Abstract
Epidemiological studies have shown a strong correlation between tumor and AF. However, the molecular link between tumor and AF remains unknown. ECRG4, a tumor suppressor gene that is expressed in the A-V node and in sporadic ventricular myocytes, inhibits tumorigenesis and monitors tissue homeostasis by functioning as a ‘sentinel’ molecule gauging inflammatory and cell proliferative responses. To explore the potential physiological function of Ecrg4 in heart, we evaluated its distribution in heart, analyzed its expression in patients with persistent AF and in a canine AF model, and dissected the molecular events downstream of Ecrg4. The results showed that the level of Ecrg4 expression is homogenously high in atria and the conduction systems and in sporadic ventricular myocytes. Importantly, the expression of Ecrg4 was significantly decreased in atrial appendages of AF patients than patients with SR. Moreover, in rapid pacing canine AF models, the expression of ECRG4 in atria was significantly decreased compared to that of the controls. Mechanistically, knockdown ECRG4 in atrial myocytes significantly shortened the APDs, inhibited the expression of Gja1, and activated pro-inflammatory cascades and genes involved in cardiac remodeling. These results suggest that Ecrg4 may play a critical role in the pathogenesis of AF.
Similar content being viewed by others
Introduction
Atrial fibrillation (AF), the most common form of cardiac arrhythmia, affects 1.5–2% population <65 years old and up to 10% in those >80 years old. It is becoming an epidemic as population ages1. Persistent or inadequately treated AF may cause blood coagulation in heart, leading to stroke, congestive heart failure, myocardial infarction and other grave medical conditions. AF is a heterogeneous disease and current treatment strategies aiming at symptom relief through rate and rhythm controls and prevention of complications are far from satisfactory and sometimes even precipitate arrhythmia occurrence2. Therefore, studies that explore the molecular mechanisms underlying AF are urgently needed.
Although significant progress has been achieved in the management of AF, the pathogenesis of AF remains incompletely understood. Traditional AF predisposing factors include ageing, cardiovascular diseases, lung disease, thyroid disease and cardiac surgery. Recently, some systemic diseases such as obesity, diabetes mellitus, tumor, inflammatory bowel diseases, gout, and even psoriasis have been associated with increased risk of AF. Most, if not all, these predisposing medical conditions have been found to have a sterile, persistent, and mild systemic inflammation3. Indeed, substantial evidence has shown that inflammation promotes the initiation and perpetuation of AF4,5,6,7. Atrial myocarditis was first discovered in patients with lone non-postoperative AF, which was not observed in subjects in Sinus rhythm8. Subsequently, epidemiological studies showed that serum C-reactive protein (CRP) and inflammatory cytokines such as TNFα, IL1, IL8, IL6, and monocyte chemoattractant protein-1 (MCP1) were significantly up-regulated in AF patients compared to those in Sinus rhythm. The increased level of CRP has been reported to predict the occurrence/re-occurrence of AF9,10,11. Consistent with the role of inflammation in the development of AF, drugs that have anti-inflammatory effect such as glucocorticoids, statins, poly-unsaturated fatty acid, colchicine, angiotensin converting enzyme inhibitors and angiotensin receptor inhibitors can reduce the risk of AF, and some have been used to prevent and even treat postoperative AF12,13,14,15. Furthermore, inflammatory mediators have been shown to disrupt calcium homeostasis, activate pro-fibrotic pathway, inhibit gap-junction intercellular communication (GJIC), and induce cardiomyocyte necrosis and apoptosis, which can all contribute to the formation of arrhythmogenic substrate16,17,18.
Recently, the association between the incidence of AF and cancer has attracted much attention19. In a case-control study, patients with AF had a 10-fold increased incidence of colorectal cancer compared to those without AF20. In a cohort study, patients with newly-diagnosed AF had a >5-fold increased cancer diagnosis than the expected incidence of cancer in general population in a 3-month follow-up21. The association was also supported by the finding that the incidence of AF in patients admitted for cancer surgery was >2 fold than those for non-neoplastic surgery22. These results warrant further investigation of the potential link between tumor and AF.
Esophageal Cancer Related Gene-4(Ecrg4) was originally characterized as a tumor suppressor gene23. It is constitutively expressed in many tissues at quiescent state and significantly decreased in cancer24. Ecrg4 encodes a pre-propeptide that is secreted but tethered on cell surface where it is colocalized with innate immunity complex TLR4-MD2-CD1425. Upon injury, Ecrg4 is shed immediately from the surface and its gene expression is rapidly decreased in 24 hours, which is accompanied by the activation of inflammatory cascades and tissue proliferative response26. Restoration of Ecrg4 significantly inhibited cell proliferation and migration, and decreased inflammation through suppression of the key inflammatory regulator, NF-κb and its downstream target gene, cyclooxygenase-2(COX2)24, 26. In addition, naïve T cells express high level of Ecrg4 that is down-regulated upon activation27. These discoveries suggest that constitutive expression of Ecrg4 in quiescent state monitors tissue homeostasis, whereas loss of Ecrg4 activates inflammatory and tissue proliferative responses that contribute to the development of tumor, wound resolution, and possibly atrial remodeling28, 29.
Ecrg4 is expressed in atrioventricular (A-V) node, atria, and sporadic ventricular myocytes30, 31. In light of the down-regulation of Ecrg4 and high incidence of AF in cancer patients, we hypothesize that Ecrg4 may play an important role in the pathogenesis of AF.
Materials and Methods
Materials
Immunohistochemistry (IHC) kit and 2 × PCR blue-mix were purchased from Shenggong Biotech Inc. (Shanghai, China). SYBR™ Green master mixes were from Thermo Fisher. Oligo nucleotides were synthesized by Qingke Biotech Inc. (Chengdu, China). Sodium pentobarbital was the product of Merck Inc. All other chemicals, unless specified otherwise, were from Sigma.
