Abstract
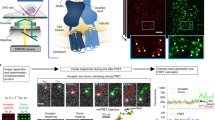
Oligomerization of membrane proteins has received intense research interest because of their importance in cellular signaling and the large pharmacological and clinical potential this offers. Fluorescence imaging methods are emerging as a valid tool to quantify membrane protein oligomerization at high spatial and temporal resolution. Here, we provide a detailed protocol for an image-based method to determine the number and oligomerization state of fluorescently labeled prototypical G-protein–coupled receptors (GPCRs) on the basis of small out-of-equilibrium fluctuations in fluorescence (i.e., molecular brightness) in single cells. The protocol provides a step-by-step procedure that includes instructions for (i) a flexible labeling strategy for the protein of interest (using fluorescent proteins, small self-labeling tags or bio-orthogonal labeling) and the appropriate controls, (ii) performing temporal and spatial brightness image acquisition on a confocal microscope and (iii) analyzing and interpreting the data, excluding clusters and intensity hot-spots commonly observed in receptor distributions. Although specifically tailored for GPCRs, this protocol can be applied to diverse classes of membrane proteins of interest. The complete protocol can be implemented in 1 month.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The original data for the experiments shown in Figs. 3–8 are available as Source Data with this protocol. Source data are provided with this paper.
Code availability
The data analysis for this study was done by using our custom-made IgorPro routine available on GitHub (https://github.com/PaoloAnnibale/MolecularBrightness). Image analysis and conversion were performed by using ImageJ, freely available at https://imagej.nih.gov/ij/. The ImageJ Stowers brightness plugins are freely available at https://research.stowers.org/imagejplugins/.
References
Sachs, J. N. & Engelman, D. M. Introduction to the membrane protein reviews: the interplay of structure, dynamics, and environment in membrane protein function. Annu. Rev. Biochem. 75, 707–712 (2006).
Jacobson, K., Sheets, E. D. & Simson, R. Revisiting the fluid mosaic model of membranes. Science 268, 1441–1442 (1995).
Klingenberg, M. Membrane protein oligomeric structure and transport function. Nature 290, 449–454 (1981).
Matthews, J. M. & Sunde, M. Dimers, oligomers, everywhere. Adv. Exp. Med. Biol. 747, 1–18 (2012).
Weis, W. I. & Kobilka, B. K. The molecular basis of G protein-coupled receptor activation. Annu. Rev. Biochem. 87, 897–919 (2018).
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schioth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Fredriksson, R., Lagerstrom, M. C., Lundin, L. G. & Schioth, H. B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 63, 1256–1272 (2003).
Pin, J. P. & Bettler, B. Organization and functions of mGlu and GABA(B) receptor complexes. Nature 540, 60–68 (2016).
Whorton, M. R. et al. A monomeric G protein-coupled receptor isolated in a high-density lipoprotein particle efficiently activates its G protein. Proc. Natl Acad. Sci. USA 104, 7682–7687 (2007).
Ferre, S. et al. G protein-coupled receptor oligomerization revisited: functional and pharmacological perspectives. Pharmacol. Rev. 66, 413–434 (2014).
Hanyaloglu, A. C. & von Zastrow, M. Regulation of GPCRs by endocytic membrane trafficking and its potential implications. Annu. Rev. Pharmacol. Toxicol. 48, 537–568 (2008).
Annibale, P. & Lohse, M. J. Spatial heterogeneity in molecular brightness. Nat. Methods 17, 273–275 (2020).
Möller, J. et al. Single-molecule analysis reveals agonist-specific dimer formation of µ-opioid receptors. Nat. Chem. Biol. 16, 946–954 (2020).
Serfling, R. et al. Quantitative single-residue bioorthogonal labeling of G protein-coupled receptors in live cells. ACS Chem. Biol. 14, 1141–1149 (2019).
Işbilir, A. et al. Advanced fluorescence microscopy reveals disruption of dynamic CXCR4 dimerization by subpocket-specific inverse agonists. Proc. Natl Acad. Sci. USA 117, 29144–29154 (2020).
Chen, Y., Muller, J. D., So, P. T. C. & Gratton, E. The photon counting histogram in fluorescence fluctuation spectroscopy. Biophys. J. 77, 553–567 (1999).
