warranted. Accumulating data on ERG negative prostate cancer will help to discover more disease progression associated and actionable driver genes. Additionally, further experimental assessments of the functional significance for recurrent genomic and gene expression alterations are also warranted.
Our study highlights new aspects of ERG-positive and ERG-negative prostate cancers at genomic, epigenetic, and expression levels. In this study, multi-omics data integration provided a methodological reference to prioritize candidate CNV genes and to evaluate the effects of overall alterations. The observed molecular differences on gene fusions, somatic mutations and copy number alterations between ERG-positive and ERG-negative prostate tumors suggest both common and distinct mechanisms of prostate tumorigenesis. Genes with recurrent alteration may act as potential drivers and contribute to patient stratification into distinct prognostic or therapeutic groups. These results will help experimental biologist and clinical doctors for further assessment of the functional significance of candidate genes. Together, our results provide new insights into prostate tumorigenesis further refining the sub-classes of ERG-negative and ERG-positive prostate tumor types.
Methods
Data collection
Somatic mutation (496 tumor samples), SCNA (492 tumor samples), methylation (497 tumor + 35 normal samples), and expression (497 tumor + 52 normal samples) data from TCGA primary prostate cancer cohort were used in this study65. Clinically actionable genes and the interactions between genes and drugs were retrieved from DGIdb (http://dgidb.org/)60.
Patient group and ethnic information
Samples were stratified into ERG-positive and ERG-negative groups based on the combined ERG fusion evidences from TCGA research article (333 samples) and TFGDP database (http://www.tumorfusions.org/, 502 samples)5,46. A patient was assigned to ERG-positive group if its ERG fusion was detected in either study. For genome wide fusion analysis and statistics except for ERG fusion, data from TFGDP database was used. The ethnic information was collected from literature in which G. Petrovics et al. determine the ancestry of TCGA cohort by principal component analysis based on SNP genotype data49.
Detection of significantly mutated genes and copy number alterations
We used MutSigCV (version: 1.2) to detect significantly mutated genes for ERG-positive and ERG-negative groups, respectively50. Chi-squared test and Fisher exact test (determined by theoretical frequencies and sample size) were used to test the significance of different alteration frequency between the two groups. GISTIC 2.0 (version 6.10) was used to identify genomic regions that are significantly amplified or deleted in ERG-positive and ERG-negative groups, respectively51,66. To find the common and specifically altered regions in the two groups, we divided the whole genome into consecutive bins (window length = 10 kb). For each bin, the SCNA status is determined by the SCNA status of majority of bases in it (that is, longer than 5 kb). For arm-level SCNA regions, the frequency was estimated by the median frequencies of all bins in that region.
Since the significant SCNA regions usually contained huge genes, we focused on the copy number alterations of tumor suppressor genes (TSGs) and oncogenes. We obtained 1217 TSGs and 232 oncogenes from TSGene Database (v2.0) and UniProtKB database (keyword:“Proto-oncogene [KW-0656]”)67,68. These genes were classified into two types based on the following filtering rules: 1) Common SCNA genes: high frequency (>20%) in both ERG-positive and ERG-negative groups; 2) Group-specific SCNA genes: TSGs (Oncogenes) whose deletion (amplification) frequencies were significantly different between two groups (P < 0.001) and the frequency difference was larger than 10%.
Selection of representative genes for ERG-negative group
We used genes with recurrent SCNA (frequency >15%) or mutation (frequency >10%) as candidate feature genes for ERG-negative group. We defined a group of genes with higher priority: genes whose alteration frequency were significantly higher in ERG-negative group than that in ERG-positive group, genes which were targetable or had interaction with drugs, and genes whose copy number alteration was significantly correlated with expression. To remove genes with similar alteration pattern, we calculated Pearson correlation coefficient between genes and did unsupervised hierarchical clustering. For each cluster, we selected genes with the highest frequency or higher priority as the representative genes. At last, six CNV genes and three mutation genes were selected as final representative feature genes for ERG-negative group. OncoPrint was used to display the mutation landscape in ERG-negative group69.
Independent validation dataset
We used an independent whole genome sequencing data (CPDR dataset) to validate the CNV candidate genes. The CPDR dataset including 7 ERG-positive and 7 ERG-negative prostate tumors. The Genomatix software suite/NGS Analysis (http://www.genomatix.de) was used for CNV calling.
Differential expression analysis
We identified the differentially expressed genes among ERG-positive (n = 201), ERG-negative (n = 296) and normal samples (n = 52). Normalized read counts were used to detect differential expression genes with R package voom and limma70. Genes with P value < 0.05 and the absolute value of fold change (FC) > 2 were considered as differentially expressed.
