Abstract
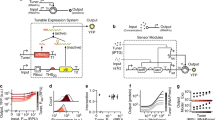
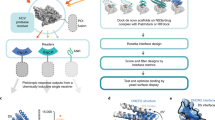
The construction and assembly of artificial allosteric protein switches into information and energy processing networks connected to both biological and non-biological systems is a central goal of synthetic biology and bionanotechnology. However, designing protein switches with the desired input, output and performance parameters is challenging. Here we use a range of reporter proteins to demonstrate that their chimeras with duplicated receptor domains produce YES gate protein switches with large (up to 9,000-fold) dynamic ranges and fast (minutes) response rates. In such switches, the epistatic interactions between largely independent synthetic allosteric sites result in an OFF state with minimal background noise. We used YES gate protein switches based on β-lactamase to develop quantitative biosensors of therapeutic drugs and protein biomarkers. Furthermore, we demonstrated the reconfiguration of YES gate switches into AND gate switches controlled by two different inputs, and their assembly into signalling networks regulated at multiple nodes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available in this paper and its supplementary information files. The following PDB entries were used in this study: BLA inhibitory protein I in complex with TEM-1 BLA (3gmw), trehalase (2jg0) and TEM-1 BLA in complex with inhibitor (1ERQ). Source data are provided with this paper.
References
Wodak, S. J. et al. Allostery in its many disguises: from theory to applications. Structure 27, 566–578 (2019).
Alberstein, R. G., Guo, A. B. & Kortemme, T. Design principles of protein switches. Curr. Opin. Struct. Biol. 72, 71–78 (2021).
Jackson, C., Anderson, A. & Alexandrov, K. The present and the future of protein biosensor engineering. Curr. Opin. Struct. Biol. 75, 102424 (2022).
Liu, G. Grand challenges in biosensors and biomolecular electronics. Front. Bioeng. Biotechnol. 9, 707615 (2021).
Merkx, M., Smith, B. & Jewett, M. Engineering sensor proteins. ACS Sens. 4, 3089–3091 (2019).
Masson, J.-F. & Pelletier, J. N. Will nanobiosensors change therapeutic drug monitoring? The case of methotrexate. Nanomedicine10, 521–524 (2015).
Katz, E. Enzyme‐Based Computing Systems (Wiley, 2019).
Stein, V. & Alexandrov, K. Protease-based synthetic sensing and signal amplification. Proc. Natl. Acad. Sci. USA 111, 15934–15939 (2014).
Fink, T. & Jerala, R. Designed protease-based signaling networks. Curr. Opin. Chem. Biol. 68, 102146 (2022).
Makhlynets, O. V., Raymond, E. A. & Korendovych, I. V. Design of allosterically regulated protein catalysts. Biochemistry 54, 1444–1456 (2015).
Nasu, Y., Shen, Y., Kramer, L. & Campbell, R. E. Structure- and mechanism-guided design of single fluorescent protein-based biosensors. Nat. Chem. Biol. 17, 509–518 (2021).
Clark, J. J., Benson, M. L., Smith, R. D. & Carlson, H. A. Inherent versus induced protein flexibility: comparisons within and between apo and holo structures. PLoS Comput. Biol. 15, e1006705 (2019).
Guo, Z. et al. Generalizable protein biosensors based on synthetic switch modules. J. Am. Chem. Soc. 141, 8128–8135 (2019).
Nadler, D. C., Morgan, S.-A., Flamholz, A., Kortright, K. E. & Savage, D. F. Rapid construction of metabolite biosensors using domain-insertion profiling. Nat. Commun. 7, 12266 (2016).
Guntas, G., Mansell, T. J., Kim, J. R. & Ostermeier, M. Directed evolution of protein switches and their application to the creation of ligand-binding proteins. Proc. Natl Acad. Sci. USA 102, 11224–11229 (2005).
Ergun Ayva, C. et al. Exploring performance parameters of artificial allosteric protein switches. J. Mol. Biol. 434, 167678 (2022).
Nishikawa, K. K., Hoppe, N., Smith, R., Bingman, C. & Raman, S. Epistasis shapes the fitness landscape of an allosteric specificity switch. Nat. Commun. 12, 5562 (2021).
Starr, T. N. & Thornton, J. W. Epistasis in protein evolution. Protein Sci. 25, 1204–1218 (2016).