Qian, H. & Elson, E. L. Distribution of molecular aggregation by analysis of fluctuation moments. Proc. Natl Acad. Sci. USA 87, 5479–5483 (1990).
Angers, S. et al. Detection of β2-adrenergic receptor dimerization in living cells using bioluminescence resonance energy transfer (BRET). Proc. Natl Acad. Sci. USA 97, 3684–3689 (2000).
Lan, T. H. et al. BRET evidence that β2 adrenergic receptors do not oligomerize in cells. Sci. Rep. 5, 10166 (2015).
Johnson, J. A., Lu, Y. Y., Van Deventer, J. A. & Tirrell, D. A. Residue-specific incorporation of non-canonical amino acids into proteins: recent developments and applications. Curr. Opin. Chem. Biol. 14, 774–780 (2010).
Tarasova, N. I. et al. Visualization of G protein-coupled receptor trafficking with the aid of the green fluorescent protein. Endocytosis and recycling of cholecystokinin receptor type A. J. Biol. Chem. 272, 14817–14824 (1997).
Barak, L. S. et al. Internal trafficking and surface mobility of a functionally intact β2-adrenergic receptor-green fluorescent protein conjugate. Mol. Pharmacol. 51, 177–184 (1997).
Chalfie, M. GFP: lighting up life. Proc. Natl Acad. Sci. USA 106, 10073–10080 (2009).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Cutrale, F. et al. Using enhanced number and brightness to measure protein oligomerization dynamics in live cells. Nat. Protoc. 14, 616–638 (2019).
Wallin, E. & von Heijne, G. Genome-wide analysis of integral membrane proteins from eubacterial, archaean, and eukaryotic organisms. Protein Sci. 7, 1029–1038 (1998).
Goldstein, B. & Perelson, A. S. Equilibrium theory for the clustering of bivalent cell surface receptors by trivalent ligands. Application to histamine release from basophils. Biophys. J. 45, 1109–1123 (1984).
Varma, R. & Mayor, S. GPI-anchored proteins are organized in submicron domains at the cell surface. Nature 394, 798–801 (1998).
Hofman, E. G. et al. Ligand-induced EGF receptor oligomerization is kinase-dependent and enhances internalization. J. Biol. Chem. 285, 39481–39489 (2010).
Serebryannyy, L. A. et al. Persistent nuclear actin filaments inhibit transcription by RNA polymerase II. J. Cell Sci. 129, 3412–3425 (2016).
Bock, A. et al. Optical mapping of cAMP signaling at the nanometer scale. Cell 182, 1519–1530.e17 (2020).
Hebert, T. E. & Bouvier, M. Structural and functional aspects of G protein-coupled receptor oligomerization. Biochem. Cell Biol. 76, 1–11 (1998).
Maggio, R., Vogel, Z. & Wess, J. Coexpression studies with mutant muscarinic/adrenergic receptors provide evidence for intermolecular “cross-talk” between G-protein-linked receptors. Proc. Natl Acad. Sci. USA 90, 3103–3107 (1993).
Kobayashi, H., Picard, L. P., Schonegge, A. M. & Bouvier, M. Bioluminescence resonance energy transfer-based imaging of protein-protein interactions in living cells. Nat. Protoc. 14, 1084–1107 (2019).
Mercier, J. F., Salahpour, A., Angers, S., Breit, A. & Bouvier, M. Quantitative assessment of β1- and β2-adrenergic receptor homo- and heterodimerization by bioluminescence resonance energy transfer. J. Biol. Chem. 277, 44925–44931 (2002).
James, J. R., Oliveira, M. I., Carmo, A. M., Iaboni, A. & Davis, S. J. A rigorous experimental framework for detecting protein oligomerization using bioluminescence resonance energy transfer. Nat. Methods 3, 1001–1006 (2006).
Lohse, M. J. G protein-coupled receptors: too many dimers? Nat. Methods 3, 972–973 (2006).
Vilardaga, J. P., Bunemann, M., Krasel, C., Castro, M. & Lohse, M. J. Measurement of the millisecond activation switch of G protein-coupled receptors in living cells. Nat. Biotechnol. 21, 807–812 (2003).
Nikolaev, V. O., Bunemann, M., Hein, L., Hannawacker, A. & Lohse, M. J. Novel single chain cAMP sensors for receptor-induced signal propagation. J. Biol. Chem. 279, 37215–37218 (2004).