Differential methylation analysis
We identified the differentially methylated genes among ERG-positive (n = 201), ERG-negative (n = 296) and normal samples (n = 35) based on TCGA methylation data. Firstly, we removed the probes on X/Y/M chromosome or NA. Secondly, we found diff-methylated sites with t-test p < 0.01 and the absolute difference of beta value > 0.2. Thirdly, we selected diff-methylated sites on promoter region (TSS200, TSS1500, 5’UTR and 1stExon). Fourthly, we retained methylation sites negatively correlated with the corresponding gene expression in a cis-regulatory manner. Fifthly, we concentrated on hyper-methylated sites whose corresponding genes have significantly lower expression in tumor samples compared to normal samples. For the comparison between ERG-positive and ERG-negative group, genes hyper-methylated in either group were taken into account.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files).
References
Torre, L. A. et al. Global cancer statistics, 2012. CA: a cancer journal for clinicians 65, 87–108 (2015).
Barbieri, C. E. et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nature genetics 44, 685–689 (2012).
Barbieri, C. E. et al. The mutational landscape of prostate cancer. European urology 64, 567–576 (2013).
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer cell 18, 11–22 (2010).
Network, C. G. A. R. The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Kan, Z. et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466, 869–873 (2010).
Berger, M. F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011).
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Tomlins, S. A. et al. TMPRSS2: ETV4 gene fusions define a third molecular subtype of prostate cancer. Cancer research 66, 3396–3400 (2006).
Helgeson, B. E. et al. Characterization of TMPRSS2: ETV5 and SLC45A3: ETV5 gene fusions in prostate cancer. Cancer research 68, 73–80 (2008).
Rubin, M. A., Maher, C. A. & Chinnaiyan, A. M. Common gene rearrangements in prostate cancer. Journal of Clinical Oncology 29, 3659–3668 (2011).
Robinson, D. et al. Integrative clinical genomics of advanced prostate cancer. Cell 161, 1215–1228 (2015).
Camacho, N. et al. Appraising the relevance of DNA copy number loss and gain in prostate cancer using whole genome DNA sequence data. PLoS genetics 13, e1007001, https://doi.org/10.1371/journal.pgen.1007001 (2017).
Attard, G. et al. Prostate cancer. The Lancet 387, 70–82, https://doi.org/10.1016/S0140-6736(14)61947-4 (2016).
Attard, G. et al. Prostate cancer. Lancet (London, England) 387, 70–82, https://doi.org/10.1016/s0140-6736(14)61947-4 (2016).
Hermans, K. G. et al. TMPRSS2: ERG fusion by translocation or interstitial deletion is highly relevant in androgen-dependent prostate cancer, but is bypassed in late-stage androgen receptor–negative prostate cancer. Cancer research 66, 10658–10663 (2006).
Lin, B. et al. Prostate-localized and androgen-regulated expression of the membrane-bound serine protease TMPRSS2. Cancer research 59, 4180–4184 (1999).
Tomlins, S. A. et al. Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia 10, 177IN171–188IN179 (2008).
Yu, J. et al. An integrated network of androgen receptor, polycomb, and TMPRSS2-ERG gene fusions in prostate cancer progression. Cancer cell 17, 443–454 (2010).
Brase, J. C. et al. TMPRSS2-ERG-specific transcriptional modulation is associated with prostate cancer biomarkers and TGF-β signaling. BMC cancer 11, 507 (2011).
Ratz, L. et al. TMPRSS2: ERG gene fusion variants induce TGF-β signaling and epithelial to mesenchymal transition in human prostate cancer cells. Oncotarget 8, 25115 (2017).
Sreenath, T. L. et al. ETS related gene mediated androgen receptor aggregation and endoplasmic reticulum stress in prostate cancer development. Scientific reports 7, 1109 (2017).
Perner, S. et al. TMPRSS2-ERG fusion prostate cancer: an early molecular event associated with invasion. The American journal of surgical pathology 31, 882–888 (2007).
Tandefelt, D. G., Boormans, J., Hermans, K. & Trapman, J. ETS fusion genes in prostate cancer. Endocrine-related cancer 21, R143–R152 (2014).
Furusato, B. et al. ERG oncoprotein expression in prostate cancer: clonal progression of ERG-positive tumor cells and potential for ERG-based stratification. Prostate cancer and prostatic diseases 13, 228 (2010).