Ethics
All animal studies were performed in accordance with the Chinese National Guidelines for the Use and Care of Experimental Animals. All protocols for animal studies were approved by the Experimental Animal Ethics Committee of Southwest Medical University. All methods for the human tissue harvest and use were performed in accordance with an approved protocol by the Ethics Committee of Southwest Medical University.
Harvest of specimens of human atrial appendage
Human right atrial appendages were harvested from 5 patients with persistent AF and 5 age- and gender-matched patients in Sinus rhythm (SR) respectively from the affiliated hospital of Southwest Medical University, during heart surgery, and an informed consent was obtained prior to the surgery.
Establishment of canine AF model
The AF model was established as described previously32. Briefly, adult mongrel dogs were anesthetized by injecting 30 mg/kg sodium pentobarbital intravenously, followed by implantation of a 6 F bipolar electrode in the right atria through the right femoral vein. AF group underwent rapid atria pacing at a frequency of 600 beats/minute for a total of 6 hours. Burst pacing (duration = 0.5 ms, interval = 100 ms, voltage = twice the diastolic threshold) was applied to atria before and 6 hour after the pacing to confirm the electrical atria re-modeling. Each burst lasted for 30 seconds, followed by a 2-minute interval. A total of 10 bursts were applied to each dog, and the EKGs were recorded using a multi-leads electrophysiological instrument (LEAD 7000A, Sichuan Jinjiang Electronic Science and Technology Co., Ltd. Chengdu). The criteria for a successful establishment of AF model were EKGs showing no P-wave, and f-wave of different sizes, intervals, and shapes that had lasted for at least 10 seconds. The duration of AF was defined as the time period from the successful induction of AF to the reversal of f-wave to the Sinus P-wave. A total of 7 AF models and 6 controls were used for gene expression analysis.
Isolation of neonatal cardiomyocytes
Neonatal Spague-Dawley (SD) rats of 1–3 days old were operated to expose their hearts using thoracotomy. The hearts were harvested, and blood was rinsed with sterile PBS in a 10 cm cell culture dish. Atria and ventricles were identified under the magnifying microscopy and harvested separately. The specimens were flushed twice with PBS, transferred into a glass culture bottle, which were then digested by trypsin at 4 °C for 12–16 hours, followed by digestion with Collagenase II for 10 min at 37 °C. Single cells were collected and plated into 100 mm culture dishes or 6-well plates containing coverslips at appropriate density in 10% DMEM containing low glucose.
Immunohistochemistry and immunofluorence
Nine SD rats of 200–250 grams were intramuscularly anesthetized with 3% pentobarbital sodium (30 mg/kg body weight) and decapitated, the hearts were quickly excised and placed into HEPES-buffered Tyrode’s solution (4 °C) consisting of the following components (in mM): NaCl 137.0, KCl 5.9, MgCl2 1.2, CaCl2 1.8, glucose 12.0, HEPES 10.0, titrated to pH 7.4 with NaOH. Atrium, ventricle, and Sinoatrial node (SAN) strip of each rat were dissected separately. The SAN strip, defined as the region bordered by its anatomic landmarks (the crista terminalis, the interatrial septum, and the superior and inferior vena cavae), was dissected from heart tissues. Tissues were washed twice with the Tyrode’s solution, and immediately fixed in 4% paraformaldeyhyde for overnight and embedded for tissue microtome. Tissues were sectioned at 4 mm and Immunohistochemistry was performed using the ICH kit (Shenggong Biotech Inc., Shanghai) following manufacturer’s instructions. The primary antibody was anti-Ecrg4 (Sigma) with a dilution of 1:500 and the secondary antibody was goat anti-rabbit-HRP at 1:10,000 dilution. Human atrial appendage specimens were processed the same way as that for adult rat heart. For immunofluorence of neonatal cardiomyocytes (CMs), newly-isolated CMs were seeded in 6-well plate containing coverslips and continued to incubate for overnight. The cells from 3–4 coverslips were then washed with PBS, fixed with 4% paraformadelhyde, permeabilized with 0.1% Triton X-100, and then proceeded for immunofluorence with anti-Ecrg4, followed by goat anti-rabbit Alexa Fluor 488 (Thermo Fisher), and nuclear counter-staining with DAPI (Thermo Fisher). The coverslips were then mounted and images were acquired with confocal microscopy.
Construction of lentivirus expressing ECRG4 small RNA and knockdown ECRG4 in neonatal cardiomyocytes
Small RNAs targeting human ECRG4 were designed using the program (http://katahdin.cshl.org:9331/homepage/siRNA/RNAi.cgi?type=shRNA) recommended in the pGreenPuroTM manual (System Biosciences). The two oligos (designated as E16 and E25) containing the core siRNA sequences, 5′-TGGCCGTTGATGAGAATAAAG-3′ and 5′-ACTACCAACGTCACTATGATG-3′ targeting ECRG4 coding sequence, were annealed with their corresponding reverse complementary strands and cloned into pGreenPuro, a bicistronic lentiviral vector co-transcribing GFP downstream of the siRNA precursor. The identity of each plasmid was confirmed by DNA sequencing. The knockdown efficiency was determined by transient transfection of PC3 cells, a prostate cancer cell line, and the expression of Ecrg4 was evaluated by real-time PCR. E16 had a better knockdown efficiency and was used in the knockdown experiment. A plasmid containing siRNA targeting firefly luciferase (Luc-siRNA) was used as a control. Lentiviruses were packaged using pPACKH1 Lentivector Packaging Kit (SBI, Cat #LV500A-1) and tittered by flow cytometry using limiting dilution. About 3 × 105 PFU virus was used to transduce neonatal CMs in the presence of 4–8 μg/ml Polybrene in a 6-well plate containing coverslips for 36 hours. The coverslips were used for patch-clamping to record action potential (AP) and the cells on the edge of the wells were used to confirm the knockdown of ECRG4 and to evaluate the expression of genes commonly involved in inflammation and atria remodeling by real-time PCR.
