Abstract
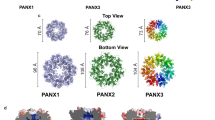
The plasma membrane adenosine triphosphate (ATP) release channel pannexin 1 (PANX1) has been implicated in many physiological and pathophysiological processes associated with purinergic signaling, including cancer progression, apoptotic cell clearance, inflammation, blood pressure regulation, oocyte development, epilepsy and neuropathic pain. Here we present near-atomic-resolution structures of human and frog PANX1 determined by cryo-electron microscopy that revealed a heptameric channel architecture. Compatible with ATP permeation, the transmembrane pore and cytoplasmic vestibule were exceptionally wide. An extracellular tryptophan ring located at the outer pore created a constriction site, potentially functioning as a molecular sieve that restricts the size of permeable substrates. The amino and carboxyl termini, not resolved in the density map, appeared to be structurally dynamic and might contribute to narrowing of the pore during channel gating. In combination with functional characterization, this work elucidates the previously unknown architecture of pannexin channels and establishes a foundation for understanding their unique channel properties.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cryo-EM maps of hPANX1 and xPANX1 have been deposited in the Electron Microscopy Data Bank with accession codes EMD-21071 and EMD-20964. Atomic coordinates for hPANX1 and xPANX1 structures have been deposited in the Protein Data Bank with accession codes 6V6D and 6UZY. Source data for Fig. 1a–d and Extended Data Figs. 2a, 4a and 5 are available with the paper online.
References
Chiu, Y. H., Schappe, M. S., Desai, B. N. & Bayliss, D. A. Revisiting multimodal activation and channel properties of pannexin 1. J. Gen. Physiol. 150, 19–39 (2018).
Dahl, G. The pannexin1 membrane channel: distinct conformations and functions. FEBS Lett. 592, 3201–3209 (2018).
Chekeni, F. B. et al. Pannexin 1 channels mediate ‘find-me’ signal release and membrane permeability during apoptosis. Nature 467, 863–867 (2010).
Qu, Y. et al. Pannexin-1 is required for ATP release during apoptosis but not for inflammasome activation. J. Immunol. 186, 6553–6561 (2011).
Billaud, M. et al. Pannexin1 regulates α1-adrenergic receptor-mediated vasoconstriction. Circ. Res. 109, 80–85 (2011).
Billaud, M. et al. A molecular signature in the pannexin1 intracellular loop confers channel activation by the α1 adrenoreceptor in smooth muscle cells. Sci. Signal. 8, ra17 (2015).
Penuela, S. et al. Loss of pannexin 1 attenuates melanoma progression by reversion to a melanocytic phenotype. J. Biol. Chem. 287, 29184–29193 (2012).
Furlow, P. W. et al. Mechanosensitive pannexin-1 channels mediate microvascular metastatic cell survival. Nat. Cell Biol. 17, 943–952 (2015).
Mousseau, M. et al. Microglial pannexin-1 channel activation is a spinal determinant of joint pain. Sci. Adv. 4, eaas9846 (2018).
Dossi, E. et al. Pannexin-1 channels contribute to seizure generation in human epileptic brain tissue and in a mouse model of epilepsy. Sci. Transl. Med. 10, eaar3796 (2018).
Thompson, R. J. et al. Activation of pannexin-1 hemichannels augments aberrant bursting in the hippocampus. Science 322, 1555–1559 (2008).
Gulbransen, B. D. et al. Activation of neuronal P2X7 receptor-pannexin-1 mediates death of enteric neurons during colitis. Nat. Med. 18, 600–604 (2012).
Karatas, H. et al. Spreading depression triggers headache by activating neuronal Panx1 channels. Science 339, 1092–1095 (2013).
Weaver, J. L. et al. Hematopoietic pannexin 1 function is critical for neuropathic pain. Sci. Rep. 7, 42550 (2017).
Sang, Q. et al. A pannexin 1 channelopathy causes human oocyte death. Sci. Transl. Med. 11, eaav8731 (2019).
Sandilos, J. K. & Bayliss, D. A. Physiological mechanisms for the modulation of pannexin 1 channel activity. J. Physiol. 590, 6257–6266 (2012).
Locovei, S., Wang, J. & Dahl, G. Activation of pannexin 1 channels by ATP through P2Y receptors and by cytoplasmic calcium. FEBS Lett. 580, 239–244 (2006).
Pelegrin, P. & Surprenant, A. Pannexin-1 mediates large pore formation and interleukin-1β release by the ATP-gated P2X7 receptor. EMBO J. 25, 5071–5082 (2006).
Iglesias, R. et al. P2X7 receptor-pannexin1 complex: pharmacology and signaling. Am. J. Physiol. - Cell Physiol. 295, C752–C760 (2008).
Lohman, A. W. et al. Pannexin 1 channels regulate leukocyte emigration through the venous endothelium during acute inflammation. Nat. Commun. 6, 7965 (2015).
