Abstract
The human facilitates chromatin transcription (FACT) complex is a chromatin remodeller composed of human suppressor of Ty 16 homologue (hSpt16) and structure-specific recognition protein-1 subunits that regulates cellular gene expression. Whether FACT regulates host responses to infection remained unclear. We identify a FACT-mediated, interferon-independent, antiviral pathway that restricts poxvirus replication. Cell culture and bioinformatics approaches suggest that early viral gene expression triggers nuclear accumulation of SUMOylated hSpt16 subunits required for the expression of E26 transformation-specific sequence-1 (ETS-1)—a transcription factor that activates virus restriction programs. However, biochemical studies show that poxvirus-encoded A51R proteins block ETS-1 expression by outcompeting structure-specific recognition protein-1 binding to SUMOylated hSpt16 and by tethering SUMOylated hSpt16 to microtubules. Furthermore, A51R antagonism of FACT enhances poxvirus replication in human cells and virulence in mice. Finally, we show that FACT also restricts rhabdoviruses, flaviviruses and orthomyxoviruses, suggesting broad roles for FACT in antiviral immunity. Our study reveals the FACT–ETS-1 antiviral response (FEAR) pathway to be critical for eukaryotic antiviral immunity and describes a unique mechanism of viral immune evasion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that the main data supporting the findings of this study are available within the article, its source data and/or its Supplementary Information. RNA-seq datasets are available under Gene Expression Omnibus accession GSE185829. The human genome assembly (hg38/GRCh38) is available under National Center for Biotechnology Information accession GCF_000001405. Source data are provided with this paper.
Code availability
Custom code used for analysis of the RNA-seq data is available in the Supplementary Information.
References
Nan, Y., Nan, G. & Zhang, Y. J. Interferon induction by RNA viruses and antagonism by viral pathogens. Viruses 6, 4999–5027 (2014).
Beachboard, D. C. & Horner, S. M. Innate immune evasion strategies of DNA and RNA viruses. Curr. Opin. Microbiol. 32, 113–119 (2016).
Reus, J. B., Rex, E. A. & Gammon, D. B. How to inhibit nuclear factor-kappa B signaling: lessons from poxviruses. Pathogens 11, 1061 (2022).
Yu, H., Bruneau, R. C., Brennan, G. & Rothenburg, S. Battle Royale: innate recognition of poxviruses and viral immune evasion. Biomedicines 9, 765 (2021).
Gammon, D. B. et al. A single vertebrate DNA virus protein disarms invertebrate immunity to RNA virus infection. eLife 3, e02910 (2014).
Orphanides, G., Wu, W. H., Lane, W. S., Hampsey, M. & Reinberg, D. The chromatin-specific transcription elongation factor FACT comprises human SPT16 and SSRP1 proteins. Nature 400, 284–288 (1999).
Orphanides, G., LeRoy, G., Chang, C. H., Luse, D. S. & Reinberg, D. FACT, a factor that facilitates transcript elongation through nucleosomes. Cell 92, 105–116 (1998).
Safina, A. et al. Complex mutual regulation of facilitates chromatin transcription (FACT) subunits on both mRNA and protein levels in human cells. Cell Cycle 12, 2423–2434 (2013).
Gammon, D. B. & Evans, D. H. The 3′-to-5′ exonuclease activity of vaccinia virus DNA polymerase is essential and plays a role in promoting virus genetic recombination. J. Virol. 83, 4236–4250 (2009).
Gammon, D. B. et al. Vaccinia virus-encoded ribonucleotide reductase subunits are differentially required for replication and pathogenesis. PLoS Pathog. 6, e1000984 (2010).
Brown, E., Senkevich, T. G. & Moss, B. Vaccinia virus F9 virion membrane protein is required for entry but not virus assembly, in contrast to the related L1 protein. J. Virol. 80, 9455–9464 (2006).
Griffiths, G., Roos, N., Schleich, S. & Locker, J. K. Structure and assembly of intracellular mature vaccinia virus: thin-section analyses. J. Virol. 75, 11056–11070 (2001).
Jordan, I. et al. A deleted deletion site in a new vector strain and exceptional genomic stability of plaque-purified modified vaccinia ankara (MVA). Virol. Sin. 35, 212–226 (2020).