Benredjem, B. et al. Exploring use of unsupervised clustering to associate signaling profiles of GPCR ligands to clinical response. Nat. Commun. 10, 4075 (2019).
Kauk, M. & Hoffmann, C. Intramolecular and intermolecular FRET sensors for GPCRs - monitoring conformational changes and beyond. Trends Pharmacol. Sci. 39, 123–135 (2018).
Pisterzi, L. F. et al. Oligomeric size of the m2 muscarinic receptor in live cells as determined by quantitative fluorescence resonance energy transfer. J. Biol. Chem. 285, 16723–16738 (2010).
Fish, K. N. Total internal reflection fluorescence (TIRF) microscopy. Curr. Protoc. Cytom. Chapter 12, Unit 12.18 (2009).
Jonkman, J., Brown, C. M., Wright, G. D., Anderson, K. I. & North, A. J. Tutorial: guidance for quantitative confocal microscopy. Nat. Protoc. 15, 1585–1611 (2020).
Felce, J. H. et al. Receptor quaternary organization explains G protein-coupled receptor family structure. Cell Rep. 20, 2654–2665 (2017).
Moerner, W. E. & Fromm, D. P. Methods of single-molecule fluorescence spectroscopy and microscopy. Rev. Sci. Instrum. 74, 3597–3619 (2003).
Scarselli, M. et al. Revealing G-protein-coupled receptor oligomerization at the single-molecule level through a nanoscopic lens: methods, dynamics and biological function. FEBS J. 283, 1197–1217 (2016).
Wang, X. et al. Single-molecule fluorescence imaging to quantify membrane protein dynamics and oligomerization in living plant cells. Nat. Protoc. 10, 2054–2063 (2015).
Calebiro, D. et al. Single-molecule analysis of fluorescently labeled G-protein-coupled receptors reveals complexes with distinct dynamics and organization. Proc. Natl Acad. Sci. USA 110, 743–748 (2013).
Sungkaworn, T. et al. Single-molecule imaging reveals receptor-G protein interactions at cell surface hot spots. Nature 550, 543–547 (2017).
Puthenveedu, M. A. & von Zastrow, M. Cargo regulates clathrin-coated pit dynamics. Cell 127, 113–124 (2006).
Hern, J. A. et al. Formation and dissociation of M-1 muscarinic receptor dimers seen by total internal reflection fluorescence imaging of single molecules. Proc. Natl Acad. Sci. USA 107, 2693–2698 (2010).
Kasai, R. S. et al. Full characterization of GPCR monomer-dimer dynamic equilibrium by single molecule imaging. J. Cell Biol. 192, 463–480 (2011).
Giannone, G. et al. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys. J. 99, 1303–1310 (2010).
Post, S. R., Hilal-Dandan, R., Urasawa, K., Brunton, L. L. & Insel, P. A. Quantification of signalling components and amplification in the beta-adrenergic-receptor-adenylate cyclase pathway in isolated adult rat ventricular myocytes. Biochem. J. 311, 75–80 (1995).
Hesselgesser, J. et al. Identification and characterization of the CXCR4 chemokine receptor in human T cell lines: ligand binding, biological activity, and HIV-1 infectivity. J. Immunol. 160, 877–883 (1998).
Hellriegel, C., Caiolfa, V. R., Corti, V., Sidenius, N. & Zamai, M. Number and brightness image analysis reveals ATF-induced dimerization kinetics of uPAR in the cell membrane. FASEB J. 25, 2883–2897 (2011).
Godin, A. G. et al. Revealing protein oligomerization and densities in situ using spatial intensity distribution analysis. Proc. Natl Acad. Sci. USA 108, 7010–7015 (2011).
Ward, R. J., Pediani, J. D., Godin, A. G. & Milligan, G. Regulation of oligomeric organization of the serotonin 5-hydroxytryptamine 2C (5-HT2C) receptor observed by spatial intensity distribution analysis. J. Biol. Chem. 290, 12844–12857 (2015).
Digman, M. A., Dalal, R., Horwitz, A. F. & Gratton, E. Mapping the number of molecules and brightness in the laser scanning microscope. Biophys. J. 94, 2320–2332 (2008).
Hinde, E. et al. Quantifying the dynamics of the oligomeric transcription factor STAT3 by pair correlation of molecular brightness. Nat. Commun. 7, 11047 (2016).
