Abstract
Pancreatic ductal adenocarcinoma (PDAC) is an aggressive disease that lacks effective biomarkers for early detection. We hypothesized that circulating long non-coding RNAs (lncRNAs) may act as diagnostic markers of incidentally-detected cystic PDAC precursors known as intraductal papillary mucinous neoplasms (IPMNs) and predictors of their pathology/histological classification. Using NanoString nCounter® technology, we measured the abundance of 28 candidate lncRNAs in pre-operative plasma from a cohort of pathologically-confirmed IPMN cases of various grades of severity and non-diseased controls. Results showed that two lncRNAs (GAS5 and SRA) aided in differentiating IPMNs from controls. An 8-lncRNA signature (including ADARB2-AS1, ANRIL, GLIS3-AS1, LINC00472, MEG3, PANDA, PVT1, and UCA1) had greater accuracy than standard clinical and radiologic features in distinguishing ‘aggressive/malignant’ IPMNs that warrant surgical removal from ‘indolent/benign’ IPMNs that can be observed. When the 8-lncRNA signature was combined with plasma miRNA data and quantitative ‘radiomic’ imaging features, the accuracy of predicting IPMN pathological classification improved. Our findings provide novel information on the ability to detect lncRNAs in plasma from patients with IPMNs and suggest that an lncRNA-based blood test may have utility as a diagnostic adjunct for identifying IPMNs and their pathology, especially when incorporated with biomarkers such as miRNAs, quantitative imaging features, and clinical data.
Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the third leading cause of cancer deaths in the United States, with a five-year survival rate of only 9%1. Most cases are diagnosed at a late, incurable stage due to the lack of accurate methods for early detection1. Serum carbohydrate antigen 19-9 (CA19-9) is used to suggest a diagnosis of PDAC and monitor disease recurrence or response to therapy. However, it’s utility as a sensitive and specific marker of early PDAC is poor2. The detection and treatment of noninvasive precursor lesions offers hope in reducing PDAC-related morbidity and mortality. Intraductal papillary mucinous neoplasms (IPMNs) of the pancreas are a morphologically distinct set of tumors located in the duct epithelium and are characterized by papillary epithelial proliferation and mucin production, leading to cystic dilation of involved ducts3. These cystic PDAC precursor lesions (‘precancers’) comprise almost half of the ~150,000 asymptomatic pancreatic cysts detected incidentally in the general population each year by computed tomography (CT) scans and magnetic resonance imaging (MRI)4, 5. Despite their radiologic detection, the only way to accurately examine their severity which ranges from noninvasive/pre-malignant (low-grade (LG), moderate-grade (MG), or high-grade (HG) dysplasia) to invasive carcinoma is through surgical resection, which carries significant risks of morbidity and mortality6,7,8. Studies reveal that the 5-year survival rate for patients with noninvasive IPMNs is greater than 70%, which is significantly higher than the 22–45% rate for individuals with resected invasive IPMNs and the 10–25% survival rate reported for individuals with resected conventional PDACs9. Thus, proper diagnosis of disease and its severity (noninvasive versus invasive IPMN versus conventional PDAC not associated with an IPMN) is important for medical management.
Consensus guidelines for IPMN management10 depend on standard radiographic and clinical features and recommend that patients with ‘high risk stigmata (HRS)’ (main pancreatic duct (MD) involvement/dilatation >10 mm, obstructive jaundice with a cystic lesion in the pancreatic head, or an enhanced solid component/nodule within the cyst) undergo resection, as most harbor high-grade or invasive disease10. More challenging to manage are IPMNs having ‘worrisome features (WF)’ (MD dilation 5–9 mm, cyst size >3 cm, thickened enhanced cyst walls, non-enhanced mural nodules, or acute pancreatitis). It is suggested that these lesions undergo endoscopic evaluation with endoscopic ultrasound with fine needle aspiration (EUS-FNA) despite suboptimal sensitivity and technical complications5, 11. The guidelines provide an important framework for management, but there is disagreement between the preoperative diagnosis and pathologic examination in a large percentage (30–70%) of cases10, 12,13,14,15,16. Novel, noninvasive markers of IPMN pathology are needed, especially for individuals who do not present with HRS.
Non-coding RNAs (ncRNAs) are outstanding candidate biomarkers of early neoplasia due to their stability in tissue17, 18 and biofluids19, 20 and their ability to regulate hundreds of genes and biological pathways. ncRNAs are typically classified according to their size; small ncRNAs are less than 200 nucleotides in length whereas long ncRNAs contain at least 200 nucleotides and resemble protein-coding transcripts but without functional open reading frames21. We22, 23 and others24,25,26 have conducted genome-wide analysis of miRNAs, the best characterized class of small ncRNAs, using tissue and plasma or serum from IPMN patients and healthy controls. Collectively, these data support miRNAs as a promising diagnostic adjunct for stratifying IPMN patients for surveillance or resection. Furthermore, in a small retrospective cohort of IPMN cases, we found diagnostic performance to improve when combining plasma miRNA data with quantitative ‘radiomic’ features invisible to the human eye extracted from routine CT scans27. LncRNAs are not as well studied as miRNAs even though they are the predominant transcribed RNAs28 and have been shown to regulate gene expression and promote carcinogenesis through mechanisms such as transcriptional regulation, initiation of chromatin remodeling, modulating alternative splicing, altering protein activity or localization, and genomic imprinting21, 29, 30. Given emerging data on the role of lncRNAs in PDAC initiation, progression, and outcomes31,32,33,34,35,36,37,38,39 and evidence to support lncRNA detection in circulation38, 40,41,42,43,44, we conducted the first study to measure the abundance of circulating lncRNAs from patients with IPMNs.
Here we quantified the abundance of 28 candidate lncRNAs in archived plasma obtained pre-operatively from individuals with noninvasive and invasive IPMNs and age- and gender-matched disease-free controls using the nCounter® technology (NanoString, Inc, Seattle, Washington) digital quantification method45. We then aimed to (1) discover circulating lncRNAs that may distinguish between patients with IPMNs and non-diseased controls, (2) identify circulating lncRNAs that may differentiate malignant/aggressive IPMNs (classified as those pathologically confirmed to have high-grade dysplasia or invasive disease) from benign/more indolent IPMNs (classified as those confirmed to have low- or moderate-grade dysplasia), and (3) determine the performance of circulating lncRNAs in predicting IPMN pathology individually and in combination with existing miRNA and quantitative imaging data described in our previous publications22, 27. Our findings provide new information on circulating lncRNAs and their potential to help individualize risk assessment and management for individuals with IPMNs.
Results
Study Population
Pre-operative plasma samples were evaluated for 57 IPMN cases and 24 non-diseased controls frequency-matched by age-group and gender. Eight samples were excluded prior to normalization and statistical analysis due to: failure to amplify (1 case) and high binding density due to possible over-seeding of the nCounter cartridge or erythrocyte contamination (5 cases, 2 controls), leaving samples from 73 participants (51 cases, 22 controls) for analysis. Study population characteristics are shown in Table 1. Cases and controls were well-matched on age (mean age: 68.5 vs 68.2 years). Most subjects were white, non-Hispanic. IPMN cases were more likely (47%) to have smoked cigarettes than controls (36%). The distribution of low-, moderate-, high-grade, and invasive IPMN cases was 12%, 29%, 24%, and 35%, respectively. Selected characteristics of the IPMN cohort are summarized in Table 2. Pre-operatively, compared to benign IPMNs (those with pathologically-confirmed low- or moderate grade dysplasia), malignant IPMNs (those with high-grade or invasive disease) were significantly (p < 0.05) more likely to be associated with: jaundice as a presenting symptom, tumors in the pancreatic head, MD involvement, and a solid component or mural nodule.