Pettersson, A. et al. The TMPRSS2: ERG rearrangement, ERG expression, and prostate cancer outcomes: a cohort study and meta-analysis. Cancer Epidemiology and Prevention Biomarkers 21, 1497–1509 (2012).
Nam, R. K. et al. Expression of TMPRSS2: ERG gene fusion in prostate cancer cells is an important prognostic factor for cancer progression. Cancer biology & therapy 6, 40–45 (2007).
Demichelis, F. et al. TMPRSS2: ERG gene fusion associated with lethal prostate cancer in a watchful waiting cohort. Oncogene 26, 4596–4599 (2007).
Perner, S. et al. TMPRSS2: ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer research 66, 8337–8341 (2006).
Fine, S. W. et al. TMPRSS2–ERG gene fusion is associated with low Gleason scores and not with high-grade morphological features. Modern pathology 23, 1325–1333 (2010).
Gopalan, A. et al. TMPRSS2-ERG gene fusion is not associated with outcome in patients treated by prostatectomy. Cancer research 69, 1400–1406 (2009).
FitzGerald, L. M. et al. Association of TMPRSS2-ERG gene fusion with clinical characteristics and outcomes: results from a population-based study of prostate cancer. BMC cancer 8, 230 (2008).
Darnel, A. D., LaFargue, C. J., Vollmer, R. T., Corcos, J. & Bismar, T. A. TMPRSS2-ERG fusion is frequently observed in Gleason pattern 3 prostate cancer in a Canadian cohort. Cancer biology & therapy 8, 125–130 (2009).
Xu, B. et al. The prognostic role of ERG immunopositivity in prostatic acinar adenocarcinoma: a study including 454 cases and review of the literature. Human pathology 45, 488–497 (2014).
Petrovics, G. et al. Frequent overexpression of ETS-related gene-1 (ERG1) in prostate cancer transcriptome. Oncogene 24, 3847 (2005).
Hu, Y. et al. Delineation of TMPRSS2-ERG splice variants in prostate cancer. Clinical Cancer Research 14, 4719–4725 (2008).
Cullen, J. et al. Predicting Prostate Cancer Progression as a Function of ETS-related Gene Status, Race, and Obesity in a Longitudinal Patient Cohort. European urology focus (2017).
Moniri, M. R., Hsing, M., Rennie, P. S., Cherkasov, A. & Cox, M. E. The future of prostate cancer precision medicine: anti-ERG therapies. Translational Cancer Research 6, S1136–S1138 (2017).
Wang, X. et al. Development of Peptidomimetic Inhibitors of the ERG Gene Fusion Product in Prostate Cancer. Cancer cell 31, 532–548. e537, https://doi.org/10.1016/j.ccell.2017.05.001 (2017).
Mohamed, A. A. et al. Identification of a small molecule that selectively inhibits ERG-positive cancer cell growth. Cancer Res, https://doi.org/10.1158/0008-5472.can-17-2949 (2018).
Sedarsky, J., Degon, M., Srivastava, S. & Dobi, A. Ethnicity and ERG frequency in prostate cancer. Nature Reviews Urology. Preprint at https://doi.org/10.1038/nrurol.2017.140 (2017).
Magi‐Galluzzi, C. et al. TMPRSS2–ERG gene fusion prevalence and class are significantly different in prostate cancer of caucasian, african‐american and japanese patients. The Prostate 71, 489–497 (2011).
Dobi, A. et al. ERG-based stratification of prostate cancer highlights ethnicity associated biological differences. https://doi.org/10.1158/1538-7445.AM2015-5277 (AACR, 2015).
Ren, S. et al. Whole-genome and transcriptome sequencing of prostate cancer identify new genetic alterations driving disease progression. European urology. Preprint at https://doi.org/10.1016/j.eururo.2017.08.027 (2017).
Alumkal, J. J. & Herman, J. G. Distinct Epigenetic Mechanisms Distinguish TMPRSS2–ERG Fusion-Positive and-Negative Prostate Cancers. Cancer discovery 2, 979–981 (2012).
Hu, X. et al. TumorFusions: an integrative resource for cancer-associated transcript fusions. Nucleic Acids Research. Preprint at https://doi.org/10.1093/nar/gkx1018 (2017).
Han, B. et al. A fluorescence in situ hybridization screen for E26 transformation–specific aberrations: identification of DDX5-ETV4 fusion protein in prostate cancer. Cancer research 68, 7629–7637 (2008).
Esgueva, R. et al. Prevalence of TMPRSS2–ERG and SLC45A3–ERG gene fusions in a large prostatectomy cohort. Modern Pathology 23, 539–546 (2010).