Action potential recording using patch-clamp
Action potential was recorded using patch-clamp amplifier (EPC-10, HEKA, and Germany). Single-pipette whole-cell patch-clamp technique under current clamp configuration was used to record the APs of single cardiomyocyte. The pipette solution contained (in mmol/L): KCl 140, MgCl2 1, HEPES 5, Glucose 10, EGTA 10, Mg2ATP 5, GTP 0.1, phosphocreatine 5 (pH 7.2, adjusted with KOH). The bath solution contained (in mmol/L): NaCl 137, KCl 5.4, MgCl2 1.2, NaH2PO4 1.2, HEPES 20, Glucose 10, CaCl2 1.5 (pH 7.4, adjusted with NaOH). APs were elicited by depolarizing pulses with width of 2 ms, intensity of 800 pA and frequency of 1 Hz. Data were sampled at 20 kHz and filtered at 1 kHz. A total of 8 GFP-positive CMs from Ecrg4-siRNA and Luc-siRNA knocking down, respectively, were used to record APs. Action potential durations at 50% repolarization (APD50) and 90% repolarization (APD90) were analyzed. All the experiments were conducted at room temperature (22–25 °C).
Gene expression analysis
After harvesting, rat and canine hearts were washed with Tyrode’s solution, and atria, ventricles, and Sinoatrial node (SAN) strips were dissected separately, which were immediately snap-frozen in liquid nitrogen and stored at −80 °C freezer. For RNA preparation, neonatal CMs were immediately processed after experiments and frozen tissues were first grinded with an electric grinder, followed by total RNA extraction with Trizol following the protocol recommended by the vendor (Life Technologies). One μg of total RNA was reverse transcribed (BioRad) in 20 μl reaction containing 5 μl of 5x reaction buffer and 1 μl of reverse transcriptase. Of the 20 μl cDNA, 1 μl was used for real-time PCR. Real-time PCR was performed in a 25 μl of reaction consisting of 12.5 μl CYBR mix, 1 μl of each primer at 10 μM, 1 μl of cDNA, and 9.5 μl of water. The cycler conditions for both genes of interest and GAPDH were 95 °C for 5 min., 45 cycles of 94 °C for 30 sec., 62 °C for 30 sec., and 72 °C for 30 sec. The PCR efficiency for all primer pairs were greater than 95%. Relative expression was calculated using ΔΔCt method as described in BioRad’s real time PCR manual. Primers for ECRG4 are sense: 5′-CCCAGGTGGCATAAGTGGAA-3′ and anti-sense: 5′-ACTGCTGGTACCACTGCTGCACC-3′. Primers for GAPDH are sense: 5′-GAGCGAGATCCCTCCAA-3′ and anti-sense: 5′-ACTGTGGTCATGAGTCCTTC-3′. Primers for others are available upon request.
Statistical analysis
The data, expressed as ‘Mean ± SD’, were analyzed using Student’s t test with SPSS19.0 software. *P < 0.05 was considered to be statistically significant.
Data availability statement
Data published in this article are available upon request.
Results
Distribution of Ecrg4 in heart
It has been reported that Ecrg4 is expressed in the A-V node and sporadic ventricular myocytes of adult rat heart30, 31. To explore the potential function of Ecrg4 in heart, we analyzed its distribution in adult SD rat heart by immunohistochemistry (IHC) and real-time PCR. Rats were anesthetized and hearts were harvested, washed in PBS, and dissected into Sinus node, left and right atria, and left and right ventricles. Each tissue was split into two parts, one for IHC and the other for real-time PCR. Figure 1 are representative IHC images showing a homogenous stronger Ecrg4 immunopositive staining (brown color) in Sinus node (1A), A-V node (1B) and left atrium (1C), and sporadic strong Ecrg4 immunopositive myocytes (1D) can be seen in left ventricle. When the expression of ECRG4 was evaluated by real-time PCR (n = 9) (Fig. 2), Sinus node (SN) and left atria (L. At.) express the highest level of ECRG4, followed by right atria (R. At.), and then right and left ventricles (R. Vt. and L. Vt.). Atria express significantly higher level of ECRG4 than that of ventricles (P < 0.05), and left atria express higher level of ECRG4 than that of right atria (P < 0.05).
Distribution of Ecrg4 in adult rat heart by immunohistochemistry. Specimens of Sinus node, A-V node, left atrium, and left ventricle were dissected from adult SD rat hearts, and processed for IHC as described. Representative images showing that Ecrg4 (brown staining) is expressed in Sinus node (A), A-V node (B), left atrium (C), and sporadically in ventricular myocytes of left ventricle (D). Scale bar, 200 µm.
Distribution of Ecrg4 in adult rat heart by real-time PCR. Hearts were harvested from 9 adult rats, specimens of Sinus node, left and right atria (L. and R. At.), and left and right ventricles (L. and R. Vt.) were dissected, which were processed for total RNA isolation and real-time PCR as described in materials and methods. SN and L. At. express the highest level of ECRG4, followed by R. At. and both ventricles. Atria express significantly higher level of ECRG4 than ventricles (n = 9, P < 0.05), and L. At. express higher level of ECRG4 than R. At. (n = 9, P < 0.05). Data were presented as “Mean ± SD”, and the experiments were performed in triplicate and repeated three times.