Analysis of plasma lncRNAs in IPMN cases versus non-diseased controls
Each of the 28 lncRNAs had signals that were detectable above background and could be included in analyses. No statistically significant differences were observed between IPMN and control samples in the frequency or the amount of erythrocyte contamination. After normalization, two lncRNAs (GAS5 and SRA) differentiated the 51 IPMN samples from the 22 control samples using a threshold of p < 0.05. Compared to controls, IPMN cases had 0.9-fold lower GAS5 expression and 1.2 fold higher SRA expression (Supplementary Figure S1). The 2-lncRNA signature, represented by PC1, explained 60% of the variability in the data, suggesting it represents the signature well. The area under the curve (AUC) value was 72.9 (95% CI: 60.9–84.9). The sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) for IPMN detection were 82%, 59%, 82% and 59%, respectively. Expression results for cases and controls are in Supplementary Table S1.
Analysis of plasma lncRNAs in malignant versus benign IPMNs
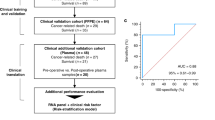
We evaluated the ability of the 2-lncRNA signature (identified in the analysis of IPMN cases versus non-diseased controls) to discriminate between the 30 malignant and 21 benign IPMNs and observed it did not perform well (AUC = 55.1 (95% CI: 38.1–72.1)). However, a focused analysis of the 51 IPMN cases showed that a signature of eight lncRNAs (ADARB2-AS1, ANRIL, GLIS3-AS1, LINC00472, MEG3, PANDA, PVT1, and UCA1) discriminated between the malignant and benign IPMNs (p < 0.05) (Table 3; Fig. 1). The 8-lncRNA signature was also characterized by a PC1 score and explained 67% of the variability. The overall expression of the 8-lncRNA signature was higher in malignant compared to benign IPMNs (p = 0.001, Fig. 2), was significantly associated with malignant status (OR (95% CI): = 1.56 (1.11–2.21), p = 0.011), and had an AUC value of 0.77 (95% CI: 0.62–0.92) in discriminating between groups (Fig. 2). Estimates of sensitivity, specificity, PPV, and NPV of the 8-lncRNA signature in discriminating malignant from benign IPMNs were 77%, 76%, 82%, and 70%, respectively. The 8-lncRNA signature also had a high specificity (91%) and PPV (90%) when comparing all IPMN cases to healthy controls. However, when comparing only malignant IPMN cases to healthy controls, neither individual lncRNAs or the 8-lncRNA signature clearly differentiated between these two groups (Fig 2 and 3), suggesting further interrogation will be necessary in larger datasets.
The 8-lncRNA signature associates with IPMN pathology. (A) Box plots of the distribution of the 8-lncRNA signature score (designated by the first principal component (PC1)) within the malignant and benign groups and healthy controls. (B) Receiver operating characteristic (ROC) curve analysis showed that the 8-lncRNA signature PC1 yielded an area under the curve (AUC) value of 0.77 (95% CI: 0.62–0.92) in differentiating between malignant and benign IPMN groups.
Relationship between standard clinical and radiologic data, IPMN pathology, and lncRNA expression
Clinical factors associated (p < 0.05) with an increased risk of malignancy included cyst size greater >3 cm (OR (95% CI) = 2.60 (0.76–8.86, p = 0.127), main duct involvement (OR (95% CI) = 11.00 (2.26–53.63, p = 0.003), and serum albumin levels (OR (95% CI) = 0.35 (0.11–1.15, p = 0.083). Cyst size positively correlated with the 8-lncRNA signature (Spearman correlation = 0.243, p = 0.125), while serum albumin levels negatively correlated with the 8-lncRNA signature (Spearman correlation = −0.158, p = 0.356). Mean serum CA 19-9 levels were higher in the malignant compared to benign group, but the difference was not statistically significant (p = 0.130). Of the identified lncRNAs, only PANDA was associated with serum CA 19-9 levels (p = 0.018), but the spearman correlation coefficient was not strong (r = 0.41). The 8-lncRNA signature was independently associated with malignant status after adjustment for MD involvement (p = 0.033) and presence of jaundice (p = 0.0095).
Combining lncRNA data with miRNA and radiomic data
Our group recently evaluated the possibility of using a multimodal approach to predict IPMN pathology that combines a 5-miRNA signature and a 14-feature radiomic CT signature generated using previously described procedures27 (see methods). We showed that a model which integrates miRNA and radiomic data had an area under the curve (AUC) value of 0.92 in discriminating 18 malignant from 20 benign cases; this was far superior to the AUC value for standard radiologic worrisome features (AUC = 0.54). In the current investigation, we integrated the 8-lncRNA signature with the 5-miRNA signature and the 14-feature radiomic CT signature and re-assessed diagnostic performance.
Of the 51 IPMN cases represented in the current investigation of lncRNAs, 31 (14 malignant, 17 benign) have available paired pre-operative plasma miRNA data and CT data. As summarized in Table 4, diagnostic performance increased from an AUC value of 0.76 when considering the 8-lncRNA signature alone to 0.90 when combining the 8-lncRNA signature, the 5-miRNA signature, and the 14-feature radiomic signature due to an increase in specificity. Finally, by incorporating standard radiologic features (worrisome features in the absence of high risk stigmata), gender, and presence of jaundice, the AUC increased to 0.92 (95% CI: 0.82–1.00) and revealed estimates of sensitivity, specificity, PPV and NPV of 93%, 82%, 81%, and 93%, respectively (Fig. 3 and Table 4). Importantly, the integrated discrimination improvement (IDI) index significantly increased with the addition of the 8-lncRNA signature, the 5-miRNA signature, and the radiomics signature, with respective IDI indices of 0.12, 0.14, and 0.13. Collectively, when the two molecular signatures and the radiomic signature were added to the model containing WF, gender, and jaundice, the IDI increased to 0.41 (95% CI: 0.23–0.59), p < 0.001. As anticipated, models that considered presence of high-risk stigmata individually or in conjunction with other data types performed well.
ROC analysis suggests that genomic data (the 8-lncRNA signature and a 5-miRNA signature) and quantitative radiomic features are more accurate in predicting malignant IPMN pathology than standard worrisome radiologic features and certain demographic and clinical variables. A final model combining the 8-lncRNA signature, the 5-miRNA signature, radiomic features, standard worrisome features (WF), gender, and presence of jaundice has potential to have high accuracy in predicting malignant pathology, with an AUC value approximating 0.92.
Cross-validation analysis
Evaluation of uncertainty by 10-fold cross-validation showed fairly robust estimates of diagnostic performance with AUC above 0.70 for most models that incorporated lncRNAs (Supplementary Table S2). A model that combined lncRNAs, miRNAs, radiomics, and worrisome features had an AUC = 0.77 (95% CI: 0.68–0.84) in cross-validation and was more accurate than standard demographic (gender), clinical (presence of jaundice), and radiologic variables (worrisome features) in predicting malignant pathology.