Petrovics, G. et al. A novel genomic alteration of LSAMP associates with aggressive prostate cancer in African American men. EBioMedicine 2, 1957–1964 (2015).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Mermel, C. H. et al. GISTIC2. 0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome biology 12, R41, https://doi.org/10.1186/gb-2011-12-4-r41 (2011).
Lee, J. H., Song, S. Y., Kim, M. S., Yoo, N. J. & Lee, S. H. Frameshift mutations of a tumor suppressor gene ZNF292 in gastric and colorectal cancers with high microsatellite instability. Apmis 124, 556–560 (2016).
Fabbri, G. & Dalla-Favera, R. The molecular pathogenesis of chronic lymphocytic leukaemia. Nature Reviews Cancer 16, 145–162 (2016).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell systems 1, 417–425 (2015).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Myers, J. S., von Lersner, A. K., Robbins, C. J. & Sang, Q.-X. A. Differentially expressed genes and signature pathways of human prostate cancer. PloS one 10, e0145322, https://doi.org/10.1371/journal.pone.0145322 (2015).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Burkhardt, L. et al. CHD1 is a 5q21 tumor suppressor required for ERG rearrangement in prostate cancer. Cancer research 73, 2795–2805 (2013).
Hua, L. et al. FRK suppresses the proliferation of human glioma cells by inhibiting cyclin D1 nuclear accumulation. Journal of neuro-oncology 119, 49–58 (2014).
Wagner, A. H. et al. DGIdb 2.0: mining clinically relevant drug–gene interactions. Nucleic acids research 44, D1036–D1044 (2015).
Cowin, P. A. et al. LRP1B deletion in high-grade serous ovarian cancers is associated with acquired chemotherapy resistance to liposomal doxorubicin. Cancer research 72, 4060–4073 (2012).
Tornesello, M. L. et al. Mutations in TP53, CTNNB1 and PIK3CA genes in hepatocellular carcinoma associated with hepatitis B and hepatitis C virus infections. Genomics 102, 74–83 (2013).
Schulze, K. et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nature genetics 47, 505–511 (2015).
Totoki, Y. et al. Trans-ancestry mutational landscape of hepatocellular carcinoma genomes. Nature genetics 46, 1267–1273 (2014).
Grossman, R. L. et al. Toward a shared vision for cancer genomic data. New England Journal of Medicine 375, 1109–1112 (2016).
Reich, M. et al. GenePattern 2.0. Nature genetics 38, 500–501 (2006).
Zhao, M., Kim, P., Mitra, R., Zhao, J. & Zhao, Z. TSGene 2.0: an updated literature-based knowledgebase for tumor suppressor genes. Nucleic acids research 44, D1023–D1031 (2015).
Boutet, E. et al. UniProtKB/Swiss-Prot, the manually annotated section of the UniProt KnowledgeBase: how to use the entry view. Plant bioinformatics: methods and protocols, 23–54, https://doi.org/10.1007/978-1-4939-3167-5_2 (2016).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. Voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome biology 15, R29, https://doi.org/10.1186/gb-2014-15-2-r29 (2014).
Acknowledgements
This work was supported by the National Key Research and Development Program on Precision Medicine (2016YFC0902201, 2016YFC0902400), National Natural Science Foundation of China (31771472), National Grand Program on Key Infectious Diseases (2015ZX10004801-005), and Chinese Academy of Sciences (ZDBS-SSW-DQC-02).
Author information
Authors and Affiliations
Contributions
Q.X. and H.L. participated in the design of the study and drafted the manuscript; Q.X. and Y.S. collected the data and carried out data analysis; H.L. and Y.L. conceived and directed the study; Y.J., J.H., A.D., S.S., W.W. and S.S. participated in the design and coordination of the study; H.L., Y.L. and G.Z. supervised the project. All of the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xiao, Q., Sun, Y., Dobi, A. et al. Systematic analysis reveals molecular characteristics of ERG-negative prostate cancer. Sci Rep 8, 12868 (2018). https://doi.org/10.1038/s41598-018-30325-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30325-9
This article is cited by
-
Characterization and evaluation of gene fusions as a measure of genetic instability and disease prognosis in prostate cancer
BMC Cancer (2023)
-
CDC20 and PTTG1 are Important Biomarkers and Potential Therapeutic Targets for Metastatic Prostate Cancer
Advances in Therapy (2021)
-
Whole-genome sequencing of glioblastoma reveals enrichment of non-coding constraint mutations in known and novel genes
Genome Biology (2020)
-
A genomic and epigenomic atlas of prostate cancer in Asian populations
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.