Motlagh, H. N., Wrabl, J. O., Li, J. & Hilser, V. J. The ensemble nature of allostery. Nature 508, 331–339 (2014).
Gruber, R. & Horovitz, A. Unpicking allosteric mechanisms of homo-oligomeric proteins by determining their successive ligand binding constants. Phil. Trans. R. Soc. B 373, 20170176 (2018).
Aroul-Selvam, R., Hubbard, T. & Sasidharan, R. Domain insertions in protein structures. J. Mol. Biol. 338, 633–641 (2004).
Salverda, M. L. M., De Visser, J. A. G. M. & Barlow, M. Natural evolution of TEM-1 β-lactamase: experimental reconstruction and clinical relevance. FEMS Microbiol. Rev. 34, 1015–1036 (2010).
Guo, Z. et al. Design of a methotrexate-controlled chemical dimerization system and its use in bio-electronic devices. Nat. Commun. 12, 7137 (2021).
Rochelet, M. et al. Amperometric detection of extended-spectrum β-lactamase activity: application to the characterization of resistant E. coli strains. Analyst 140, 3551–3556 (2015).
Banaszynski, L. A., Liu, C. W. & Wandless, T. J. Characterization of the FKBP·rapamycin·FRB ternary complex. J. Am. Chem. Soc. 127, 4715–4721 (2005).
Gräwe, A. & Merkx, M. Bioluminescence goes dark: boosting the performance of bioluminescent sensor proteins using complementation inhibitors. ACS Sens. 7, 3800–3808 (2022).
Dincer, C., Bruch, R., Kling, A., Dittrich, P. S. & Urban, G. A. Multiplexed point-of-care testing —xPOCT. Trends Biotechnol. 35, 728–742 (2017).
Acknowledgements
We thank C. Pretorius from Pathology Queensland for carrying out the quantification of biomarkers in human samples for this study. We thank D. Massana Roquero from Stanford University School of Medicine for help with experiments related to alginate gel. We also thank I. Berezovsky from A-STAR Singapore and S. Fleishman from Weizmann Institute for stimulating discussions. This work was supported in part by the Australian Research Council Discovery Projects (DP160100973 and DP150100936), as well as ARC Centre of Excellence in Synthetic Biology (CE200100029 to K.A.). The work was also supported by National Health and Medical Research Council Development Grants (APP1113262 to K.A. and APP1179001 to K.A. and J.P.J.U.). This work was also in part supported by the Human Frontier Science Program (grant RGP0002/2018 to O.S., K.A. and E.K.) and the US Department of Defense (grant W81XWH-20-1-0708 to K.A. and E.K.). K.A. gratefully acknowledges the financial support of the Synthetic Biology Alliance between the Commonwealth Scientific and Industrial Research Organisation (CSIRO) and Queensland University of Technology.
Author information
Authors and Affiliations
Contributions
Z.G. designed experiments, performed experiments, analysed data and contributed to writing the paper. O.S. designed experiments, performed experiments, analysed data and contributed to writing the paper. C.E.A. designed experiments, performed experiments, analysed data and contributed to writing the paper. P.W. performed experiments. J.P. designed experiments, analysed data and contributed to writing the paper. J.W. designed experiments, performed experiments, analysed data and contributed to writing the paper. C.E.V. designed experiments, performed experiments, analysed data and contributed to writing the paper. J.P.J.U. designed experiments and analysed data. E.K. designed experiments, analysed data and contributed to writing the paper. K.A. designed experiments, analysed data and contributed to writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: Z.G. and K.A. are named inventors on PCT patent application PCT/IB2023/020001 entitled ‘AND-gate protein-based switches’ filed by Queensland University of Technology that covers the YES biosensor architecture used in this study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Tables 1–6.
Supplementary Data 1
Data used to generate plots and graphs in Supplementary Figs. 1–10. Unprocessed SDS–PAGE gels used to generate Fig. 3i.
Source data
Source Data Figs. 1–4
Data used to generate plots and graphs in Figs. 1–4.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, Z., Smutok, O., Ayva, C.E. et al. Development of epistatic YES and AND protein logic gates and their assembly into signalling cascades. Nat. Nanotechnol. 18, 1327–1334 (2023). https://doi.org/10.1038/s41565-023-01450-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-023-01450-y