Discussion
This is the first investigation to report on lncRNA quantification using plasma from individuals newly-diagnosed with commonly detected PDAC precursors known as IPMNs. nCounter technology® was used to measure lncRNAs as an alternative to microarray and PCR-based methods to more accurately detect and quantify low lncRNA levels present in blood46. Similar to our previous study of circulating miRNAs22, an extensive quality control and data analysis pipeline was implemented to control for pre-analytical and technical factors that may affect circulating lncRNA levels and result in biases that do not reflect underlying biology. We show that lncRNAs can be detected in plasma, and provide data to support the possibility that this class of noninvasive biomarkers may serve as an adjunct to help predict IPMN severity/pathology.
To evaluate whether lncRNAs may serve as diagnostic markers of PDAC, recent studies37, 39, 47,48,49,50 have evaluated lncRNA expression in PDAC tumor tissue and adjacent normal tissue, and a few51, 52 have evaluated lncRNA expression in the circulation of PDAC patients, control patients, and healthy individuals using blood51, 53 or saliva52. Of the three biofluid-based studies51, 52, only one51 focused on detection of early-stage PDAC and none studied PDAC precursors. Hence, to our knowledge, we are unaware of previous published reports of lncRNA signatures derived from tissues, blood, or other biofluids from patients with IPMNs. Using plasma, we show that two lncRNAs (GAS5 and SRA) can partially discriminate IPMN cases from non-diseased controls (AUC) = 0.73 (95% CI:0.61–0.85, p = 0.004). GAS5 (growth arrest-specific 5) was found to have lower expression in IPMN cases compared to controls (p = 0.028), consistent with data33 showing that GAS5 expression is significantly decreased in pancreatic cancer tissues compared with normal pancreatic tissues. The authors33 demonstrate that GAS5 inhibition induces a significant decrease in G0/G1 phase and an increase in S phase, whereas overexpression in PDAC cells inhibits cell proliferation by negatively regulating CDK6 (cyclin-dependent kinase 6) expression in vitro and in vivo. Furthermore, a recent study showed that decreased GAS5 levels in serum were associated with type 2 diabetes in a cohort of US military veteran54, suggesting this lncRNA may help to identify individuals ‘at-risk’ for diabetes. GAS5 expression was not associated with presence of diabetes among IPMN cases in our small dataset (p = 0.36), but given that diabetes is an established risk factor for PDAC and ‘new-onset’ diabetes may serve as a potential marker of early PDAC55, further research may be indicated to explore a possible role for GAS5 in the molecular pathogenesis of diabetes-mediated PDAC. SRA (steroid receptor RNA activator) is responsible for coordinating functions of transcription factors and enhancing steroid receptor-dependent gene expression56. Specifically, as a nuclear receptor coactivator, SRA can coactivate androgen receptor (AR), estrogen receptor alpha (ERalpha), ERbeta, progesterone receptor (PR), glucocorticoid receptor (GR), thyroid hormone receptor and retinoic acid receptor (RAR). Emerging studies have revealed that SRA plays a key role in biological processes (such as myogenesis and steroidogenesis) and pathological changes (such as obesity and tumorigenesis)56, 57. Thus, it is biologically plausible SRA could be expressed at higher levels in IPMNs compared to normal controls. Moreover, in our dataset, patients with malignant IPMNs had higher levels of SRA than those with benign IPMNs. Further investigation is needed in larger cohorts before conclusions can be reached.
Analysis also revealed an 8-lncRNA signature (comprising ADARB2-AS1, ANRIL, GLIS3-AS1, LINC00472, MEG3, PANDA, PVT1, and UCA1) that partially discriminates between malignant and benign IPMNs AUC = 0.77 (95% CI: 0.62–0.92, p = 0.006). These lncRNAs were upregulated in malignant compared to benign cases. Consistent with our findings, several of the identified lncRNAs (UCA1, PVT1) appear to have oncogenic functions in pancreatic carcinogenesis47, 48, 52, 53, inferring biological plausibility of the 8-lncRNA signature. Urothelial cancer-associated 1 (UCA1) is known for its role in bladder cancer progression and embryologic development, and in two separate investigations47, 48 has recently been shown to be upregulated in PDAC tissues (versus matched adjacent normal pancreas tissue), to be associated with several prognostic factors (such as stage, tumor size, and grade), and shorter survival. Furthermore, functional experiments have shown that UCA1 promotes invasion and proliferation of PDAC cells and that down-regulation of UCA1 inhibits cell proliferation and induces apoptosis and cell cycle arrest47, 48. PVT1 (plasmacytoma variant translocation 1) has been shown to be upregulated in PDAC tissues, to be correlated with clinical stage and poor survival, to be overexpressed in the saliva of PDAC cases versus healthy controls, and to contribute to susceptibility to PDAC as part of a genome-wide association study53, 54, 58. Furthermore, PVT1 has been identified as a regulator of gemcitabine sensitivity; PVT1 inactivation led to enhanced gemcitabine sensitivity in human PDAC cells59. Thus, in addition to serving as prognostic markers and therapeutic targets, it is biologically plausible that UCA1 and PVT1 may also contribute to early pancreatic carcinogenesis (and malignant potential) as observed in this study. ANRIL (antisense non-coding RNA in the INK4 locus) was also shown to have higher expression in malignant versus benign IPMNs. Although we are unaware of reports of ANRIL dysregulation and PDAC, increased expression of this well-known lncRNA has been shown to contribute to the risk of diabetes60, a risk factor for PDAC55 and IPMNs61.
Several of the lncRNAs highlighted in our investigation appear to have a tumor suppressor role in pancreatic neoplasms. For example, maternally expressed gene 3 (MEG3) was recently shown to inhibit PDAC proliferation via activation of p53 and to play a key role in the anti-tumor effects of fenofibrate, a PPAR-α agonist62. Additionally, epigenetic activation of MEG3 (and inactivation of its target c-MET) with DNA methylating drugs has been shown to have a therapeutic effect on pancreatic neuroendocrine tumors63. Analyses in breast64 and ovarian tumors65 support a tumor suppressor role for intergenic lncRNA, LINC00472, but functional investigations of this gene are lacking. A recent phenome-genome association study of pancreatic cancer66 revealed several open reading frames associated with pancreatic neoplasms, and included candidates LINC00472, GLI3-AS1, and ADARB2-AS1. Finally, although reports of the candidate lncRNA PANDA (P21associated noncoding RNA DNA damage-activated) and PDAC are lacking, data suggest it plays a role in stabilizing p53 in response to DNA damage67 and in regulating senescence68.
To increase diagnostic performance, we combined the 8-lncRNA signature with a 5-miRNA plasma signature, ‘radiomic’ imaging features, and clinical characteristics available through previous studies22, 27. We showed that integration of multiple data types improves prediction of IPMN pathology beyond that provided by standard clinical and radiologic characteristics, especially worrisome features considered in consensus guidelines10. Although model overfitting may have contributed to our findings and warrants further interrogation in larger datasets, results of the integrated discrimination improvement (IDI) test and the cross-validation analysis suggest that these novel data types do have potential to add value in IPMN risk assessment.