Herrick-Davis, K., Grinde, E., Cowan, A. & Mazurkiewicz, J. E. Fluorescence correlation spectroscopy analysis of serotonin, adrenergic, muscarinic, and dopamine receptor dimerization: the oligomer number puzzle. Mol. Pharmacol. 84, 630–642 (2013).
Parmar, V. K., Grinde, E., Mazurkiewicz, J. E. & Herrick-Davis, K. Beta2-adrenergic receptor homodimers: role of transmembrane domain 1 and helix 8 in dimerization and cell surface expression. Biochim. Biophys. Acta Biomembr. 1859(9 Pt A), 1445–1455 (2016).
Hink, M. A., Shah, K., Russinova, E., de Vries, S. C. & Visser, A. J. Fluorescence fluctuation analysis of Arabidopsis thaliana somatic embryogenesis receptor-like kinase and brassinosteroid insensitive 1 receptor oligomerization. Biophys J 94, 1052–1062 (2008).
Unruh, J. R. & Gratton, E. Analysis of molecular concentration and brightness from fluorescence fluctuation data with an electron multiplied CCD camera. Biophys. J. 95, 5385–5398 (2008).
Nagy, P., Claus, J., Jovin, T. M. & Arndt-Jovin, D. J. Distribution of resting and ligand-bound ErbB1 and ErbB2 receptor tyrosine kinases in living cells using number and brightness analysis. Proc. Natl Acad. Sci. USA 107, 16524–16529 (2010).
Zamai, M. et al. Number and brightness analysis reveals that NCAM and FGF2 elicit different assembly and dynamics of FGFR1 in live cells. J. Cell Sci. 132, jcs220624 (2019).
Eichel, K., Jullie, D. & von Zastrow, M. β-Arrestin drives MAP kinase signalling from clathrin-coated structures after GPCR dissociation. Nat. Cell Biol. 18, 303–310 (2016).
Pediani, J. D., Ward, R. J., Godin, A. G., Marsango, S. & Milligan, G. Dynamic regulation of quaternary organization of the M-1 muscarinic receptor by subtype-selective antagonist drugs. J. Biol. Chem. 291, 13132–13146 (2016).
Golebiewska, U., Johnston, J. M., Devi, L., Filizola, M. & Scarlata, S. Differential response to morphine of the oligomeric state of μ-opioid in the presence of δ-opioid receptors. Biochemistry 50, 2829–2837 (2011).
Dunsing, V. et al. Optimal fluorescent protein tags for quantifying protein oligomerization in living cells. Sci. Rep. 8, 10634 (2018).
Dalal, R. B., Digman, M. A., Horwitz, A. F., Vetri, V. & Gratton, E. Determination of particle number and brightness using a laser scanning confocal microscope operating in the analog mode. Microsc. Res. Tech. 71, 69–81 (2008).
Kobilka, B. K. G protein coupled receptor structure and activation. Biochim. Biophys. Acta 1768, 794–807 (2007).
Bohme, I. & Beck-Sickinger, A. G. Illuminating the life of GPCRs. Cell Commun. Signal. 7, 16 (2009).
Daly, C. J. & McGrath, J. C. Fluorescent ligands, antibodies, and proteins for the study of receptors. Pharmacol. Ther. 100, 101–118 (2003).
Lohse, M. J., Nuber, S. & Hoffmann, C. Fluorescence/bioluminescence resonance energy transfer techniques to study G-protein-coupled receptor activation and signaling. Pharmacol. Rev. 64, 299–336 (2012).
Naganathan, S., Ye, S., Sakmar, T. P. & Huber, T. Site-specific epitope tagging of G protein-coupled receptors by bioorthogonal modification of a genetically encoded unnatural amino acid. Biochemistry 52, 1028–1036 (2013).
Dorsch, S., Klotz, K. N., Engelhardt, S., Lohse, M. J. & Bunemann, M. Analysis of receptor oligomerization by FRAP microscopy. Nat. Methods 6, 225–230 (2009).
Emami-Nemini, A. et al. Time-resolved fluorescence ligand binding for G protein-coupled receptors. Nat. Protoc. 8, 1307–1320 (2013).
Hoffmann, C. et al. A FlAsH-based FRET approach to determine G protein-coupled receptor activation in living cells. Nat. Methods 2, 171–176 (2005).
