Abstract
Deep-sea hydrothermal vents are usually considered as extreme environments with high pressure, high temperature, scarce food, and chemical toxicity, while many local inhabitants have evolved special adaptive mechanisms for residence in this representative ecosystem. In this study, we constructed a high-quality genome assembly for a novel deep-sea anemone species (Actinostola sp.) that was resident at a depth of 2,971 m in an Edmond vent along the central Indian Ocean ridge, with a total size of 424.3 Mb and a scaffold N50 of 383 kb. The assembled genome contained 265 Mb of repetitive sequences and 20,812 protein-coding genes. Taken together, our reference genome provides a valuable genetic resource for exploring the evolution and adaptive clues of this deep-sea anemone.
Similar content being viewed by others
Background & Summary
Deep-sea hydrothermal vents are a representative ecosystem, where hot and chemical fluids exit the seafloor from black smoker chimneys1. These vents are considered as extremely harsh environments with high pressure, high temperature, low oxygen, and high concentrations of methane (CH4), heavy metals and hydrogen sulfide (H2S)2,3. Many species live within and around these hydrothermal vents, including various crabs, shrimps, fishes, octopus, as well as diverse sessile creatures such as sea anemones, barnacles, and tube worms4,5. These special organisms arouse many interests to developers for drugs, enzymes, cosmetics, biofuel, and other products. However, the genetic basis of evolution and adaptation of deep-sea hydrothermal vents animals is still lacking.
Sea anemones, a group of primitive Cnidarians, are widely distributed across the whole ocean depth6. Their unique adaptive strategies help them live in a variety of marine habitats from shallow waters to deep-sea trenches. During a recent expedition, an anemone (Fig. 1a) was collected at 2,971 m depth in certain hydrothermal vents of Indian Ocean (E60.5, N6.4). In this area, Actinostolidae anemones showed the highest abundance reported from previous research7. Morphological and molecular analyses suggest that this deep-sea anemone belongs to the genus Actinostola. Here, whole genome sequencing was performed to construct a high-quality genome assembly for this newfound Actinostola sp., which will help to elucidate adaptive clues to deep-sea hydrothermal environments.
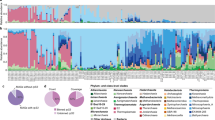
A total of 44.23-Gb paired-end reads produced by an Illumina sequencing platform were used for a genome survey (Fig. 1b). The sequencing depth with the highest frequency was identified at 54, and the total number of 17-mer reads was 19,503,242,454. Therefore, the estimated genome size of Actinostola sp. was about 487 Mb. Meanwhile, the heterozygosity rate of this genome was predicted to be 0.9% (see more details in Fig. 1b).
A 424.3-Mb draft genome was subsequently assembled based on 112.37-Gb long reads generated from a PacBio sequencing platform and 26.10-Gb short reads generated from an Illumina Hiseq Xten platform, with a contig N50 of 373 kb, a scaffold N50 of 383 kb and GC content of 38.7% (Table 1). The routine BUSCO (Benchmarking Universal Single-Copy Orthologs) method was applied to evaluate the completeness of our assembled genome, using the eukaryota_odb9 database as the reference. Finally, 252 (83.2%) BUSCO core genes were completely identified.
For further repeat annotation, a total of 265-Mb data covering 62.4% of the total assembled genome were predicted to be repeat sequences. Among them, 25.5% of the genome (108.2 Mb) was DNA repeat elements, 8.4% (35.6 Mb) was long interspersed nuclear elements (LINE), 14.3% (60.6 Mb) was long terminal repeats (LTR), and 0.8% (3.6 Mb) was short interspersed nuclear elements (SINE). After masking those repetitive regions, we applied an integrated method of homologous sequence search and de novo gene prediction to obtain annotations of 20,812 protein-coding genes in the assembled genome. By searching four public databases including GO (Gene ontology)8, KEGG (Kyoto Encyclopedia of Genes and Genomes)9, SwissProt10 and TrEMBL11, we found that 97.89% (19,111 in total) of these predicted genes were functionally annotated.
The coding sequences (CDS), predicted from assembled genomes of Actinostola sp. (this study) and other seven representative species (Fig. 1c), were utilized for clustering of gene families. Eventually, the 20,812 protein-coding genes of Actinostola sp. were clustered into 10,327 gene families, among them 3,526 were single-copy orthologous. A phylogenetic tree (Fig. 1c) was constructed based on these single-copy orthologous gene families with the maximum likelihood method, predicting that the divergence of our newfound Actinostola sp. from another sea anemone Exaiotasia diaphana occurred 305 million years ago (Mya). This high-quality reference genome for Actinostola sp. can also provide novel insights for enhancing wild resource conservation, discovering new functional genes, developing novel marine drugs, and elucidating special adaptive mechanisms.