Despite the strengths and novelty of this study, there are limitations that merit consideration. Given that lncRNAs are typically expressed at low concentrations in circulation, we used the robust nCounter platform (as opposed to quantitative real time RT-PCR as others have done) and performed a pre-amplification procedure at the suggestion of the manufacturer and available literature. Although pre-amplification has potential to introduce amplification bias or assay cartridge saturation, head-to-head comparisons of plasma samples with and without pre-amplification have shown a higher level of linearity in samples with pre-amplification40. Furthermore, circulating lncRNAs have been shown to be stable under different experimental conditions40. Although mechanisms accounting for lncRNA stability are incompletely understood, data suggest they may be protected by exosome encapsulation and/or complex formation with proteins and miRNAs38, 69. Thus, integrity of our findings may not have been significantly influenced by pre-amplification or sample instability. Our study is also limited in that we evaluated only 28 candidate lncRNAs. It is possible that lncRNAs not evaluated here (such as Linc-pint, a lncRNA observed in plasma to be a possible biomarker of early PDAC by Li et al.51) could be important to IPMN pathogenesis.
Other limitations of the current study are that we were unable to examine post-operative levels of plasma lncRNAs to see if they normalize following the removal of the lesion and we were unable to evaluate whether IPMN tissue is the origin of the circulating lncRNAs. Plasma is now being collected and processed pre- and post-surgery and during surveillance as part of our multi-institutional cohort study known as the Florida Pancreas Collaborative70 so that we can assess changes in biomarker levels over time. Due to the small or focal nature of IPMNs and the lack of ample tissue for molecular analyses for most cases, tissue microarrays (rather than whole sections) are being created to perform in situ hybridization for the most promising lncRNAs. Finally, the relatively small sample size of our study population limits the ability to draw meaningful conclusions. External validation in a large, multi-center prospective investigation of serial plasma lncRNA measurements is indicated for individuals newly-diagnosed with various types of pancreatic cysts and early-stage PDAC, and those at high genetic risk for developing PDAC, and healthy individuals without cancer.
In summary, lncRNAs can be detected in plasma and have potential to be incorporated clinically as part of a blood-based diagnostic adjunct to aid in IPMN management, especially in conjunction with other types of biomarkers (such as miRNAs) and quantitative radiologic features. Large-scale studies with rigorous designs and incorporation of epidemiologic and clinical data are needed to further explore the potential for circulating lncRNAs to be utilized as novel biomarkers for IPMN diagnosis and monitoring and as targets for intervention using RNA interference (RNA-i)-mediated approaches.
Materials and Methods
Study population and biospecimens
A prospectively maintained clinical database was retrospectively reviewed to identify individuals who underwent a pancreatic resection for an IPMN between 2006 and 2011 at Moffitt Cancer Center and Research Institute (Moffitt) and had provided written consent for blood to be donated pre-operatively for research through protocols approved by the Institutional Review Board (IRB) of the University of South Florida, including Total Cancer Care71. IRB approval was granted for study participation and the research described herein (IRB#Pro4971), written informed consent was obtained from study participants, and all methods were performed in accordance with relevant guidelines and regulations. The diagnosis and degree of dysplasia was pathologically confirmed using World Health Organization (WHO) guidelines3. The final diagnosis represented the most severe grade of dysplasia observed in the neoplastic epithelium. None of the included cases received pre-operative chemotherapy or radiation. Also eligible for inclusion were age- and gender- matched healthy controls with no current or prior history of pancreatic disease or symptoms who presented to Moffitt’s Cancer Screening and Prevention Center during the same time period and donated blood through a related IRB-approved protocol using the same procedures.
Blood was collected from consented participants via phlebotomy in a 7-mL EDTA tube and processed for plasma within two hours using standard procedures72. The tube was inverted 3 times and spun at 3600 rpm for 8 minutes and then aliquoted into 0.5 mL bar-coded cryovials and banked at −80 °C. Demographic, clinical, and epidemiologic data was collected from an electronic questionnaire, the medical record, Moffitt’s cancer registry, and other sources.
RNA isolation and multiplexed target enrichment (MTE)
One 0.5 mL cryovial of plasma was retrieved and thawed for each study participant. To assess hemolysis, samples were visually inspected and spectrophotometric analysis was performed at 414, 541, and 576 nm73. Samples were classified as hemolyzed if the A414, A541, or A576 value exceeded 0.2. Synthetic spike-in RNA representing NEFL (neurofilament 1), ENO2 (neuron-specific endonuclease) and GFAP (glial fibrillary acidic protein) were added to plasma to control for variance in the starting material and the efficiency of RNA extraction, according to vendor recommendations. Cell-free RNA was then isolated from 500 µl of plasma using the Plasma/Serum RNA Purification Midi Kit with a final elution volume of 50 µl (Norgen Biotek Corp, Ontario, CA). The extracted RNA was further purified and concentrated down to 30 µl using the Zymo RNA Clean and Concentrator-5 kit (Zymo Research Corp, Irvine, CA). The concentrated RNA was qualitatively assessed on the Agilent BioAnalyzer Total RNA Pico chip (Agilent Technologies, Santa Clara, CA). As the RNA was expected to be degraded, a BioAnalyzer peak from 25–200 nt indicated successful RNA recovery. Four µl of RNA was reverse transcribed into single-stranded cDNA using the Invitrogen SuperScript VILO cDNA Synthesis Kit and Master Mix (Thermo Fisher Scientific, Waltham, MA) following the nCounter Single Cell Gene Expression protocol (NanoString Technologies, Seattle, WA). The single-stranded cDNA was enriched using a highly multiplexed pool of target-specific PCR primer pairs targeting 28 lncRNAs. The MTE process was carried out using 22 cycles of PCR using the ABI TaqMan PreAmp MasterMix (Thermo Fisher Scientific, Waltham, MA) with the conditions described in the NanoString protocol.
Quality control of the multiplexed target enrichment and hybridization
The resulting PCR products from the Multiplexed Target Enrichment were assessed using quantitative RT-PCR (qPCR) to evaluate the expression of several target genes and normalize the amount of MTE DNA input into the NanoString hybridization. The MTE PCR products were diluted 1:100 and assayed on an Illumina Eco using ABI SYBR Green PCR Master Mix for genes, GNAS and VIM, which were confirmed to be expressed stably based on initial data generated from the custom NanoString CodeSet (data not shown). The resulting Cq values for both genes were used to generate a dilution factor for each sample. Dilution factors were calculated by targeting each sample’s effective Cq value (post-dilution) to equal 16 for GNAS and 19 for VIM, which were the approximate values generated by the successful pilot samples (those with binding densities between 0.05 and 2.25 spots per square micron). Samples with Cq values greater than the target values were not diluted, and in samples where calculations resulted in significantly different dilution factors, the higher dilution factor was selected in order to avoid cartridge overloading. Following the dilution of the MTE PCR product, the DNA was denatured for 2 minutes at 94 °C then snap cooled on ice for 5 minutes prior to hybridization. The samples were hybridized overnight at 65 °C with the custom codeset described below.