Kuder, K. & Kiec-Kononowicz, K. Fluorescent GPCR ligands as new tools in pharmacology. Curr. Med. Chem. 15, 2132–2143 (2008).
Magalhaes, A. C., Dunn, H. & Ferguson, S. S. Regulation of GPCR activity, trafficking and localization by GPCR-interacting proteins. Br. J. Pharmacol. 165, 1717–1736 (2012).
Al-Sabah, S., Adi, L., Bunemann, M. & Krasel, C. The effect of cell surface expression and linker sequence on the recruitment of arrestin to the GIP receptor. Front. Pharmacol. 11, 1271 (2020).
Schoneberg, T. et al. Mutant G-protein-coupled receptors as a cause of human diseases. Pharmacol. Ther. 104, 173–206 (2004).
Guan, X. M., Kobilka, T. S. & Kobilka, B. K. Enhancement of membrane insertion and function in a type IIIb membrane protein following introduction of a cleavable signal peptide. J. Biol. Chem. 267, 21995–21998 (1992).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128–136 (2008).
Los, G. V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Maurel, D. et al. Cell-surface protein-protein interaction analysis with time-resolved FRET and snap-tag technologies: application to GPCR oligomerization. Nat. Methods 5, 561–567 (2008).
Nikic, I. et al. Minimal tags for rapid dual-color live-cell labeling and super-resolution microscopy. Angew. Chem. Int. Ed. Engl. 53, 2245–2249 (2014).
Pandy-Szekeres, G. et al. GPCRdb in 2018: adding GPCR structure models and ligands. Nucleic Acids Res. 46, D440–D446 (2018).
Işbilir, A. et al. Visualization of class A GPCR oligomerization by image-based fluorescence fluctuation spectroscopy. Preprint at bioRxiv https://doi.org/10.1101/240903 (2017).
Arai, R., Ueda, H., Kitayama, A., Kamiya, N. & Nagamune, T. Design of the linkers which effectively separate domains of a bifunctional fusion protein. Protein Eng. 14, 529–532 (2001).
Fricke, F., Beaudouin, J., Eils, R. & Heilemann, M. One, two or three? Probing the stoichiometry of membrane proteins by single-molecule localization microscopy. Sci. Rep. 5, 14072 (2015).
Latty, S. L. et al. Referenced single-molecule measurements differentiate between GPCR oligomerization states. Biophys. J. 109, 1798–1806 (2015).
Bhatia, S., Edidin, M., Almo, S. C. & Nathenson, S. G. Different cell surface oligomeric states of B7-1 and B7-2: implications for signaling. Proc. Natl Acad. Sci. USA 102, 15569–15574 (2005).
Kraetke, O. et al. Dimerization of corticotropin-releasing factor receptor type 1 is not coupled to ligand binding. J. Recept. Signal Transduct. Res. 25, 251–276 (2005).
Teichmann, A. et al. The specific monomer/dimer equilibrium of the corticotropin-releasing factor receptor type 1 is established in the endoplasmic reticulum. J. Biol. Chem. 289, 24250–24262 (2014).
Moller, T. C. et al. Oligomerization of a G protein-coupled receptor in neurons controlled by its structural dynamics. Sci. Rep. 8, 10414 (2018).
Trullo, A., Corti, V., Arza, E., Caiolfa, V. R. & Zamai, M. Application limits and data correction in number of molecules and brightness analysis. Microsc. Res. Tech. 76, 1135–1146 (2013).
Digman, M. A. et al. Measuring fast dynamics in solutions and cells with a laser scanning microscope. Biophys. J. 89, 1317–1327 (2005).
D’Agostino, R. B. & Stephens, M. A., eds. Goodness-Of-Fit Techniques (Marcel Dekker, 1986).
Ulbrich, M. H. & Isacoff, E. Y. Subunit counting in membrane-bound proteins. Nat. Methods 4, 319–321 (2007).
Jonas, K. C., Huhtaniemi, I. & Hanyaloglu, A. C. Single-molecule resolution of G protein-coupled receptor (GPCR) complexes. Methods Cell Biol. 132, 55–72 (2016).
Serfling, R., Seidel, L., Bottke, T. & Coin, I. Optimizing the genetic incorporation of chemical probes into GPCRs for photo-crosslinking mapping and bioorthogonal chemistry in live mammalian cells. J. Vis. Exp. 134, 57069 (2018).