Neonatal cardiomyocytes express higher level of Ecrg4 in atria than ventricles
To confirm that atria express relatively higher level of Ecrg4 than that of ventricles, neonatal rat atrial and ventricular myocytes were isolated separately and expression of Ecrg4 was evaluated by immunofluorence. As shown in Fig. 3, a much stronger Ecrg4-positive immunofluorence signal (green color) localized to peri-nucleus and endoplasmic reticulum (ER) regions was seen in atrial myocytes (A, left panel) compared to a faint homogeneous immunofluorence in the cytoplasm of ventricular myocytes (B, right panel). This result suggests that atria express higher level of Ecrg4.
Ecrg4 is expressed in neonatal rat cardiomyocytes. Neonatal atrial and ventricular myocytes were prepared separately as described in materials and methods, which were seeded in a 6-well plate containing coverslips for overnight. Cells were fixed, permeabilized, and proceeded for immunofluorence using anti-Ecrg4 as primary antibody, goat anti-rabbit Alexa Fluor 488 as secondary antibody, and DAPI for nuclear staining. Representative images showing relative higher level of Ecrg4 (bright green) in perinuclear region and in endoplasmic reticulum in neonatal atrial myocytes (A, left panel) compared to relative lower level of Ecrg4 (faint green) diffused in the cytoplasm of ventricular myocytes (B, right panel). Scale bar = 50 µm.
Expression of Ecrg4 was down-regulated in appendages of AF patients
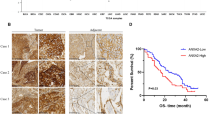
To explore the clinical relevance of Ecrg4 expression in atria and the conduction systems, we analyzed the expression of Ecrg4 by IHC in patients of rheumatic heart disease (RHD) with or without AF. Specimens of atrial appendage of 5 female patients with persistent AF (labeled AF1, 2, 3, 4, and 5) and 5 age-, and gender-matched patients with Sinus rhythm (SR, labeled SR1, 2, 3, 4, and 5) were harvested after surgery. Table 1 lists the age and the diagnosis of each patient. Specimens were fixed, paraffin embedded, microtomed, and proceeded for IHC as described in materials and methods. As shown in Fig. 4, the Ecrg4-specific immunoreactive staining shown as brown color, with 200x magnification on the left side and 400x magnification on the right side of (A) and (B) respectively, was significantly less intense in the appendages of patients with persistent AF (A, left panel) than control patients with SR (B, right panel), suggesting that Ecrg4 may participate in the pathogenesis of AF.
Expression of Ecrg4 is down-regulated in appendages of AF patients. Specimens of atrial appendage from right atrium were harvested from 5 RHD patients with persistent AF (A, labeled AF1–5 on left panel) and 5 RHD patients with SR (B, labeled SR1–5 on right panel) respectively. The specimen were fixed, microtomed, and proceeded for IHC using anti-Ecrg4 as primary antibody and goat anti-rabbit-HRP as secondary antibody per instructions in materials and methods. Representative images (with 200x magnification on the left side and 400x magnification on the right side of A and B respectively) showing a light brown staining of cardiomyocytes (low level of Ecrg4) in all 5 specimens of atrial appendages from patients with AF (A, left panel) relative to a dark brown staining of cardiomyocytes (high level of Ecrg4) in all 5 specimens of atrial appendages from patients with SR (B, right panel).
Expression of Ecrg4 was down-regulated in atria in a canine AF model
To confirm the potential role of Ecrg4 in the pathogenesis of AF, a canine AF model was established by rapid atria pacing (RAP) as described previously32 and the successful establishment of AF model was confirmed by burst pacing as described in materials and methods. After continuous RAP for 6 hours, dogs were euthanized and left atrial appendages were collected for gene expression analysis. Figure 5A shows representative recordings of the regular normal AP waves in a control dog and 5B shows fast and irregular atrial waves in an AF model induced by a burst pacing. Wthen the expression of ECRG4 was analyzed by real-time PCR (Fig. 5C), it showed that ECRG4 was significantly down-regulated in AF models (black bar, n = 8) compared to control dogs (Open Bar) (P < 0.05). These results indicate that Ecrg4 may participate in the development of AF.
Establishment of canine AF model and expression of Ecrg4 in atria. AF was induced in canines using rapid atria pacing as described in materials and methods. AF was confirmed using surface and intracardiac electrograms showing a normal regular P and atrial waves (A) in a control dog relative to fast and irregular atrial waves (B) in an AF model dog. When gene expression was analyzed by real-time PCR (C), ECRG4 was significantly down-regulated (n = 8, P < 0.05) in AF models (AF, black bar) compared to controls in SR (SR, Open Bar). Data were presented as “Mean ± SD”, and the experiments were performed in triplicate and repeated three times.
Knockdown Ecrg4 affects the APDs and genes commonly involved in atrial remodeling
To explore the potential molecular mechanisms of Ecrg4 in the pathogenesis of AF, neonatal atrial myocytes were seeded in a 6-well plate containing coverslips, and transduced on the next day with lentiviruses expressing Ecrg4-siRNA to knockdown Ecrg4 or luciferase-siRNA (Luc-siRNA) as a control as described in materials and methods. Cells on the coverslips were used for action potential recording, and cells on the edge of each well were used for real-time PCR confirming the knockdown of ECRG4 and analyzing the expression of pro-inflammatory cytokines and genes commonly involved in atrial remodeling. Thirty six hours after lentivirus transduction, the expression of ECRG4 was 74% lower in Ecrg4-siRNA (A, black bar) than Luc-siRNA (A, open bar) transduced atrial myocytes (P < 0.05). Figure 6C is a representative AP recording curve from Luc-siRNA knockdown and Fig. 6D is a representative AP recording curve from Ecrg4-siRNA knockdown atrial myocytes. When AP durations were analyzed, there were a 53% and 81% decrease of APD50 (Fig. 6B, left panel) and APD90 (Fig. 6B, right panel) respectively in Ecrg4-siRNA (black bars) compared to Luc-siRNA (open bar) transduced atrial myocytes (P < 0.05 in both APD50 and APD90). To further identify the downstream targets of Ecrg4 in AF, we profiled genes commonly implicated in inflammation and in cardiac remodeling. Compared to Luc-siRNA (arbitrarily set as 1), knockdown ECRG4 significantly up-regulated the expression of proinflammatory genes IL1a, IL6, MCP1 (P < 0.05 in all) but not that of NF-kb p50 (ns, non-significant) (Fig. 7A), down-regulated Gap junction alpha-1 protein (Gja1) (P < 0.05), and up-regulated MMP3, s100a1 and s100a8 (Fig. 7B) (P < 0.05 in all). These results suggest that Ecrg4 may play a critical role in the pathogenesis of AF, and loss of Ecrg4 may initiate the cascades of atrial remodeling.