High-throughput measurement of lncRNA abundance
A custom nCounter™ Expression Assay codeset (Nanostring Technologies, Seattle, WA, USA) was used to quantify the abundance of 28 lncRNAs selected because of their role in the development or progression of pancreatic and other cancers after review of published literature31,32,33,34,35,36,37 ADARB2-AS1, ANRIL, AS1DHRS4, BCYRN1, DDX6P, GAS5, GLIS3-AS1, H19, HOTAIR, HOTTIP, HOXD-AS1, HULC, LINC00244, LINC00469, LINC00472, LINC00491, lncRNA-p21, MALAT1, MEG3, PANDA, PPP3CB, PTENP1, PVT1, SRA, TERC, UCA1, XIST, αHIF. The codeset also included positive controls, negative controls, three mRNA spike-in targets, four WBC cellular contamination targets (APOE, CD68, CD2, and CD3) three hemolysis/erythrocyte targets (MB, NGB, CYGB), and messenger RNA (mRNA) housekeeping genes (ACTB, PGK1, and PPIB). The NanoString cartridge was processed on the NanoString nCounter Prep Station using the high sensitivity protocol and scanned at the highest sensitivity setting (550 fields of view) on the nCounter Digital Analyzer. Data quality control was performed using the NanoString nSolver software, and were exported for normalization and analysis.
Data processing and quality control
For each sample, background-corrected measures of lncRNA expression were estimated by subtracting the negative control average plus two standard deviation (SD) cut-point from the raw lncRNA counts. lncRNAs with less than 20% of samples above the negative control cut-point (ie. low-expression probes) were removed from downstream analysis. Possible WBC and erythrocyte sample contamination was evaluated using built-in controls. Based on manufacturer recommendations, data for each sample was normalized using the geometric mean of the three housekeeping genes (ACTB, PGK1 and PPIB), each of which appeared to be relatively invariant/stable in the dataset and to have less variability as compared to the mRNA spike-in genes. Normalized data was log2-transformed prior to signature selection.
Statistical Analysis
Descriptive statistics were calculated using frequencies and percents for categorical variables and means and standard deviations (SD) for continuous variables. Data analysis was performed to identify a panel of lncRNAs that (a) differentiate between IPMN cases and non-diseased controls and (b) distinguish malignant (pathologically-confirmed as HG or invasive) from benign IPMNs (pathologically-confirmed as LG or MG).
Identification of plasma lnRNA signatures
Linear models for microarray data (LIMMA)74 was used to identify lncRNAs that differentiate between IPMN cases and controls and between malignant and benign IPMNs, respectively. Since lncRNAs can be over- or under-expressed, we used principal component analysis (PCA) to combine the most deregulated lncRNAs and generate an overall ‘IPMN-risk score’ based on the first principal component (PC1), which accounts for the largest variability in the data and represents the overall combined effect of an IPMN-risk lncRNA signature. Specifically, IPMN-risk score, defined by PC1 as \(\sum {w}_{i}{x}_{i}\), is a weighted average expression among the IPMN-risk lncRNAs, where x i represents lncRNA i expression level, w i is the corresponding weight (PC1’s loading coefficient for lncRNA i) with \(\sum {w}_{i}^{2}=1\), and the w i values maximize the variance of \(\sum {w}_{i}{x}_{i}\). This approach has been used to derive gene signatures previously22, 75,76,77,78.
Receiver operating characteristic (ROC) curves were generated to measure the predictive power of the IPMN-risk signatures in discriminating between groups. Youdon method was used to determine the best threshold value for each model based on maximum of sum of sensitivities and specificities as the optimality criterion79. Estimates of sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were calculated. For the analysis of malignant versus benign cases, we also conducted multivariable logistic regression analysis to assess whether the identified lncRNA signature was associated with malignant IPMN status independent of known prognostics factors (ie. main-duct involvement, lesion size, serum CA-19-9 level)10. Finally, to assess the extent to which data types in addition to plasma lncRNAs and standard clinical and radiologic features may augment correct prediction of malignant versus benign pathology, logistic regression models and ROC curves were also generated for a subset of cases with existing pre-operative plasma miRNA and radiomic data produced according to methods described in previous studies22, 27 and reviewed below. P values were adjusted with a false discovery rate (FDR) approach using the Benjamini and Hochberg method80. Additionally, to determine the ‘added value’ of the lncRNA, miRNA, and radiomic data, we calculated the integrated discrimination improvement (IDI) index, a magnitude of the reclassification probability improvement or worsening by a new test(s) over probability thresholds81.
miRNA expression data
Preoperative plasma miRNA expression data was generated previously22 for 42 surgically-resected, pathologically-confirmed IPMN cases (21 malignant and 21 benign) using one 0.5-mL cryovial of plasma per case. Briefly, RNA spike-in miRNAs (synthetic control templates) were used and total RNA isolation was performed on 500 uL of plasma using the Plasma/Serum Circulating and Exosomal RNA Purification Mini Kit (Slurry Format) from Norgen Biotek (Ontario, Canada). The nCounter™ Human v2 miRNA Expression Assay Codeset (NanoString Technologies, Seattle, WA, USA) was used to quantify the abundance of a pre-defined panel of 800 human miRNAs and built-in controls, and raw miRNA counts underwent technical and biological normalization and log2-transformation. The most deregulated miRNAs were identified using the linear models for microarray data (LIMMA) method and a principal component analysis (PCA) approach (14). A focused analysis of the 42 IPMN cases showed that five miRNAs (miR-200a-3p, miR-1185-5p, miR-33a-5p, miR-574-3p, and miR-663b) had an AUC value of 0.73 (95% CI: 0.58–0.89) in discriminating between groups.
CT acquisition and radiomic feature selection and extraction
CT and MRI-MRCP scans can be used to diagnose IPMNs, though MRI-MRCP is believed to be superior for optimal management82. In our series of surgically-resected, pathologically-confirmed IPMN cases, more cases had available pre-operative CT images versus pre-operative MRI-MRCP images. Specifically, preoperative CT images were available for 38 of the 42 pathologically-confirmed IPMN cases who also had available matched preoperative miRNA expression data generated previously22. CT images were obtained from Moffitt’s GE Centricity Picture Archiving and Communication System (PACS). CT images were reviewed for standard radiologic features encompassing ‘high-risk stigmata’ and ‘worrisome features’ represented in consensus guidelines10. Axial venous phase images (3 mm) were used for most patients the region of interest (ROI) by helping to outline the peripheral margin of tumors in their entirety, capturing both solid (nodular) and cystic components. The radiomics team then marked the ROI using Definiens/GE AWS Advanced Visualization software. The entire tumors were identified (solid and cystic components) using a semi-manual version of a single click semi-automated ensemble segmentation algorithm within the Definiens Developer XD (Munich, Germany) software platform. Target lesions were segmented, with a second radiologist finalizing the segmentation boundaries on the CT slices. We then extracted categories of 18 non-texture and 94 texture features. Non-texture features measure tumor size (volume, diameter, border length), shape (compactness, asymmetry), and location, whereas texture features measure properties such as smoothness, coarseness, and regularity. We focused on evaluating two-dimensional (2D) quantitative features in the middle CT slice. In house algorithms for feature extraction and quantification of segmented regions were implemented by custom routines in the Definiens Platform. Logistic regression, PCA, and cross-validation analyses were used to examine associations between features and IPMN pathology. Fourteen features, most of which were textural, differentiated malignant from benign IPMNs and collectively had an AUC value of 0.77.
