Abstract
T cells are important in tumour immunity but a better understanding is needed of the differentiation of antigen-specific T cells in human cancer1,2. Here we studied CD8 T cells in patients with human papillomavirus (HPV)-positive head and neck cancer and identified several epitopes derived from HPV E2, E5 and E6 proteins that allowed us to analyse virus-specific CD8 T cells using major histocompatibility complex (MHC) class I tetramers. HPV-specific CD8 T cells expressed PD-1 and were detectable in the tumour at levels that ranged from 0.1% to 10% of tumour-infiltrating CD8 T lymphocytes (TILs) for a given epitope. Single-cell RNA-sequencing analyses of tetramer-sorted HPV-specific PD-1+ CD8 TILs revealed three transcriptionally distinct subsets. One subset expressed TCF7 and other genes associated with PD-1+ stem-like CD8 T cells that are critical for maintaining T cell responses in conditions of antigen persistence. The second subset expressed more effector molecules, representing a transitory cell population, and the third subset was characterized by a terminally differentiated gene signature. T cell receptor clonotypes were shared between the three subsets and pseudotime analysis suggested a hypothetical differentiation trajectory from stem-like to transitory to terminally differentiated cells. More notably, HPV-specific PD-1+TCF-1+ stem-like TILs proliferated and differentiated into more effector-like cells after in vitro stimulation with the cognate HPV peptide, whereas the more terminally differentiated cells did not proliferate. The presence of functional HPV-specific PD-1+TCF-1+CD45RO+ stem-like CD8 T cells with proliferative capacity shows that the cellular machinery to respond to PD-1 blockade exists in HPV-positive head and neck cancer, supporting the further investigation of PD-1 targeted therapies in this malignancy. Furthermore, HPV therapeutic vaccination efforts have focused on E6 and E7 proteins; our results suggest that E2 and E5 should also be considered for inclusion as vaccine antigens to elicit tumour-reactive CD8 T cell responses of maximal breadth.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The following protein sequences were used for predicting and generating HPV peptides: E2 (Uniprot P03120), E5 (Uniprot P06927), E6 (Uniprot P03126), and E7 (Uniprot P03129). scRNA-seq data are available in the NCBI Gene Expression Omnibus (GEO) database under the accession number GSE180268. Other relevant data are available from the corresponding authors upon reasonable request.
Code availability
Custom code for scRNA-seq is available from the corresponding authors upon reasonable request.
References
Hashimoto, M. et al. CD8 T cell exhaustion in chronic infection and cancer: opportunities for interventions. Annu. Rev. Med. 69, 301–318 (2018).
McLane, L. M., Abdel-Hakeem, M. S. & Wherry, E. J. CD8 T cell exhaustion during chronic viral infection and cancer. Annu. Rev. Immunol. 37, 457–495 (2019).
Gallimore, A. et al. Induction and exhaustion of lymphocytic choriomeningitis virus-specific cytotoxic T lymphocytes visualized using soluble tetrameric major histocompatibility complex class I-peptide complexes. J. Exp. Med. 187, 1383–1393 (1998).
Zajac, A. J. et al. Viral immune evasion due to persistence of activated T cells without effector function. J. Exp. Med. 188, 2205–2213 (1998).
Barber, D. L. et al. Restoring function in exhausted CD8 T cells during chronic viral infection. Nature 439, 682–687 (2006).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8+ T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
He, R. et al. Follicular CXCR5- expressing CD8+ T cells curtail chronic viral infection. Nature 537, 412–428 (2016).
Jadhav, R. R. et al. Epigenetic signature of PD-1+TCF1+CD8 T cells that act as resource cells during chronic viral infection and respond to PD-1 blockade. Proc. Natl Acad. Sci. USA 116, 14113–14118 (2019).
Zander, R. et al. CD4+ T cell help is required for the formation of a cytolytic CD8+ T cell subset that protects against chronic infection and cancer. Immunity 51, 1028–1042 (2019).
Hudson, W. H. et al. Proliferating transitory T cells with an effector-like transcriptional signature emerge from PD-1+ stem-like CD8+ T cells during chronic infection. Immunity 51, 1043–1058 (2019).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013 (2018).
Brummelman, J. et al. High-dimensional single cell analysis identifies stem-like cytotoxic CD8+ T cells infiltrating human tumors. J. Exp. Med. 215, 2520–2535 (2018).
Jansen, C. S. et al. An intra-tumoral niche maintains and differentiates stem-like CD8 T cells. Nature 576, 465–470 (2019).