Herrick-Davis, K. & Mazurkiewicz, J. E. Fluorescence correlation spectroscopy and photon-counting histogram analysis of receptor-receptor interactions. Methods Cell Biol. 117, 181–196 (2013).
Stoneman, M. R. et al. A general method to quantify ligand-driven oligomerization from fluorescence-based images. Nat. Methods 16, 493–496 (2019).
Balleza, E., Kim, J. M. & Cluzel, P. Systematic characterization of maturation time of fluorescent proteins in living cells. Nat. Methods 15, 47–51 (2018).
Rizzo, M. A., Springer, G. H., Granada, B. & Piston, D. W. An improved cyan fluorescent protein variant useful for FRET. Nat. Biotechnol. 22, 445–449 (2004).
Goedhart, J. et al. Structure-guided evolution of cyan fluorescent proteins towards a quantum yield of 93%. Nat. Commun. 3, 751 (2012).
Ormo, M. et al. Crystal structure of the Aequorea victoria green fluorescent protein. Science 273, 1392–1395 (1996).
Kremers, G. J., Goedhart, J., van Munster, E. B. & Gadella, T. W. Jr. Cyan and yellow super fluorescent proteins with improved brightness, protein folding, and FRET Forster radius. Biochemistry 45, 6570–6580 (2006).
Bajar, B. T. et al. Improving brightness and photostability of green and red fluorescent proteins for live cell imaging and FRET reporting. Sci. Rep. 6, 20889 (2016).
Shaner, N. C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Shen, Y., Chen, Y., Wu, J., Shaner, N. C. & Campbell, R. E. Engineering of mCherry variants with long Stokes shift, red-shifted fluorescence, and low cytotoxicity. PloS ONE 12, e0171257 (2017).
Bindels, D. S. et al. mScarlet: a bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 14, 53–56 (2017).
Acknowledgements
We acknowledge funding by the Deutsche Forschungsgemeinschaft (German Research Foundation, DFG) CRC 1423, project number 421152132, subproject C03 (to P.A. and M.J.L.); DFG Cluster of Excellence EXC2046 Math+ (to P.A. and M.J.L.); DFG CRC 1423, project number 421152132, subproject C05 (to I.C. and A.B.); the National Institutes of Health (R01-DA038882; to M.J.L.); the Elite Network Bavaria graduate program ‘Receptor Dynamics’ (to M.J.L.); and DFG Grant CO822/2-1 (to I.C.).
Author information
Authors and Affiliations
Contributions
A.I., R.S., J.M. and P.A. performed data collection and data analysis and developed the data acquisition and analysis procedure. A.I., A.B. and U.Z. developed the cloning procedure. P.A., A.G.B.-S., R.S. and U.Z. contributed analysis tools and materials used in the research. P.A. wrote the manuscript and initiated the research. A.B. and A.I. contributed to writing the manuscript. R.T., C.D.F., M.S. and M.J.L. contributed edits to the manuscript. P.A., I.C. and M.J.L. supervised the primary research. All authors reviewed the manuscript and approved the final article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Andrew Clayton, Catherine Ann Royer and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Serfling, R. et al. ACS Chem. Biol. 14, 1141–1149 (2019): https://doi.org/10.1021/acschembio.8b01115
Annibale, P. & Lohse, M. J. Nat. Methods 17, 273–275 (2020): https://doi.org/10.1038/s41592-020-0732-0
Möller, J. et al. Nat. Chem. Biol. 16, 946–954 (2020): https://doi.org/10.1038/s41589-020-0566-1
Işbilir, A. et al. Proc. Natl Acad. Sci. USA 117, 29144–29154: https://doi.org/10.1073/pnas.2013319117
Extended data
Extended Data Fig. 1 Use of fluorescent ligands to label receptor of interest.
Confocal image of the basolateral membrane of a HEK293-AD cell expressing the Y2-receptor C-terminally labeled with EYFP (left) and labeled with 1 μM TAMRA-Ahx(5-24)NPY (center) followed by washout. Right, Upon displacement with 10 μM unlabeled Ahx(5-24)NPY, the fluorescent ligand is almost entirely displaced within tens of seconds. Scale bar, 10 μm.