Cross-validation analysis
To evaluate model performance, repeated (10,000 times) 10-fold cross validation was performed. The average and 95% confidence intervals of accuracy, sensitivity, specificity, PPV and NPV were estimated. In each 10-fold cross-validation, data were split into 10 subsets. By holding one subset of data (test set), the remaining 9 subsets were used as a training set to build a model for prediction evaluation in the test set. The process continued until each subset was used as the test set. By testing the model on a test set (not used in estimation), cross-validation aimed to reduce over-fitting. All statistical analyses were performed using SAS version 9.4 and R version 3.2.5.
References
American Cancer Society. Cancer Facts and Figures 2017. Atlanta: American Cancer Society (2017).
Szajda, S. D., Waszkiewicz, N., Chojnowska, S. & Zwierz, K. Carbohydrate markers of pancreatic cancer. Biochem Soc Trans 39, 340–343, doi:10.1042/bst0390340 (2011).
Adsay, N. V. et al. Intraductal Papillary Mucinous Neoplasm of the Pancreas. In: Bosman FT, Carneiro F, Hruban RH, Theise ND, editors. WHO classification of tumors of the digestive system. Lyon: WHO Press; p.304–313 (2010).
Megibow, A. J., Baker, M. E., Gore, R. M. & Taylor, A. The incidental pancreatic cyst. Radiol Clin North Am 49, 349–359, doi:10.1016/j.rcl.2010.10.008 (2011).
Farrell, J. J. Prevalence, Diagnosis and Management of Pancreatic Cystic Neoplasms: Current Status and Future Directions. Gut and liver 9, 571–589, doi:10.5009/gnl15063 (2015).
Sachs, T., Pratt, W. B., Callery, M. P. & Vollmer, C. M. Jr. The incidental asymptomatic pancreatic lesion: nuisance or threat? J Gastrointest Surg 13, 405–415, doi:10.1007/s11605-008-0788-0 (2009).
Matthaei, H., Schulick, R. D., Hruban, R. H. & Maitra, A. Cystic precursors to invasive pancreatic cancer. Nat Rev Gastroenterol Hepatol 8, 141–150, doi:10.1038/nrgastro.2011.2 (2011).
Hines, O. J. & Reber, H. A. Pancreatic surgery. Curr Opin Gastroenterol 24, 603–611, doi:10.1097/MOG.0b013e32830b112e (2008).
Koh, Y. X. et al. Systematic review and meta-analysis of the spectrum and outcomes of different histologic subtypes of noninvasive and invasive intraductal papillary mucinous neoplasms. Surgery 157, 496–509, doi:10.1016/j.surg.2014.08.098 (2015).
Tanaka, M. et al. International consensus guidelines 2012 for the management of IPMN and MCN of the pancreas. Pancreatology 12, 183–197, doi:10.1016/j.pan.2012.04.004 (2012).
Panarelli, N. C. et al. Commercial molecular panels are of limited utility in the classification of pancreatic cystic lesions. Am J Surg Pathol 36, 1434–1443, doi:10.1097/PAS.0b013e31825d534a (2012).
Kim, K. W. et al. Imaging features to distinguish malignant and benign branch-duct type intraductal papillary mucinous neoplasms of the pancreas: a meta-analysis. Ann Surg 259, 72–81, doi:10.1097/SLA.0b013e31829385f7 (2014).
Roch, A. M. et al. International Consensus Guidelines parameters for the prediction of malignancy in intraductal papillary mucinous neoplasm are not properly weighted and are not cumulative. HPB: the official journal of the International Hepato Pancreato Biliary Association 16, 929–935, doi:10.1111/hpb.12305 (2014).
Sahora, K. et al. Branch duct intraductal papillary mucinous neoplasms: does cyst size change the tip of the scale? A critical analysis of the revised international consensus guidelines in a large single-institutional series. Ann Surg 258, 466–475, doi:10.1097/SLA.0b013e3182a18f48 (2013).
Fritz, S. et al. Pancreatic main-duct involvement in branch-duct IPMNs: an underestimated risk. Ann Surg 260, 848–855; discussion 855–846, doi:10.1097/sla.0000000000000980 (2014).
Goh, B. K. et al. Utility of the sendai consensus guidelines for branch-duct intraductal papillary mucinous neoplasms: a systematic review. J Gastrointest Surg 18, 1350–1357, doi:10.1007/s11605-014-2510-8 (2014).
Xi, Y. et al. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. Rna 13, 1668–1674, doi:10.1261/rna.642907 (2007).
Li, J. et al. Comparison of miRNA expression patterns using total RNA extracted from matched samples of formalin-fixed paraffin-embedded (FFPE) cells and snap frozen cells. BMC Biotechnol 7, 36 (2007).
Mostert, B., Sieuwerts, A. M., Martens, J. W. & Sleijfer, S. Diagnostic applications of cell-free and circulating tumor cell-associated miRNAs in cancer patients. Expert Rev Mol Diagn 11, 259–275, doi:10.1586/erm.11.11 (2011).
Cortez, M. A. et al. MicroRNAs in body fluids–the mix of hormones and biomarkers. Nat Rev Clin Oncol 8, 467–477, doi:10.1038/nrclinonc.2011.76 (2011).
Shi, X., Sun, M., Liu, H., Yao, Y. & Song, Y. Long non-coding RNAs: A new frontier in the study of human diseases. Cancer Lett, doi:10.1016/j.canlet.2013.06.013 (2013).
Permuth-Wey, J. et al. Plasma MicroRNAs as Novel Biomarkers for Patients with Intraductal Papillary Mucinous Neoplasms of the Pancreas. Cancer Prev Res (Phila). doi:10.1158/1940-6207.capr-15-0094 (2015).
Permuth-Wey, J. et al. A Genome-Wide Investigation of MicroRNA Expression Identifies Biologically-Meaningful MicroRNAs That Distinguish between High-Risk and Low-Risk Intraductal Papillary Mucinous Neoplasms of the Pancreas. PLoS One 10, e0116869, doi:10.1371/journal.pone.0116869 (2015).
Li, A. et al. MicroRNA Array Analysis Finds Elevated Serum miR-1290 Accurately Distinguishes Patients with Low-Stage Pancreatic Cancer from Healthy and Disease Controls. Clin Cancer Res, doi:10.1158/1078-0432.ccr-12-3092 (2013).
Lubezky, N. et al. MicroRNA expression signatures in intraductal papillary mucinous neoplasm of the pancreas. Surgery 153, 663–672, doi:10.1016/j.surg.2012.11.016 (2013).
Matthaei, H. et al. miRNA biomarkers in cyst fluid augment the diagnosis and management of pancreatic cysts. Clin Cancer Res 18, 4713–4724, doi:10.1158/1078-0432.ccr-12-0035 (2012).
Permuth, J. B. et al. Combining radiomic features with a miRNA classifier may improve prediction of malignant pathology for pancreatic intraductal papillary mucinous neoplasms. Oncotarget, doi:10.18632/oncotarget.11768 (2016).