Mann, T. H. & Kaech, S. M. Tick-TOX, it’s time for T cell exhaustion. Nat. Immunol. 20, 1092–1094 (2019).
Bhatt, K. H. et al. Profiling HPV-16-specific T cell responses reveals broad antigen reactivities in oropharyngeal cancer patients. J. Exp. Med. 217, e20200389 (2020).
Krishna, S. et al. Human papilloma virus specific immunogenicity and dysfunction of CD8+ T cells in head and neck cancer. Cancer Res. 78, 6159–6170 (2018).
Bobisse, S. et al. Sensitive and frequent identification of high avidity neo-epitope specific CD8+ T cells in immunotherapy-naive ovarian cancer. Nat. Commun. 9, 1092 (2018).
Wieland, A. et al. T cell receptor sequencing of activated CD8 T cells in the blood identifies tumor-infiltrating clones that expand after PD-1 therapy and radiation in a melanoma patient. Cancer Immunol. Immunother. 67, 1767–1776 (2018).
Simoni, Y. et al. Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Rosato, P. C. et al. Virus-specific memory T cells populate tumors and can be repurposed for tumor immunotherapy. Nat. Commun. 10, 567 (2019).
Gattinoni, L. et al. A human memory T cell subset with stem cell-like properties. Nat. Med. 17, 1290–1297 (2011).
Kamphorst, A. O. et al. Rescue of exhausted CD8 T cells by PD-1-targeted therapies is CD28-dependent. Science 355, 1423–1427 (2017).
Patel, J. J., Levy, D. A., Nguyen, S. A., Knochelmann, H. M. & Day, T. A. Impact of PD-L1 expression and human papillomavirus status in anti-PD1/PDL1 immunotherapy for head and neck squamous cell carcinoma—systematic review and meta-analysis. Head Neck 42, 774–786 (2020).
Skeate, J. G., Woodham, A. W., Einstein, M. H., Da Silva, D. M. & Kast, W. M. Current therapeutic vaccination and immunotherapy strategies for HPV-related diseases. Hum. Vaccines Immunother. 12, 1418–1429 (2016).
Ha, S. J. et al. Enhancing therapeutic vaccination by blocking PD-1-mediated inhibitory signals during chronic infection. J. Exp. Med. 205, 543–555 (2008).
de Martel, C., Georges, D., Bray, F., Ferlay, J. & Clifford, G. M. Global burden of cancer attributable to infections in 2018: a worldwide incidence analysis. Lancet Glob. Health 8, e180–e190 (2020).
Wieland, A. et al. Defining HPV-specific B cell responses in patients with head and neck cancer. Nature, https://doi.org/10.1038/s41586-020-2931-3 (2020).
NIH Tetramer Core Facility. Production Protocols: Class I MHC Tetramer Preparation https://tetramer.yerkes.emory.edu/support/protocols#10 (2006).
Vita, R. et al. The Immune Epitope Database (IEDB): 2018 update. Nucleic Acids Res. 47, D339–D343 (2019).
Sidney, J. et al. Measurement of MHC/peptide interactions by gel filtration or monoclonal antibody capture. Current Protoc. Immunol. 100, 18.3.1–18.3.36 (2013).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
DeTomaso, D. & Yosef, N. FastProject: a tool for low-dimensional analysis of single-cell RNA-seq data. BMC Bioinformatics 17, 315 (2016).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Acknowledgements
This work was supported by funding from the Ambrose Monell Foundation (R.A.); the T. J. Martell Foundation (R.A.); NIH grants 5U19AI057266 and P01AI056299 (R.A.); the Oliver S. and Jennie R. Donaldson Charitable Trust (R.A.); a Winship Invest$ Pilot grant (R.A., Z.G.C. and N.F.S.); Swiss National Science Foundation grant P300PB_174483 (C.S.E.); a Eugenio Litta Foundation grant (C.S.E.); NCI grant 1-R00-CA197891 (H.T.K.); the James M. Cox Foundation and James C. Kennedy (H.T.K.); the Prostate Cancer Foundation (H.T.K. and N.P.); and a Triological Society Research Career Development Award (M.R.P.). We thank H. Wu for technical assistance and M. Clayton for administrative support. We acknowledge the Emory Flow Cytometry Core supported by the National Center for Georgia Clinical and Translational Science Alliance of the NIH under award number UL1TR002378; the Integrated Cellular Imaging Microscopy Core of the Winship Cancer Institute of Emory University and the NIH–NCI under award number, 2P30CA138292-04; and the Yerkes NHP Genomics Core, which is supported in part by NIH P51 OD011132. This research was supported in part by the Intramural Research Program of the NIH, the Frederick National Laboratory for Cancer Research and the Center for Cancer Research. The project has been funded in part with federal funds from the Frederick National Laboratory for Cancer Research, under contract no. HHSN261200800001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products or organizations imply endorsement by the US Government.
