Abstract
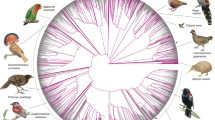
Despite tremendous efforts in the past decades, relationships among main avian lineages remain heavily debated without a clear resolution. Discrepancies have been attributed to diversity of species sampled, phylogenetic method, and the choice of genomic regions 1–3. Here, we address these issues by analyzing genomes of 363 bird species 4 (218 taxonomic families, 92% of total). Using intergenic regions and coalescent methods, we present a well-supported tree but also a remarkable degree of discordance. The tree confirms that Neoaves experienced rapid radiation at or near the Cretaceous–Paleogene (K–Pg) boundary. Sufficient loci rather than extensive taxon sampling were more effective in resolving difficult nodes. Remaining recalcitrant nodes involve species that challenge modeling due to extreme GC content, variable substitution rates, incomplete lineage sorting, or complex evolutionary events such as ancient hybridization. Assessment of the impacts of different genomic partitions showed high heterogeneity across the genome. We discovered sharp increases in effective population size, substitution rates, and relative brain size following the K–Pg extinction event, supporting the hypothesis that emerging ecological opportunities catalyzed the diversification of modern birds. The resulting phylogenetic estimate offers novel insights into the rapid radiation of modern birds and provides a taxon-rich backbone tree for future comparative studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Supplementary Results.
Supplementary Data
Table of all sequenced species with taxonomic grouping according to Howard & Moore. 4th Edition and accession numbers of the used genome assemblies. Given as a separate tab-delimited text file.
Rights and permissions
About this article
Cite this article
Stiller, J., Feng, S., Chowdhury, AA. et al. Complexity of avian evolution revealed by family-level genomes. Nature (2024). https://doi.org/10.1038/s41586-024-07323-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41586-024-07323-1
This article is cited by
-
Adaptive expansion of ERVK solo-LTRs is associated with Passeriformes speciation events
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.