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science (New York, N.Y.) 316, 1484–1488, doi:10.1126/science.1138341 (2007).
Gibb, E. A., Brown, C. J. & Lam, W. L. The functional role of long non-coding RNA in human carcinomas. Molecular cancer 10, 38, doi:10.1186/1476-4598-10-38 (2011).
Guil, S. & Esteller, M. Cis-acting noncoding RNAs: friends and foes. Nat Struct Mol Biol 19, 1068–1075, doi:10.1038/nsmb.2428 (2012).
Li, J. et al. Long non-coding RNAs expressed in pancreatic ductal adenocarcinoma and lncRNA BC008363 an independent prognostic factor in PDAC. Pancreatology 14, 385–390, doi:10.1016/j.pan.2014.07.013 (2014).
Liu, J. H., Chen, G., Dang, Y. W., Li, C. J. & Luo, D. Z. Expression and prognostic significance of lncRNA MALAT1 in pancreatic cancer tissues. Asian Pacific journal of cancer prevention: APJCP 15, 2971–2977 (2014).
Lu, X. et al. Downregulation of gas5 increases pancreatic cancer cell proliferation by regulating CDK6. Cell and tissue research 354, 891–896, doi:10.1007/s00441-013-1711-x (2013).
Pang, E. J., Yang, R., Fu, X. B. & Liu, Y. F. Overexpression of long non-coding RNA MALAT1 is correlated with clinical progression and unfavorable prognosis in pancreatic cancer. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine, doi:10.1007/s13277-014-2850-8 (2014).
Peng, W., Gao, W. & Feng, J. Long noncoding RNA HULC is a novel biomarker of poor prognosis in patients with pancreatic cancer. Medical oncology (Northwood, London, England) 31, 346, doi:10.1007/s12032-014-0346-4 (2014).
Sun, Y. W. et al. A novel long non-coding RNA ENST00000480739 suppresses tumour cell invasion by regulating OS-9 and HIF-1alpha in pancreatic ductal adenocarcinoma. Br J Cancer 111, 2131–2141, doi:10.1038/bjc.2014.520 (2014).
Tahira, A. C. et al. Long noncoding intronic RNAs are differentially expressed in primary and metastatic pancreatic cancer. Molecular cancer 10, 141, doi:10.1186/1476-4598-10-141 (2011).
Kishikawa, T. et al. Circulating RNAs as new biomarkers for detecting pancreatic cancer. World journal of gastroenterology: WJG 21, 8527–8540, doi:10.3748/wjg.v21.i28.8527 (2015).
Zhou, M. et al. Construction and analysis of dysregulated lncRNA-associated ceRNA network identified novel lncRNA biomarkers for early diagnosis of human pancreatic cancer. Oncotarget 7, 56383–56394, doi:10.18632/oncotarget.10891 (2016).
Arita, T. et al. Circulating long non-coding RNAs in plasma of patients with gastric cancer. Anticancer research 33, 3185–3193 (2013).
Isin, M. et al. Investigation of circulating lncRNAs in B-cell neoplasms. Clinica chimica acta; international journal of clinical chemistry 431, 255–259, doi:10.1016/j.cca.2014.02.010 (2014).
Ren, S. et al. Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 derived miniRNA as a novel plasma-based biomarker for diagnosing prostate cancer. Eur J Cancer 49, 2949–2959, doi:10.1016/j.ejca.2013.04.026 (2013).
Wu, Y. et al. A serum-circulating long noncoding RNA signature can discriminate between patients with clear cell renal cell carcinoma and healthy controls. Oncogenesis 5, e192, doi:10.1038/oncsis.2015.48 (2016).
Xie, H., Ma, H. & Zhou, D. Plasma HULC as a Promising Novel Biomarker for the Detection of Hepatocellular Carcinoma. BioMed research international 2013, 136106, doi:10.1155/2013/136106 (2013).
Veldman-Jones, M. H. et al. Evaluating Robustness and Sensitivity of the NanoString Technologies nCounter Platform to Enable Multiplexed Gene Expression Analysis of Clinical Samples. Cancer Res 75, 2587–2593, doi:10.1158/0008-5472.can-15-0262 (2015).
Schultz, N. A. et al. MicroRNA biomarkers in whole blood for detection of pancreatic cancer. JAMA 311, 392–404, doi:10.1001/jama.2013.284664 (2014).
Fu, X. L. et al. Analysis of long non-coding RNA expression profiles in pancreatic ductal adenocarcinoma. Sci Rep 6, 33535, doi:10.1038/srep33535 (2016).
Chen, P. et al. Long non-coding RNA UCA1 promotes the tumorigenesis in pancreatic cancer. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie 83, 1220–1226, doi:10.1016/j.biopha.2016.08.041 (2016).
Wang, Y. et al. Expression profile of long non-coding RNAs in pancreatic cancer and their clinical significance as biomarkers. Oncotarget 6, 35684–35698, doi:10.18632/oncotarget.5533 (2015).
Zhou, Y. et al. Microarray expression profile analysis of long non-coding RNAs in pancreatic ductal adenocarcinoma. International journal of oncology 48, 670–680, doi:10.3892/ijo.2015.3292 (2016).
Li, L. et al. Plasma and tumor levels of Linc-pint are diagnostic and prognostic biomarkers for pancreatic cancer. Oncotarget, doi:10.18632/oncotarget.12365 (2016).
Xie, Z. et al. Salivary HOTAIR and PVT1 as novel biomarkers for early pancreatic cancer. Oncotarget 7, 25408–25419, doi:10.18632/oncotarget.8323 (2016).
Huang, C. S. et al. Increased expression of the lncRNA PVT1 is associated with poor prognosis in pancreatic cancer patients. Minerva medica (2015).
Carter, G. et al. Circulating long noncoding RNA GAS5 levels are correlated to prevalence of type 2 diabetes mellitus. BBA clinical 4, 102–107, doi:10.1016/j.bbacli.2015.09.001 (2015).
Yeo, T. P. Demographics, epidemiology, and inheritance of pancreatic ductal adenocarcinoma. Seminars in oncology 42, 8–18, doi:10.1053/j.seminoncol.2014.12.002 (2015).
Liu, C. et al. Steroid receptor RNA activator: Biologic function and role in disease. Clinica chimica acta; international journal of clinical chemistry 459, 137–146, doi:10.1016/j.cca.2016.06.004 (2016).
Xu, B. et al. Multiple roles for the non-coding RNA SRA in regulation of adipogenesis and insulin sensitivity. PLoS One 5, e14199, doi:10.1371/journal.pone.0014199 (2010).
Wolpin, B. M. et al. Genome-wide association study identifies multiple susceptibility loci for pancreatic cancer. Nat Genet 46, 994–1000, doi:10.1038/ng.3052 (2014).
You, L., Chang, D., Du, H. Z. & Zhao, Y. P. Genome-wide screen identifies PVT1 as a regulator of Gemcitabine sensitivity in human pancreatic cancer cells. Biochemical and biophysical research communications 407, 1–6, doi:10.1016/j.bbrc.2011.02.027 (2011).
Kong, Y., Sharma, R. B., Nwosu, B. U. & Alonso, L. C. Islet biology, the CDKN2A/B locus and type 2 diabetes risk. Diabetologia 59, 1579–1593, doi:10.1007/s00125-016-3967-7 (2016).