Knockdown ECRG4 in neonatal atrial myocytes shortens APD. Neonatal atrial CMs were transduced with lentivirus expressing Ecrg4-siRNA or luciferase-siRNA for 36 hours and the knock down of ECRG4 was confirmed by real-time PCR showing a 74% decreased ECRG4 expression in Ecrg4-siRNA (black bar) relative to Luc-siRNA transduction (open bar) (A, P < 0.05). Thirty-six hours after transduction, AP was recorded using patch-clamp on GFP positive CMs. Representative recordings showing APs in Luc-siRNA (C) and Ecrg4-siRNA (D) transduced CMs. Quantitative analysis of APD50 (B, left panel) and APD90 (B, right panel) showed 53% and 81% decrease, respectively, in Ecrg4-siRNA (black bars) relative to Luc-siRNA (open bar) transduced atrial myocytes (n = 8, P < 0.05 in both APD50 and APD90). Data were presented as “Mean ± SD”, and the knockdown experiment was performed in triplicate and repeated three times.
Knockdown ECRG4 affects the expression of genes implicated in inflammation and cardiac remodeling. Knockdown ECRG4 was performed as described in materials and methods and gene expression normalized to GAPDH was expressed as relative expression of the knockdown using Luc-siRNA, which was arbitrarily set as 1. Knockdown ECRG4 significantly increased the expression of IL1a, IL6, and MCP1 (A, P < 0.05 in all) but not that of NF-kb p50 (ns, non-significant) (A), inhibited the expression of Gja1 (B, P < 0.05) and activated the expression of MMP3, s100a1 and s100a8 (B, P < 0.05 in all). Data were presented as “Mean ± SD”, and the experiments were performed in triplicate and repeated three times.
Discussion
The findings presented here show that Ecrg4 is expressed in Sinus node, A-V node, atrium and sporadic CMs of ventricle (Fig. 1), and levels of ECRG4 expression are higher in SN and atria than ventricles (Fig. 2A). In neonatal CMs, expression of Ecrg4 in atrial myocytes seems high and localized in the endoplasmic reticulum versus low and homogenous cytoplasmic distribution in ventricular myocytes (Fig. 3). Significantly, expression of Ecrg4 in atrial appendages was significantly lower in patients with persistent AF than patients with SR (Fig. 4). Moreover, in a canine AF model induced by rapid atria pacing, expression of ECRG4 in atria was significantly decreased compared to that of controls (Fig. 5C). Finally, knockdown ECRG4 in neonatal atrial myocytes significantly shortened the APD50 and APD90 (Fig. 6B–D), increased the expression of pro-inflammatory cytokines IL1a, IL6, and MCP1 (Fig. 7A), decreased the expression of Gja1, and increased the expression of MMP3, s100a1 and s100a8 (Fig. 7B). These discoveries point to the involvement of Ecrg4 in the pathogenesis of AF.
Since the cloning of ECRG4 from epithelial cells of normal esophagus23, Ecrg4 has now been characterized as a tumor suppressor gene that is constitutively expressed in normal tissues of many organs24, 26, 29, 33. Ecrg4 is not only expressed in the typical epithelial cells, but also expressed in specialized epithelial-derived cells including pituicytes, oligodendrocytes, choroid plexus epithelium, middle ear mucosa, cornea and airway epithelia, and even in adrenal gland and human leukocytes34. These wide tissue distributions are in consistent with the multiple tissue-dependent functions of Ecrg4. Mirabeau first reported that Ecrg4 was expressed in A-V node of heart by in situ hybridization31. Later, Porzionato et al. showed that Ecrg4 is homogenously expressed in atria and in sporadic ventricular myocytes of rat by immunohistochemistry30. Using both immunohistochemistry and real-time PCR, we showed that expression of Ecrg4 is high in Sinus node, A-V node and atria relative to ventricles, which agrees with microarray data that ECRG4 expression was higher in artial chamber/atria than ventricles during mouse embryo development (PubMed GEO Profile ID: 4938867) and in adult rats (PubMed GEO Profile ID: 62698595). Together, these discoveries suggest that Ecrg4 may play a critical role in normal cardiac physiology.