Methods
Sample collection, library construction, and genome sequencing
A specimen of the Actinostola sp. was collected from an Edmond vent along the central Indian Ocean ridge for whole genome sequencing. Genomic DNA (gDNA) was extracted using QIAwave DNA Blood & Tissue Kit (Qiagen, Germantown, MD, USA). The genome was sequenced using a combination of sequencing techniques, including paired-end sequencing with a 500-bp inserted library on an Illumina Hiseq Xten platform (Illumina Inc., San Diego, CA, USA), and a PacBio library with an insert-size of 20 kb on a PacBio sequencing platform (Pacific Biosciences, Menlo Park, CA, USA).
Genome size estimation
The Illumina short reads were filtered with SOAPfilter v2.212. Clean reads were then used for estimation of the Actinostola sp. genome size with a 17-mer frequency distribution analysis according to the following formula13: Genome Size = Kmer_num/peak_depth, where k-mer_num is the total number of reads and peak_depth denotes the estimated peak frequency of 17-mers.
Genome assembly
Before assembly, the PacBio long sequencing reads were calibrated using LoRDEC14, along with the clean Illumina short reads. After correction, DBG2OLC15 was applied to assemble these long reads to contigs with assistance of the clean short reads. To further improve the genome accuracy, two rounds of polishing was performed with different strategies. First, Racon v1.3.116 was employed for contigs polishing based on the uncorrected PacBio long reads. Second, the clean short reads were used to polish the contigs with pilon17. After heterozygosity reducing with Redundans18, we obtained a polished genome assembly for the sequenced Actinostola sp. BUSCO19 v5.22 provided quantitative measurements for the completeness of this assembly with the popular eukaryota_odb9 database as the reference.
Genome annotation
We predicted repeat elements by de novo and homology annotations. RepeatModeler20 and LTR-FINDER21 were employed for the de novo prediction to build a repeat library. Then, the two libraries were combined and aligned to the assembled genome with RepeatMasker22. For the homology prediction, a known repeat library (Repbase23) was employed to identify repeats with RepeatMasker and RepeatProteinMask22. Tandem repeats were detected using Tandem Repeat Finder24. Finally, by integrating these data from both methods, a nonredundant set of repeat elements were obtained.
To predict protein-coding genes, protein sequences form nine representative species including California sea hare (Aplysia californica), nematode (Caenorhabditis elegans), sacoglossan sea slug (Elysia chlorotica), limpet (Lottia gigantea), two-spot octopus (Octopus bimaculoides), invasive apple snail (Pomacea canaliculata), glass anemone (Exaiptasia pallida), starlet sea anemone (Nematostella vectensis), and human (Homo sapiens), were downloaded from Ensembl25, and then they were mapped to our assembled genome with TBLASTn26. Subsequently, gene structures were predicted by GeneWise27. Finally, we integrated all these predicted results using MAKER28 to obtain a consistent gene set.
For functional annotation, BLASTp29 was applied to align the predicted protein sequences against four public databases (including SwissProt10, TrEMBL10, KEGG30 and InterPro8), and then these results were retrieved to obtain GO31 terms.
Data Records
Our final assembly and annotation data have been deposited at the NCBI with accession number JAUJYZ00000000032. Protein and gene coding sequences are uploaded into FigShare depository for public accession33. The raw reads of PacBio and Illumina sequencing were also uploaded at the NCBI with accession numbers SRR25988563- SRR2598856734.
Technical Validation
The genome assembly was 424.3 Mb with a scaffold N50 of 383 kb. For quantitative assessment of this genome assembly, we showed that 83.2% of the reference BUSCO genes (insecta_db9) were successfully identified in the final genome assembly version, suggesting remarkable completeness of this Actinostola sp. genome assembly.
Code availability
No custom scripts or code was used in this study. All software and pipelines were executed according to the manuals and protocols of related published bioinformatic tools. Corresponding versions and codes/parameters of software have been described in Methods.
References
Van Dover, C. L. & Trask, J. L. Diversity at deep-sea hydrothermal vent and intertidal mussel beds. Marine Ecology Progress Series 195, 169–178 (2000).
Little, C. T. S. & Vrijenhoek, R. C. Are hydrothermal vent animals living fossils? Trends in Ecology & Evolution 18, 582–588 (2003).
Sun, S. E., Sha, Z. & Xiao, N. The first two complete mitogenomes of the order Apodida from deep-sea chemoautotrophic environments: New insights into the gene rearrangement, origin and evolution of the deep-sea sea cucumbers. Comparative Biochemistry and Physiology Part D: Genomics and Proteomics 39, 100839- (2021).