Capurso, G. et al. Risk factors for intraductal papillary mucinous neoplasm (IPMN) of the pancreas: a multicentre case-control study. The American journal of gastroenterology 108, 1003–1009, doi:10.1038/ajg.2013.42 (2013).
Hu, D. et al. Fenofibrate inhibited pancreatic cancer cells proliferation via activation of p53 mediated by upregulation of LncRNA MEG3. Biochemical and biophysical research communications 471, 290–295, doi:10.1016/j.bbrc.2016.01.169 (2016).
Modali, S. D., Parekh, V. I., Kebebew, E. & Agarwal, S. K. Epigenetic regulation of the lncRNA MEG3 and its target c-MET in pancreatic neuroendocrine tumors. Molecular endocrinology (Baltimore, Md.) 29, 224–237, doi:10.1210/me.2014-1304 (2015).
Shen, Y. et al. LINC00472 expression is regulated by promoter methylation and associated with disease-free survival in patients with grade 2 breast cancer. Breast cancer research and treatment 154, 473–482, doi:10.1007/s10549-015-3632-8 (2015).
Fu, Y. et al. Long non-coding RNAs, ASAP1-IT1, FAM215A, and LINC00472, in epithelial ovarian cancer. Gynecologic oncology, doi:10.1016/j.ygyno.2016.09.021 (2016).
Narayanan, R. Phenome-Genome Association Studies of Pancreatic Cancer: New Targets for Therapy and Diagnosis. Cancer genomics & proteomics 12, 9–19 (2015).
Kotake, Y. et al. Long Non-coding RNA, PANDA, Contributes to the Stabilization of p53 Tumor Suppressor Protein. Anticancer research 36, 1605–1611 (2016).
Puvvula, P. K. et al. Long noncoding RNA PANDA and scaffold-attachment-factor SAFA control senescence entry and exit. Nature communications 5, 5323, doi:10.1038/ncomms6323 (2014).
Dong, L. et al. Circulating Long RNAs in Serum Extracellular Vesicles: Their Characterization and Potential Application as Biomarkers for Diagnosis of Colorectal Cancer. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology 25, 1158–1166, doi:10.1158/1055-9965.epi-16-0006 (2016).
Permuth, J. B., Trevino, J., Merchant, N. & Malafa M. Partnering to advance early detection and prevention efforts for pancreatic cancer: the Florida Pancreas Collaborative. Future Oncology 12(8), 997–1000 (2016).
Fenstermacher, D. A., Wenham, R. M., Rollison, D. E. & Dalton, W. S. Implementing personalized medicine in a cancer center. Cancer J 17, 528–536, doi:10.1097/PPO.0b013e318238216e (2011).
Blondal, T. et al. Assessing sample and miRNA profile quality in serum and plasma or other biofluids. Methods 59, 164–169, doi:10.1016/j.ymeth.2012.09.015 (2013).
Kirschner, M. B. et al. The Impact of Hemolysis on Cell-Free microRNA Biomarkers. Front Genet 4, 94, doi:10.3389/fgene.2013.00094 (2013).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3, Article3, doi:10.2202/1544-6115.1027 (2004).
Chen, D. T. et al. Prognostic and predictive value of a malignancy-risk gene signature in early-stage non-small cell lung cancer. Journal of the National Cancer Institute 103, 1859–1870, doi:10.1093/jnci/djr420 (2011).
Chen, D. T. et al. Proliferative genes dominate malignancy-risk gene signature in histologically-normal breast tissue. Breast cancer research and treatment 119, 335–346, doi:10.1007/s10549-009-0344-y (2010).
Marchion, D. C. et al. BAD phosphorylation determines ovarian cancer chemosensitivity and patient survival. Clinical cancer research: an official journal of the American Association for Cancer Research 17, 6356–6366, doi:10.1158/1078-0432.CCR-11-0735 (2011).
Hopewell, E. L. et al. Lung tumor NF-kappaB signaling promotes T cell-mediated immune surveillance. The Journal of clinical investigation. doi:10.1172/JCI67250 (2013).
Youden, W. J. Index for rating diagnostic tests. Cancer 3, 32–35 (1950).
Benjamini, Y. H. & Controlling, Y. the False Discovery Rate: a Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society B 57, 289–300 (1995).
Pencina, M. J., D’Agostino, R. B. Sr., D’Agostino, R. B. Jr. & Vasan, R. S. Evaluating the added predictive ability of a new marker: from area under the ROC curve to reclassification and beyond. Statistics in medicine 27, 157–172; discussion 207–112, doi:10.1002/sim.2929 (2008).
Waters, J. A. et al. CT vs MRCP: optimal classification of IPMN type and extent. J Gastrointest Surg 12, 101–109, doi:10.1007/s11605-007-0367-9 (2008).
Acknowledgements
This study was supported in part by: the State of Florida and the Florida Academic Cancer Center Alliance (FACCA), Innovation Award Funding from Moffitt Cancer Center (J.B.P), and Institutional Research Grant number 93-032-16 from the American Cancer Society (J.B.P). The research was also made possible through the Total Cancer Care™ Protocol at the H. Lee Moffitt Cancer Center & Research Institute and by the Collaborative Data Services, Tissue, Molecular Genomics, and Biostatistics Core Facilities at the H. Lee Moffitt Cancer Center & Research Institute, an NCI designated Comprehensive Cancer Center (P30-CA076292).
Author information
Authors and Affiliations
Contributions
J.B.P., S.J.Y., and D.-T.C. conceived and designed the study, S.J.Y. and A.T.S. performed the experiments, D.-T.C. and J.L. analyzed the data, K.J., D.C., B.A.C., M.P.M. and P.H. contributed clinical samples and data, and J.B.P. and S.J.Y. wrote the initial draft of the manuscript. J.B.P.,D.-T.C., S.J.Y., A.T.S., J.W.C., J.K., Y.B., K.J., D.C., B.A.C., J.K., P.H., F.K., J.G.T., N.M., A.M., M.P.M., and R.G. reviewed, edited, and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
A provisional patent application pertinent to this manuscript content was filed on May 3, 2017 (serial number: 62/501,040).
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Permuth, J.B., Chen, DT., Yoder, S.J. et al. Linc-ing Circulating Long Non-coding RNAs to the Diagnosis and Malignant Prediction of Intraductal Papillary Mucinous Neoplasms of the Pancreas. Sci Rep 7, 10484 (2017). https://doi.org/10.1038/s41598-017-09754-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09754-5
This article is cited by
-
Blood-derived lncRNAs as biomarkers for cancer diagnosis: the Good, the Bad and the Beauty
npj Precision Oncology (2022)
-
High-throughput analysis of ANRIL circRNA isoforms in human pancreatic islets
Scientific Reports (2022)
-
Association study of the functional variants of the GLIS3 gene with risk of knee osteoarthritis
Clinical Rheumatology (2021)
-
Research progress on long non-coding RNAs and their roles as potential biomarkers for diagnosis and prognosis in pancreatic cancer
Cancer Cell International (2020)
-
The multifaceted roles of long noncoding RNAs in pancreatic cancer: an update on what we know
Cancer Cell International (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.