Author information
Authors and Affiliations
Contributions
C.S.E. performed most of the experiments and analysed the data. H.T.K. analysed the scRNA-seq data. M.R.P. collected and provided human specimens, and analysed patient data. M.A.C. analysed scRNA-seq data. M.A.C., N.P. and A.W. performed in vitro proliferation experiments. R.C.O. performed and analysed multiplex immunohistochemistry experiments. T.H.N. performed flow cytometry experiments. C.C.G. and X.W., supervised by D.M.S., handled human specimens. M.C. performed HLA-typing analyses. J.S. and A.S. performed peptide–MHC affinity measurements. D.M.S., N.F.S. and Z.G.C. initiated the clinical specimen protocol. A.W. processed human specimens. A.W. and R.A. conceived, designed and supervised the project, and contributed equally to this work. C.S.E., H.T.K., A.W. and R.A. wrote the manuscript, with all authors contributing to the revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.A. holds patents related to the PD-1 inhibitory pathway. C.S.E., A.W. and R.A. are inventors on a patent application filed by Emory University relating to the use of HPV-specific TCR sequences. All other authors declare no competing interests.
Additional information
Peer review information Nature thanks Benny Chain, Evan Newell and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Analysis of PD-1+ CD8 TILs in patients with HPV-positive HNSCC.
a, Frequency of CD8 T cells among TILs and number of CD8+ TILs per gram of HPV-positive HNSCC primary tumours (pink) and metastatic lymph nodes (metLN, grey), with mean and SD. b, Representative flow plots showing Granzyme B, Ki-67, and CD39 expression among TCF-1+Tim-3− and Tim-3+TCF-1− CD8 T cell subsets in TILs. c, Frequency of Granzyme B+, Ki-67+ and CD39+ cells among PD-1+ CD8 TIL subsets, with means and SD. Two-sided Wilcoxon matched-pairs rank test. d, Geometric mean fluorescence intensity (MFI) of TOX in CD8 TIL subsets and patient-matched naive peripheral blood CD8 T cells (CD3+CD4−CD8+CCR7+CD45RA+), with means and SD. Friedman test with two-sided Dunn’s multiple comparisons test. e, f, Multiplex immunohistochemistry of primary tumour (e) and (f) metastatic lymph nodes (metLN) showing CD8+ (red) and PD-1+ (green) cells infiltrating the tumour parenchyma (cytokeratin+; blue) and stroma (cytokeratin negative; black areas highlighted by the orange dashed lines). TCF-1+ cells (white) are predominantly found in the stromal regions of the metastasis. Composite image on the right shows stem-like CD8 T cells (CD8+PD-1+TCF-1+; yellow membranous staining with white nuclear marker) within the stroma at a higher magnification of the area corresponding to the white rectangle. g, h, Frequency (g) and density (h) of PD-1+TCF-1− CD8 TILs in the tumour parenchyma and stroma, with mean and SD. n = 7 (five primary tumours (pink) and two metastatic lymph nodes (grey)). ns = not significant, two-sided paired t-tests.
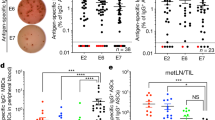
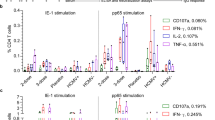
Extended Data Fig. 2 Mapping of HPV-specific CD8 T cell epitopes in patients with HNSCC.
a, Summary graph showing the numbers of positive peptides from HPV E2 (blue), E5 (green), E6 (red) and E7 (grey) proteins in expanded T cells of patients with HPV-positive HNSCC (n = 17) as measured by IFNγ ELISpot. b, Flow plots of expanded PBMC re-stimulated with the identified peptides and analysed by ICCS for IFNγ and TNFα. Gated on CD3+CD4−CD8+ cells. c, Flow plots of expanded PBMC stained with HPV-specific MHC-I tetramers. Gated on CD3+CD4−CD8+ cells. d, Table summarizing results of the HLA-binding assays (IC50) of identified HPV epitopes. Indicated in bold are epitopes for which ex vivo tetramer staining was performed.