Extended Data Fig. 2 Microscope calibration.
a, Theoretical autocorrelation function (ACF) for a sample of 10 particles diffusing in 2D through a point spread function (PSF) with a waist of 0.3 μm and D = 0.1 μm2/s. The pixel dwell time should allow for an accurate recording of the fluctuations. A general guideline is for the dwell time to be ~10 times smaller than the decay time of the diffusing species. In this case, the decay time is of the order of 300 ms, so any pixel dwell time smaller than 1 ms would be a very safe choice. The characteristic dwell time used in our temporal brightness acquisitions (2.4 μs) is way below this value. However, because the associated frame time is of the order of 640 ms on the Leica microscope, we felt that this was the best compromise between an acceptable photon collection and a not-too-slow acquisition time (about >1 min for 100 frames). b, Apparent brightness B versus intensity scatter plot originating from a movie (256 × 256 pixels, 100 frames) of a homogeneous mixture of Alexa488 imaged in a 90% (wt/wt) glycerol/water solution, for increasing values of the laser power. c, Change of brightness (fold change) as a function of the increase in intensity (fold change). As the mean pixel intensity increases (almost linearly with the laser power), the mean brightness scales proportionally. The linear fit (constrained to 0), has a slope of 0.9, indicating that the increase in intensity is matched by a proportional increase in brightness, as expected. d, Example of a dark count histogram for the Leica SP8 HyD photon-counting detector and for the analog PMT, the latter superposed to Gaussian + exponential fits (black dashed lines) to determine calibration parameters (see Box 2).
Extended Data Fig. 3 Example of labeling efficiency quantification (Box 2).
a, Export dual-color movies from the Leica LAS X software as tiff multipage files. b, Run the N&B processing macro. It relies on a set of plugins developed by Jay Unruh at the Stowers Institute for Biomedical Research. The macro guides the user in a step-by-step fashion to convert the tiff images to 16 bits, apply a moving average detrend to remove effects from photobleaching (optional) and calculate the average intensity and brightness value for each pixel of the movie. c, As a result, the sum intensity image is displayed in parallel to the brightness versus intensity plot (highlighted here as B/S vs I/S plot). Note that this plugin also allows for processing images that were collected with an analog detector (in this case, Slope, Zero Variance and Offset need to be adjusted to the properties of the detector of interest. For a photon-counting acquisition, leave as indicated here). A cursor selection allows for selecting those pixels belonging to cluster in as a B/S vs I/S plot. These pixels are highlighted in red in the intensity image above. The x avg value provides the average intensity, while the y avg value provides the average brightness value extracted from the selected pixels. d, For each cell analyzed, these values can be noted and then exported in the software of choice (in this case, Excel), where the number of emitters for each channel (Cy3 versus EGFP) can be plotted as a scatter and fit to a line (constrained through the origin, because in the absence of detectable aspecific binding for EGFP = 0, we assume Cy3 = 0) to determine the labeling efficiency.
Supplementary information
Source data
Source Data Fig. 3
Tabulated brightness values for Fig. 3b,c
Source Data Fig. 4
Raw acquisition move used in Fig. 4c–h
Source Data Fig. 5
Spatial brightness acquisition for Fig. 5b–d
Source Data Fig. 6
Spatial brightness for Fig. 6a–c
Source Data Fig. 7
Tabulated brightness values for Fig. 7a,b
Source Data Fig. 8
Tabulated number values for Fig. 8a,b
Source Data Extended Data Fig. 2
Combined tiff file containing acquisition of dye in solution at 5% laser power (100 frames), 10% laser power (100 frames) and 20% laser power (100 frames). Dark acquisition with HyD (100 frames) and with PMT (100 frames)
Rights and permissions
About this article
Cite this article
Işbilir, A., Serfling, R., Möller, J. et al. Determination of G-protein–coupled receptor oligomerization by molecular brightness analyses in single cells. Nat Protoc 16, 1419–1451 (2021). https://doi.org/10.1038/s41596-020-00458-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00458-1
This article is cited by
-
Insights into receptor structure and dynamics at the surface of living cells
Nature Communications (2023)
-
Unraveling the hidden temporal range of fast β2-adrenergic receptor mobility by time-resolved fluorescence
Communications Biology (2022)
-
Evaluation of sted super-resolution image quality by image correlation spectroscopy (QuICS)
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.