Tunnicliffe, V., McArthur, A. G. & McHugh, D. in Advances in marine biology Vol. 34 353–442 (Elsevier, 1998).
Zierenberg, R. A., Adams, M. W. W. & Arp, A. J. Life in extreme environments: Hydrothermal vents. Proceedings of the National Academy of Sciences 97, 12961–12962 (2000).
Jamieson, A. The hadal zone: life in the deepest oceans. (Cambridge University Press, 2015).
Zhou, Y. et al. Characterization of vent fauna at three hydrothermal vent fields on the Southwest Indian Ridge: Implications for biogeography and interannual dynamics on ultraslow-spreading ridges. Deep Sea Research Part I Oceanographic Research Papers 137(JUL.), 1–12 (2018).
Hunter, S. et al. InterPro: the integrative protein signature database. Nucleic acids research 37, D211–D215 (2009).
Ogata, H. et al. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic acids research 27, 29–34 (1999).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic acids research 31, 365–370 (2003).
Kulikova, T. et al. The EMBL nucleotide sequence database. Nucleic Acids Research 32, D27–D30 (2004).
Chen, Y. et al. SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience 7, gix120 (2018).
Hequan, S., Jia, D., Mathieu, P. & Korbinian, S. findGSE: estimating genome size variation within human and Arabidopsis using k-mer frequencies. Bioinformatics 34, 550–557 (2018).
Salmela, L. & Rivals, E. LoRDEC: accurate and efficient long read error correction. Bioinformatics 30, 3506–3514 (2014).
Ye, C., Hill, C. M., Wu, S., Ruan, J. & Ma, Z. DBG2OLC: efficient assembly of large genomes using long erroneous reads of the third generation sequencing technologies. Scientific reports 6, 31900 (2016).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome research 27, 737–746 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PloS one 9, e112963 (2014).
Pryszcz, L. P. & Gabaldón, T. Redundans: an assembly pipeline for highly heterozygous genomes. Nucleic acids research 44, e113–e113 (2016).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Smit, A., Hubley, R. & Green, P. RepeatModeler Open-1.0. 2008–2010. Access date Dec (2014).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic acids research 35, W265–W268 (2007).
Chen, N. Using Repeat Masker to identify repetitive elements in genomic sequences. Current protocols in bioinformatics 5, 4.10. 11–14.10. 14 (2004).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenetic genome research 110, 462–467 (2005).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic acids research 27, 573–580 (1999).
Flicek, P. et al. Ensembl 2013. Nucleic acids research 41, D48–D55 (2012).
Gertz, E. M., Yu, Y.-K., Agarwala, R., Schäffer, A. A. & Altschul, S. F. Composition-based statistics and translated nucleotide searches: improving the TBLASTN module of BLAST. BMC biology 4, 1–14 (2006).
Birney, E., Clamp, M. & Durbin, R. GeneWise and genomewise. Genome research 14, 988–995 (2004).
Cantarel, B. L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome research 18, 188–196 (2008).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. Journal of molecular biology 215, 403–410 (1990).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic acids research 28, 27–30 (2000).
Consortium, G. O. The Gene Ontology (GO) database and informatics resource. Nucleic acids research 32, D258–D261 (2004).
Bian, C. NCBI GenBank https://identifiers.org/ncbi/insdc:JAUJYZ000000000 (2023).
Bian, C. Actinostola_sp genome and annotation. figshare https://doi.org/10.6084/m9.figshare.23659923.v1 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP459375 (2023).
Acknowledgements
The authors are grateful to the laboratory members at Dalian Ocean University for their technical assistance and helpful discussion. This research was supported by the National key R & D program of China (no. 2018YFC0310702), Outstanding Talents and Innovative Teams of Agricultural Scientific Research in the MOAA of China, the innovation team of Aquaculture Environment Safety from Liaoning Province (no. LT202009), and funds from the Research Foundation for Talented Scholars at Dalian Ocean University.
Author information
Authors and Affiliations
Contributions
L.S. and L.W. designed the project. C.L., Q.G. and C.B. prepared the DNA for sequencing; Y.H. and Z.G. analyzed the genome size and annotated the genome; C.L., C.B., Z.G. and Y.H. performed comparative genomics analysis and gene family identification; C.L., C.B. and L.W. drafted the manuscript; L.S. and Q.S. improved and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, C., Bian, C., Gao, Q. et al. Whole genome sequencing of a novel sea anemone (Actinostola sp.) from a deep-sea hydrothermal vent. Sci Data 11, 102 (2024). https://doi.org/10.1038/s41597-024-02944-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-02944-7