Extended Data Fig. 3 Direct ex vivo tetramer staining of TILs and PBMCs from patients with HNSCC.
TILs from primary tumours and metastatic lymph nodes as well as matched PBMC samples were tetramer-stained ex vivo with HPV-specific tetramers. Flow plots showing tetramer+ CD8 T cells in primary tumours, metastatic LN and PBMC. The cells were stained with the following tetramers: a, A01/E2 151-159. b, A01/E2 329-337, c, A02/E5 46-54 and d, B35/E6 52-61. Gated on CD3+CD4−CD8+ cells.
Extended Data Fig. 4 MHC-I tetramer sorting and scRNA-seq analysis of HPV-specific PD-1+ CD8 T cells from primary tumour and metastatic lymph nodes.
a, b, Flow cytometry plots showing pre- and post-sorted tetramer-positive cells from primary and metastatic tumours. c–f, scRNA-seq data of the 13 tetramer-sorted samples showing the relative distribution among the three different clusters (stem-like, transitory and exhausted) in HPV-specific CD8 T cells in each sample.
Extended Data Fig. 5 Comparison of HPV-specific PD-1+ CD8 T cells in primary tumour and metastatic lymph nodes.
HPV tetramer-specific CD8 T cells from 13 samples including seven primary tumours and six metastatic lymph nodes were sorted and subjected to single cell RNAseq. a, UMAP clustering of HPV tetramer-specific CD8 T cells combined from all 13 samples, irrespective of tumour site, identified three distinct clusters; #1 stem-like, #2 transitory and #3 terminally differentiated/exhausted. b, Pairwise comparison of identified clusters among all HPV tetramer-specific CD8 T cells. Volcano plots show average fold-change by all cells in the cluster by -log10 p-value. The number of differentially expressed genes (≥0.25 Log2 fold-change in each pairwise comparison) are indicated in each plot. c, Heat map showing the top differentially expressed genes. The top 25 most significant genes from each cluster are shown. d, To assess gene expression differences of HPV-specific CD8 T cells in the two sites, we performed UMAP clustering of cells isolated from primary tumour and metastatic lymph node samples, respectively. We identified three clusters in primary tumours and four in metastatic lymph nodes. Gene expression was compared between the respective subsets (Green -> Green; Yellow -> Yellow; Blue -> Blue) in each tissue. Volcano plots highlight the differences between these clusters. The orange cluster identified in the metastatic lymph nodes consisted of very few cells and was thus not included in the comparisons. e, VISION analysis of HPV-specific CD8 T cells for enrichment of gene signatures associated with LCMV-specific stem-like and terminally differentiated/exhausted CD8 T cells. UMAP plots show the top quintile of cells enriched for the signature in blue. f, FACS analysis of various markers for HPV tetramer-specific intratumoral CD8 T cells. Plots are gated on PD-1+ HPV-specific CD8 T cells and show the respective marker versus TCF-1, the defining transcription factor of stem-like CD8 T cells. Summary plots show the frequency of TCF-1+ and TCF-1− cells expressing the respective marker for six patient samples.
Extended Data Fig. 6 Comparing the transcriptional program of total PD-1+ CD8 T cells with HPV-specific PD-1+ CD8 T cells in the tumour.
To compare total PD-1+ CD8 T cells with HPV-specific PD-1+ CD8 T cells in the TME, we performed scRNA-seq of total PD-1+ CD8 T cells (depleted of identified HPV-specific CD8 T cell reactivities) and used analysis techniques similar to those in Fig. 3. a, UMAP clustering of PD-1+ CD8 T cells. Cells from six samples (three patients, primary tumour and metastatic lymph nodes) were computationally combined, and four distinct clusters were identified. b, Distribution of individual samples among the four identified clusters. c, Comparison of gene expression differences between total PD-1+ CD8 T cells and HPV tetramer-positive CD8 T cells. The corresponding clusters of cells from HPV tetramer-specific CD8 T cells were compared to the cells found among total PD-1+ CD8 T cells. Clusters 1-3 mapped very closely to what was found in the HPV-specific cells, with relatively few differentially expressed genes. d, Comparison of cluster 4 in PD-1+ cells to clusters of HPV-specific cells. Cluster 4 was compared to the other clusters to identify specific gene differences. Volcano plots show fold change versus -log(p-value) for each gene. e, UMAP plots show selected genes that are significantly up- or down-regulated in this cluster versus others from the PD-1+ cells.