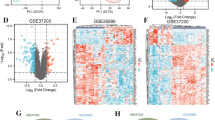
Epidemiogical studies have demonstrated a strong correlation between tumor and the incidence of AF. Predisposing factors for cancer patients to develop AF include advanced age, chronic inflammation, and cancer-induced dys-regulation of metabolism, radio- and chemo-therapy, disturbance of autonomous nerve system caused by pain and psychological stress, and surgery. However, whether tumor suppressor genes play a role in the pathogenesis of AF remains largely unknown. Kim et al. compared gene expression profiles in human atrial tissues between patients with chronic AF and those with Sinus rhythm (SR) and reported that tumor suppressor, p27 expression was significantly lower in patient with AF than patients with SR35. Genome-wide association studies (GWAS) have also associated transcription factor, zinc finger homeobox 3 (ZFHX3) with AF36, 37. ZFHX3 is a well-known tumor suppressor gene encoding AT-motif binding factor-1 (ATBF1)38. Rapid electric stimulation (RES) significantly activated STAT3 signaling pathway that has been shown to contribute to atrial remodeling39. Knockdown ZFHX3 in atrial myocytes activated STAT3, whereas over-expression of ZFHX3 inhibited the activation of STAT3 induced by RES, supporting the involvement of ATBF1 in the pathogenesis of AF40, 41. Moreover, ATBF1 interacts with Runt domain transcription factor 3 (Runt-3), and upon activation of TGFβ signaling pathway the ATBF1-Runt3 complex translocates to nucleus, where it transactivates pro-fibrotic genes, leading to increased fibrosis42. We showed that expression of Ecrg4 is decreased in atrial appendages of AF patients and in atria of a canine AF model, and knockdown ECRG4 in atrial myocytes significantly increased the expression of pro-inflammatory cykotines (IL1a, IL6, MCP1), down-regulated Gap junction alpha-1 expression (Gja1), and up-regulated the expression of s100a1, s100a8, and MMP3. S100a1 and s100a8 are members of the s100 calcium binding protein family expressed in cardiac muscle cells, which are critical regulators of cardiomyocyte Ca2+ cycling43. Together, these results support that the constitutively expressed Ecrg4 in atria monitors atrial homeostasis, and loss of Ecrg4 may promote the formation of arrhythmogenic substrate, increasing the risk of AF.
Rapid electric stimulation of cultured atrial myocytes produces electrical remodeling that mimic the typical features of atrial tachycardia remodeling in vivo. Yang et al. demonstrated that rapid electric stimulation of HL1 cells for 24 hours significantly decreased ECRG4 expression44, suggesting the involvement of Ecrg4 in atrial electric remodeling. In agreement, we found that knockdown ECRG4 in atrial myocytes significantly shortened APD. Atrial electric properties are maintained by the tightly-regulated calcium level and the coordination among ion channels. The shortening of APD can be caused by either increased outward current carried mainly by K+ and/or decreased inward current carried by Ca2+ during repolarization45. The decrease of APD can facilitate initiation and maintenance of reentrant circuits in AF46, 47. Knockdown ZFHX3 in atrial myocytes significantly abbreviated the APD, which were attributed to the increased expression of sarco/endoplasmic reticulum Ca2+-ATPase 2a (SERCA2a), ryanodine receptor, Kv1.4, Kv1.5, and Kir3, leading to larger SERCA2a activity, ultra-rapid delayed rectifier potassium currents, and transient outward currents46. However, the molecular mechanisms underlying the shortened APD in ECRG4 knockdown remain to be studied.
In summary, we have provided evidence to show that Ecrg4 is constitutively expressed in atria and the conduction systems and is down-regulated in AF. Moreover, loss of Ecrg4 modulated the expression of molecules promoting the formation of arrhythmogenic substrate, which tends to support that Ecrg4 plays a critical role in the pathogenesis of AF. However there are limitations in this study: (1) the relatively small sample size in specimens of human atrial appendage may overestimate the observed decrease of Ecrg4 expression in AF, (2) specimens of atrial appendages came from patients with RHD and at least two more other comorbidities, which might have confounded the effect of atrial remodelings caused by loss of Ecrg4, (3) although in vitro knockdown ECRG4 seems to suggest that loss of ECRG4 leads to AF, in vivo the causal relationship between Ecrg4 and AF remains to be determined, (4) Ecrg4 is a secreted molecule that may affect other cell types than CMs in heart through a paracrine mechanism, which may contribute significantly to atrial remodeling, and (5) whether restoration of Ecrg4 in the canine AF model decreases the incidence of AF and atrial remodeling process remains to be tested. Nevertheless, to our knowledge this is the first report to show that Ecrg4 is involved in the pathogenesis of AF, which may shed new light on a long-held mystery that why the incidence of AF is higher in cancer patients.
References
Sankaranarayanan, R., Kirkwood, G., Dibb, K. & Garratt, C. J. Comparison of Atrial Fibrillation in the Young versus That in the Elderly: A Review. Cardiology research and practice 2013, 976976–16, doi:10.1155/2013/976976 (2013).
Lazzerini, P. E., Capecchi, P. L., Galeazzi, M. & Laghi-Pasini, F. Biologic drugs and arrhythmic risk in chronic inflammatory arthritis: the good and the bad. Immunologic research. doi:10.1007/s12026-016-8833-7 (2016).
Lappegard, K. T., Hovland, A., Pop, G. A. & Mollnes, T. E. Atrial fibrillation: inflammation in disguise? Scandinavian journal of immunology 78, 112–119, doi:10.1111/sji.12061 (2013).
Harada, M., Van Wagoner, D. R. & Nattel, S. Role of inflammation in atrial fibrillation pathophysiology and management. Circulation journal: official journal of the Japanese Circulation Society 79, 495–502, doi:10.1253/circj.CJ-15-0138 (2015).
Tamariz, L., Hernandez, F., Bush, A., Palacio, A. & Hare, J. M. Association between serum uric acid and atrial fibrillation: a systematic review and meta-analysis. Heart rhythm: the official journal of the Heart Rhythm Society 11, 1102–1108, doi:10.1016/j.hrthm.2014.04.003 (2014).
Jiang, Z., Dai, L., Song, Z., Li, H. & Shu, M. Association between C-reactive protein and atrial fibrillation recurrence after catheter ablation: a meta-analysis. Clinical cardiology 36, 548–554, doi:10.1002/clc.22157 (2013).