Weilinger, N. L., Tang, P. L. & Thompson, R. J. Anoxia-induced NMDA receptor activation opens pannexin channels via Src family kinases. J. Neurosci. 32, 12579–12588 (2012).
Silverman, W. R. et al. The pannexin 1 channel activates the inflammasome in neurons and astrocytes. J. Biol. Chem. 284, 18143–18151 (2009).
Wang, J. et al. The membrane protein pannexin1 forms two open-channel conformations depending on the mode of activation. Sci. Signal. 7, ra69 (2014).
Qiu, F., Wang, J., Spray, D. C., Scemes, E. & Dahl, G. Two non-vesicular ATP release pathways in the mouse erythrocyte membrane. FEBS Lett. 585, 3430–3435 (2011).
Sandilos, J. K. et al. Pannexin 1, an ATP release channel, is activated by caspase cleavage of its pore-associated C-terminal autoinhibitory region. J. Biol. Chem. 287, 11303–11311 (2012).
Chiu, Y. H. et al. A quantized mechanism for activation of pannexin channels. Nat. Commun. 8, 14324 (2017).
Sridharan, M. et al. Pannexin 1 is the conduit for low oxygen tension-induced ATP release from human erythrocytes. Am. J. Physiol. Heart Circ. Physiol. 299, H1146–H1152 (2010).
Bao, L., Locovei, S. & Dahl, G. Pannexin membrane channels are mechanosensitive conduits for ATP. FEBS Lett. 572, 65–68 (2004).
Locovei, S., Bao, L. & Dahl, G. Pannexin 1 in erythrocytes: function without a gap. Proc. Natl Acad. Sci. USA 103, 7655–7659 (2006).
Bruzzone, R., Hormuzdi, S. G., Barbe, M. T., Herb, A. & Monyer, H. Pannexins, a family of gap junction proteins expressed in brain. Proc. Natl Acad. Sci. USA 100, 13644–13649 (2003).
Michalski, K., Henze, E., Nguyen, P., Lynch, P. & Kawate, T. The weak voltage dependence of pannexin 1 channels can be tuned by N-terminal modifications. J. Gen. Physiol. 150, 1758–1768 (2018).
Thompson, R. J., Zhou, N. & MacVicar, B. A. Ischemia opens neuronal gap junction hemichannels. Science 312, 924–927 (2006).
Ma, W. et al. Pannexin 1 forms an anion-selective channel. Pflugers Arch. 463, 585–592 (2012).
Romanov, R. A. et al. The ATP permeability of pannexin 1 channels in a heterologous system and in mammalian taste cells is dispensable. J. Cell Sci. 125, 5514–5523 (2012).
Chiu, Y. H., Ravichandran, K. S. & Bayliss, D. A. Intrinsic properties and regulation of pannexin 1 channel. Channels 8, 103–109 (2014).
Nielsen, B. S. et al. Pannexin 1 activation and inhibition is permeant-selective. J. Physiol. 598, 361–379 (2019).
Boassa, D. et al. Pannexin1 channels contain a glycosylation site that targets the hexamer to the plasma membrane. J. Biol. Chem. 282, 31733–31743 (2007).
Epp, A. L. et al. A novel motif in the proximal C-terminus of pannexin 1 regulates cell surface localization. Sci. Rep. 9, 9721 (2019).
Michalski, K. & Kawate, T. Carbenoxolone inhibits pannexin1 channels through interactions in the first extracellular loop. J. Gen. Physiol. 147, 165–174 (2016).
Qiu, F. & Dahl, G. A permeant regulating its permeation pore: Inhibition of pannexin 1 channels by ATP. Am. J. Physiol. Cell Physiol. 296, C250–C255 (2009).
Boyce, A. K. J. & Swayne, L. A. P2X7 receptor cross-talk regulates ATP-induced pannexin 1 internalization. Biochem. J. 474, 2133–2144 (2017).
Michalski, K. et al. The cryo-EM structure of a pannexin 1 reveals unique motifs for ion selection and inhibition. eLife 9, e54670 (2020).
Maeda, S. et al. Structure of the connexin 26 gap junction channel at 3.5 Å resolution. Nature 458, 597–602 (2009).
Oshima, A., Tani, K. & Fujiyoshi, Y. Atomic structure of the innexin-6 gap junction channel determined by cryo-EM. Nat. Commun. 7, 13681 (2016).
Deneka, D., Sawicka, M., Lam, A. K. M., Paulino, C. & Dutzler, R. Structure of a volume-regulated anion channel of the LRRC8 family. Nature 558, 254–259 (2018).
Wang, J. & Dahl, G. SCAM analysis of Panx1 suggests a peculiar pore structure. J. Gen. Physiol. 136, 515–527 (2010).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996). 376.
Schnorf, M., Potrykus, I. & Neuhaus, G. Microinjection technique: routine system for characterization of microcapillaries by bubble pressure measurement. Exp. Cell. Res. 210, 260–267 (1994).