Extended Data Fig. 7 HPV-specific CD8 T cell clonotypes exist in multiple differentiation states.
a, UMAP clustering of HPV tetramer-specific CD8 T cells. b, TCR repertoire of tetramer-positive CD8 TILs in primary tumours (Prim.) and metastatic lymph nodes (Met.) with the top four clonotypes highlighted. Colours do not indicate the same clonotype between patients or epitope reactivities. c, TCR repertoire of tetramer-positive CD8 TILs in matched primary tumour and metastatic lymph node of the same patient with the top 4 clonotypes highlighted. Colours indicate the same clonotype within a patient and epitope reactivity. d, TCR frequency in patients with matched primary and metastatic tissue. e, TCR diversity of the identified clusters. f, Overlap between clusters for each patient as determined by Morisita Horn index. g, Distribution of the most frequent TCR clonotypes for each patient and epitope across gene expression clusters. UMAP plots show the distribution of the most frequent TCR clonotype across clusters. Accompanying bar charts showing the distribution of the eight most prominent TCR clonotypes across all clusters (clonotypes with less than 10 cells are not shown). The number of cells of the respective TCR clonotype is indicated below.
Extended Data Fig. 8 Pseudotime analysis to investigate the lineage relationship of HPV-specific PD-1+ CD8 T cell subsets.
a–c, Pseudotime analysis of HPV tetramer-positive CD8 T cells showing a differentiation trajectory where cells start as stem-like PD-1+TCF-1+ cells, transition through an intermediate stage, before taking on a terminally differentiated state. Selected genes (b) and enrichments for stem- and terminally differentiated gene signatures (c) are shown through this trajectory. d, Pseudotime analysis showing the distribution of the immunodominant clonotype of patients 7 and 51 in primary tumour and metastatic site through pseudotime.
Extended Data Fig. 9 Proliferation and differentiation potential of HPV-specific stem-like CD8 T cells.
a, Flow plots showing the gating strategy to isolate stem-like (CD39−Tim-3−) and terminally differentiated cells (CD39+Tim-3+) PD-1+CD45RA− CD8 T cells. Histograms showing TCF-1 expression to validate that the sorting strategy using CD39 and Tim-3 as surrogate markers enriches for stem-like (green) and terminally differentiated (blue) cells. b, Representative plots showing CTV dilution and expression of CD45RO, CD25 and CD28 after five days of culturing stem-like and terminally differentiated CD8 T cells alone or with peptide-pulsed PBMCs. Summary plots show percentage of cells positive for the indicated markers on day five. Means and their SD are represented, ** < 0.01, ns = not significant, unpaired Mann-Whitney U test.
Extended Data Fig. 10 Gating strategies.
a, Flow plots showing the gating strategy for bulk TIL staining and gating on live CD3+CD8+PD-1+ (shown in Fig. 1a) and Tim-3/TCF-1+ cells (shown in Fig. 1b–d). b, Flow plots showing the gating strategy for ICCS of expanded T cells measuring IFNγ and TNFα expression of live CD3+CD4−CD8+ cells (shown in Fig. 2b). c, Flow plots showing the sort gating strategy for ex vivo TIL staining (shown in Fig. 2c). Tetramer-positive CD8 T cells were gated as live CD3+ CD4− CD8+ double-tetramer+ cells.
Supplementary information
Supplementary Table 1
List of reagents.
Supplementary Table 2
Predicted HPV-peptides used for T cell expansion and epitope mapping.
Rights and permissions
About this article
Cite this article
Eberhardt, C.S., Kissick, H.T., Patel, M.R. et al. Functional HPV-specific PD-1+ stem-like CD8 T cells in head and neck cancer. Nature 597, 279–284 (2021). https://doi.org/10.1038/s41586-021-03862-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03862-z
This article is cited by
-
FOXP1 and KLF2 reciprocally regulate checkpoints of stem-like to effector transition in CAR T cells
Nature Immunology (2024)
-
Reusability report: Leveraging supervised learning to uncover phenotype-relevant biology from single-cell RNA sequencing data
Nature Machine Intelligence (2024)
-
Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics
Nature Communications (2024)
-
Semi-supervised integration of single-cell transcriptomics data
Nature Communications (2024)
-
PD-1 defines a distinct, functional, tissue-adapted state in Vδ1+ T cells with implications for cancer immunotherapy
Nature Cancer (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.