Liu, T., Li, G., Li, L. & Korantzopoulos, P. Association between C-reactive protein and recurrence of atrial fibrillation after successful electrical cardioversion: a meta-analysis. Journal of the American College of Cardiology 49, 1642–1648, doi:10.1016/j.jacc.2006.12.042 (2007).
Frustaci, A. et al. Histological substrate of atrial biopsies in patients with lone atrial fibrillation. Circulation 96, 1180–1184, doi:10.1161/01.CIR.96.4.1180 (1997).
Guo, Y., Lip, G. Y. & Apostolakis, S. Inflammation in atrial fibrillation. Journal of the American College of Cardiology 60, 2263–2270, doi:10.1016/j.jacc.2012.04.063 (2012).
Gedikli, O. et al. Inflammatory markers according to types of atrial fibrillation. International journal of cardiology 120, 193–197, doi:10.1016/j.ijcard.2006.09.015 (2007).
Boos, C. J., Anderson, R. A. & Lip, G. Y. Is atrial fibrillation an inflammatory disorder? European heart journal 27, 136–149, doi:10.1093/eurheartj/ehi645 (2006).
Rahimi, K. et al. Effect of statins on atrial fibrillation: collaborative meta-analysis of published and unpublished evidence from randomised controlled trials. BMJ 342, d1250–d1250, doi:10.1136/bmj.d1250 (2011).
Levine, M. J. & Schweitzer, P. Angiotensin converting enzyme inhibitors and angiotensin II receptor blockers in atrial fibrillation. Vnitrni lekarstvi 56, 1138–1141 (2010).
Koyama, T. et al. Prevention of atrial fibrillation recurrence with corticosteroids after radiofrequency catheter ablation: a randomized controlled trial. Journal of the American College of Cardiology 56, 1463–1472, doi:10.1016/j.jacc.2010.04.057 (2010).
Deftereos, S. et al. Colchicine for prevention of early atrial fibrillation recurrence after pulmonary vein isolation: a randomized controlled study. Journal of the American College of Cardiology 60, 1790–1796, doi:10.1016/j.jacc.2012.07.031 (2012).
Mesubi, O. O. & Anderson, M. E. Atrial remodelling in atrial fibrillation: CaMKII as a nodal proarrhythmic signal. Cardiovascular research 109, 542–557, doi:10.1093/cvr/cvw002 (2016).
Hu, Y. F., Chen, Y. J., Lin, Y. J. & Chen, S. A. Inflammation and the pathogenesis of atrial fibrillation. Nature reviews. Cardiology 12, 230–243, doi:10.1038/nrcardio.2015.2 (2015).
Schotten, U., Verheule, S., Kirchhof, P. & Goette, A. Pathophysiological mechanisms of atrial fibrillation: a translational appraisal. Physiological reviews 91, 265–325, doi:10.1152/physrev.00031.2009 (2011).
Farmakis, D., Parissis, J. & Filippatos, G. Insights into onco-cardiology: atrial fibrillation in cancer. Journal of the American College of Cardiology 63, 945–953, doi:10.1016/j.jacc.2013.11.026 (2014).
Erichsen, R. et al. Colorectal cancer and risk of atrial fibrillation and flutter: a population-based case-control study. Internal and emergency medicine 7, 431–438, doi:10.1007/s11739-011-0701-9 (2012).
Ostenfeld, E. B. et al. Atrial fibrillation as a marker of occult cancer. PloS one 9, e102861, doi:10.1371/journal.pone.0102861 (2014).
Guzzetti, S., Costantino, G., Vernocchi, A., Sada, S. & Fundaro, C. First diagnosis of colorectal or breast cancer and prevalence of atrial fibrillation. Internal and emergency medicine 3, 227–231, doi:10.1007/s11739-008-0124-4 (2008).
Su, T., Liu, H. & Lu, S. Cloning and identification of cDNA fragments related to human esophageal cancer. Zhonghua zhong liu za zhi [Chinese journal of oncology] 20, 254–257 (1998).
Li, L. W. et al. Expression of esophageal cancer related gene 4 (ECRG4), a novel tumor suppressor gene, in esophageal cancer and its inhibitory effect on the tumor growth in vitro and in vivo. International journal of cancer 125, 1505–1513, doi:10.1002/ijc.24513 (2009).
Dang, X., Podvin, S., Coimbra, R., Eliceiri, B. & Baird, A. Cell-specific processing and release of the hormone-like precursor and candidate tumor suppressor gene product, Ecrg4. Cell and tissue research 348, 505–514, doi:10.1007/s00441-012-1396-6 (2012).
Kurabi, A. et al. Ecrg4 attenuates the inflammatory proliferative response of mucosal epithelial cells to infection. PloS one 8, e61394, doi:10.1371/journal.pone.0061394 (2013).
Matsuzaki, J. et al. ECRG4 is a negative regulator of caspase-8-mediated apoptosis in human T-leukemia cells. Carcinogenesis 33, 996–1003, doi:10.1093/carcin/bgs118 (2012).
Baird, A. et al. Cell surface localization and release of the candidate tumor suppressor Ecrg4 from polymorphonuclear cells and monocytes activate macrophages. Journal of leukocyte biology 91, 773–781, doi:10.1189/jlb.1011503 (2012).
Podvin, S. et al. Esophageal cancer related gene-4 is a choroid plexus-derived injury response gene: evidence for a biphasic response in early and late brain injury. PloS one 6, e24609, doi:10.1371/journal.pone.0024609 (2011).
Porzionato, A. et al. ECRG4 expression in normal rat tissues: expression study and literature review. European journal of histochemistry: EJH 59, 2458, doi:10.4081/ejh.2015.2458 (2015).
Mirabeau, O. et al. Identification of novel peptide hormones in the human proteome by hidden Markov model screening. Genome research 17, 320–327, doi:10.1101/gr.5755407 (2007).