Lee, S. Y., Letts, J. A. & MacKinnon, R. Dimeric subunit stoichiometry of the human voltage-dependent proton channel Hv1. Proc. Natl Acad. Sci. USA 105, 7692–7695 (2008).
Acknowledgements
This work was supported by startup funds from the Washington University School of Medicine (to P.Y.). M.J.R and J.A.J.F are supported by the Washington University Center for Cellular Imaging, which is funded in part by the Washington University School of Medicine through the Precision Medicine Initiative, the Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital (CDI-CORE-2015-505 and CDI-CORE-2019-813) and the Foundation for Barnes-Jewish Hospital (3770).
Author information
Authors and Affiliations
Contributions
Z.D., Z.H. and R.M.B. performed biochemical preparations, cryo-EM data processing, structural determination and analysis. G.M. conducted electrophysiology experiments. M.R., J.A.J.F., Z.D. and Z.H. collected cryo-EM data. Z.H. performed the crosslinking experiments. P.Y. designed and supervised the project. Z.D., Z.H., G.M. and P.Y. analyzed the results and prepared the manuscript with input from all authors. Correspondence and requests for materials should be addressed to P.Y.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available. Katarzyna Marcinkiewicz was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CBX inhibition of the wild-type xPANX1 and mutants in excised inside-out membrane patches.
a,b, Current-voltage relationship of wild-type xPANX1 in symmetrical NaCl (a) and KCl (b) in the absence (black) and presence (gray) of 100 µM CBX. c-e, Current-voltage relationship of the W74A (c), R75A (d), and D81A (e) mutants in symmetrical NaCl in the absence (black) and presence (gray) of 100 µM CBX.
Extended Data Fig. 2 Cryo-EM reconstruction of xPANX1.
a, Purification of xPANX1 on size-exclusion chromatography. Fractions of the monodisperse channel peak were shown in SDS-PAGE (inset). The uncropped gel image is available as source data. b, Flowchart of cryo-EM image processing. c, Fourier shell correlation before and after post-processing in RELION3. d, Fourier shell correlation between the refined model and the full map. e, Cryo-EM density map colored by local resolution in the range of 3.0-4.0 Å. f, Angular distribution plot of particles used in the final reconstruction. Only one-seventh of the sphere is shown owing to applied C7 symmetry.
Extended Data Fig. 3 Cryo-EM density map of xPANX1.
Cryo-EM density for an entire channel subunit as well as for selected regions is shown as blue mesh. Residues forming disulfide bonds are highlighted.
Extended Data Fig. 4 Cryo-EM reconstruction of human PANX1.
a, Purification of hPANX1 on size-exclusion chromatography. Fractions of the monodisperse channel peak were shown in SDS-PAGE (inset) and protein samples used for cryo-EM were indicated. The uncropped gel image is available as source data. b, Flowchart of cryo-EM image processing. c, Fourier shell correlation before and after post-processing in RELION3. d, Fourier shell correlation between the refined model and the full map. e, Cryo-EM density map colored by local resolution in the range of 3.5-6.0 Å. f, Angular distribution plot of particles used in the final reconstruction.
Extended Data Fig. 5 Crosslinking of PANX1 channels.
a, SDS-PAGE showing crosslinking of purified hPANX1 and xPANX1 proteins expressed in yeast P. pastoris. A ladder of seven distinct oligomers was detected at a medium concentration of crosslinking reagent disuccinimidyl suberate (DSS, 250 μM). At high concentrations of DSS, both xPANX1 and hPANX1 were primarily crosslinked to heptamers. Numbers on the right indicate numbers of crosslinked subunits. b, Western blots showing crosslinking of hPANX1 in the plasma membranes of mammalian cells (HEK293S). With 75 μM DSS, a ladder of crosslinked oligomers up to heptamer was observed. At high concentrations of DSS, hPANX1 protein is crosslinked to heptamers and even larger size assemblies. Uncropped images are available as source data.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 2
Full-length unprocessed SDS-PAGE gel
Source Data Extended Data Fig. 4
Full-length unprocessed SDS-PAGE gel
Source Data Extended Data Fig. 5
Full-length unprocessed SDS-PAGE gel and western blots
Rights and permissions
About this article
Cite this article
Deng, Z., He, Z., Maksaev, G. et al. Cryo-EM structures of the ATP release channel pannexin 1. Nat Struct Mol Biol 27, 373–381 (2020). https://doi.org/10.1038/s41594-020-0401-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0401-0
This article is cited by
-
Cryo-EM structures of pannexin 1 and 3 reveal differences among pannexin isoforms
Nature Communications (2024)
-
Targeting Pannexin-1 Channels: Addressing the ‘Gap’ in Chronic Pain
CNS Drugs (2024)
-
Cryo-EM structure of human heptameric pannexin 2 channel
Nature Communications (2023)
-
Direct cell extraction of membrane proteins for structure–function analysis
Scientific Reports (2023)
-
Structural and functional analysis of human pannexin 2 channel
Nature Communications (2023)