Miyata, A., Zipes, D. P., Hall, S. & Rubart, M. Kb-R7943 prevents acute, atrial fibrillation-induced shortening of atrial refractoriness in anesthetized dogs. Circulation 106, 1410–1419, doi:10.1161/01.CIR.0000028587.85711.F6 (2002).
Gonzalez, A. M. et al. Ecrg4 expression and its product augurin in the choroid plexus: impact on fetal brain development, cerebrospinal fluid homeostasis and neuroprogenitor cell response to CNS injury. Fluids and barriers of the CNS 8, 6, doi:10.1186/2045-8118-8-6 (2011).
Baird, A. et al. Esophageal cancer-related gene 4 at the interface of injury, inflammation, infection, and malignancy. Gastrointestinal cancer: targets and therapy 2014, 131–142, doi:10.2147/GICTT.S49085 (2014).
Kim, N. H. et al. Altered patterns of gene expression in response to chronic atrial fibrillation. International heart journal 46, 383–395, doi:10.1536/ihj.46.383 (2005).
Gudbjartsson, D. F. et al. Variants conferring risk of atrial fibrillation on chromosome 4q25. Nature 448, 353–357, doi:10.1038/nature06007 (2007).
Gudbjartsson, D. F. et al. A sequence variant in ZFHX3 on 16q22 associates with atrial fibrillation and ischemic stroke. Nature genetics 41, 876–878, doi:10.1038/ng.417 (2009).
Yamada, K., Miura, Y., Scheidl, T., Yoshida, M. C. & Tamaoki, T. Assignment of the human ATBF1 transcription factor gene to chromosome 16q22.3-q23.1. Genomics 29, 552–553, doi:10.1006/geno.1995.9967 (1995).
Huang, Z. et al. Signal Transducer and Activator of Transcription 3/MicroRNA-21 Feedback Loop Contributes to Atrial Fibrillation by Promoting Atrial Fibrosis in a Rat Sterile Pericarditis Model. Circulation. Arrhythmia and electrophysiology 9, doi:10.1161/CIRCEP.115.003396 (2016).
Jiang, Q. et al. Down-regulation of ATBF1 activates STAT3 signaling via PIAS3 in pacing-induced HL-1 atrial myocytes. Biochemical and biophysical research communications 449, 278–283, doi:10.1016/j.bbrc.2014.05.041 (2014).
Nojiri, S. et al. ATBF1 enhances the suppression of STAT3 signaling by interaction with PIAS3. Biochemical and biophysical research communications 314, 97–103, doi:10.1016/j.bbrc.2003.12.054 (2004).
Mabuchi, M. et al. Tumor suppressor, AT motif binding factor 1 (ATBF1), translocates to the nucleus with runt domain transcription factor 3 (RUNX3) in response to TGF-beta signal transduction. Biochemical and biophysical research communications 398, 321–325, doi:10.1016/j.bbrc.2010.06.090 (2010).
Kraus, C. et al. S100A1 in cardiovascular health and disease: closing the gap between basic science and clinical therapy. Journal of molecular and cellular cardiology 47, 445–455, doi:10.1016/j.yjmcc.2009.06.003 (2009).
Yang, Z., Shen, W., Rottman, J. N., Wikswo, J. P. & Murray, K. T. Rapid stimulation causes electrical remodeling in cultured atrial myocytes. Journal of molecular and cellular cardiology 38, 299–308, doi:10.1016/j.yjmcc.2004.11.015 (2005).
Iwasaki, Y. K., Nishida, K., Kato, T. & Nattel, S. Atrial fibrillation pathophysiology: implications for management. Circulation 124, 2264–2274, doi:10.1161/CIRCULATIONAHA.111.019893 (2011).
Kao, Y. H. et al. ZFHX3 knockdown increases arrhythmogenesis and dysregulates calcium homeostasis in HL-1 atrial myocytes. International journal of cardiology 210, 85–92, doi:10.1016/j.ijcard.2016.02.091 (2016).
Colman, M. A. et al. Pro-arrhythmogenic effects of atrial fibrillation-induced electrical remodelling: insights from the three-dimensional virtual human atria. The Journal of physiology 591, 4249–4272, doi:10.1113/jphysiol.2013.254987 (2013).
Acknowledgements
The authors would like to thank Dr. Andrew Baird, Department of Surgery, University of California San Diego, California, for his insightful input and unconditional support in the early stage of the proposed research. We would also like to thank Ms. Xuehui Fan and Ms. Linlin Chen for preparation of neonatal CMs, and professors, Guang Li, Xiaoqiu Tan and Yan Yang for critical reading of the manuscript. The research was supported by funding from National Natural Science Foundation of China (#81570277 to XD, and #30870903 to XZ), Luzhou-Sichuan Medical University joint scientific funding (2015LZCYD-S03(4/7)), and a seeding grant from Southwest Medical University.
Author information
Authors and Affiliations
Contributions
Conceived and designed the paper (X.D., X.Z.), analyzed data and wrote the manuscript (X.D.), performed experiments (H.Y., L.H., X.F., X.L., L.M., J.C.). All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, L., Yu, H., Fan, X. et al. A Potential Role of Esophageal Cancer Related Gene-4 for Atrial Fibrillation. Sci Rep 7, 2717 (2017). https://doi.org/10.1038/s41598-017-02902-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-02902-x
This article is cited by
-
Cardiac Expression of Esophageal Cancer-Related Gene-4 is Regulated by Sp1 and is a Potential Early Target of Doxorubicin-Induced Cardiotoxicity
Cardiovascular Toxicology (2022)
-
ECRG4: a new potential target in precision medicine
Frontiers of Medicine (2019)
-
Potential functions of esophageal cancer-related gene-4 in the cardiovascular system
Frontiers of Medicine (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.