Lu, Q. et al. Homeostatic control of innate lung inflammation by Vici syndrome gene Epg5 and additional autophagy genes promotes influenza pathogenesis. Cell Host Microbe 19, 102–113 (2016).
Schoggins, J. W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 (2011).
Ohlson, M. B. et al. Genome-scale CRISPR screening reveals host factors required for ribosome formation and viral replication. mBio 14, e0012723 (2023).
Dembowski, J. A. & DeLuca, N. A. Selective recruitment of nuclear factors to productively replicating herpes simplex virus genomes. PLoS Pathog. 11, e1004939 (2015).
Fox, H. L., Dembowski, J. A. & DeLuca, N. A. A herpesviral immediate early protein promotes transcription elongation of viral transcripts. mBio 8, e00745-17 (2017).
Gasparian, A. V. et al. Curaxins: anticancer compounds that simultaneously suppress NF-κB and activate p53 by targeting FACT. Sci. Transl. Med. 3, 95ra74 (2011).
Suzawa, M. et al. A gene-expression screen identifies a non-toxic sumoylation inhibitor that mimics SUMO-less human LRH-1 in liver. eLife 4, e09003 (2015).
Zhao, Q. et al. GPS-SUMO: a tool for the prediction of sumoylation sites and SUMO-interaction motifs. Nucleic Acids Res. 42, W325–W330 (2014).
Li, S. J. & Hochstrasser, M. A new protease required for cell-cycle progression in yeast. Nature 398, 246–251 (1999).
Winkler, D. D. & Luger, K. The histone chaperone FACT: structural insights and mechanisms for nucleosome reorganization. J. Biol. Chem. 286, 18369–18374 (2011).
Seo, D. et al. Poxvirus A51R proteins regulate microtubule stability and antagonize a cell-intrinsic antiviral response. Cell Rep. 43, 113882 (2024).
Dehmelt, L. & Halpain, S. The MAP2/Tau family of microtubule-associated proteins. Genome Biol. 6, 204 (2005).
Brameier, M., Krings, A. & MacCallum, R. M. NucPred—predicting nuclear localization of proteins. Bioinformatics 23, 1159–1160 (2007).
Kosugi, S., Hasebe, M., Tomita, M. & Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl Acad. Sci. USA 106, 10171–10176 (2009).
Yang, Z. et al. Deciphering poxvirus gene expression by RNA sequencing and ribosome profiling. J. Virol. 89, 6874–6886 (2015).
Pavri, R. et al. Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125, 703–717 (2006).
Fleming, A. B., Kao, C. F., Hillyer, C., Pikaart, M. & Osley, M. A. H2B ubiquitylation plays a role in nucleosome dynamics during transcription elongation. Mol. Cell 31, 57–66 (2008).
Minsky, N. et al. Monoubiquitinated H2B is associated with the transcribed region of highly expressed genes in human cells. Nat. Cell Biol. 10, 483–488 (2008).
Sanda, C. et al. Differential gene induction by type I and type II interferons and their combination. J. Interferon Cytokine Res. 26, 462–472 (2006).
Goulet, M. L. et al. Systems analysis of a RIG-I agonist inducing broad spectrum inhibition of virus infectivity. PLoS Pathog. 9, e1003298 (2013).
Garrett-Sinha, L. A. Review of Ets1 structure, function, and roles in immunity. Cell. Mol. Life Sci. 70, 3375–3390 (2013).
Schoggins, J. W. Interferon-stimulated genes: what do they all do? Annu. Rev. Virol. 6, 567–584 (2019).
Wust, S., Schad, P., Burkart, S. & Binder, M. Comparative analysis of six IRF family members in alveolar epithelial cell-intrinsic antiviral responses. Cells 10, 2600 (2021).
Urban, C. et al. Persistent innate immune stimulation results in IRF3-mediated but caspase-independent cytostasis. Viruses 12, 635 (2020).
Kolundzic, E. et al. FACT sets a barrier for cell fate reprogramming in Caenorhabditis elegans and human cells. Dev. Cell 46, 611–626.e12 (2018).
Deng, L. et al. Suppression of NF-κB activity: a viral immune evasion mechanism. Viruses 10, 409 (2018).
Shaw, A. E. et al. Fundamental properties of the mammalian innate immune system revealed by multispecies comparison of type I interferon responses. PLoS Biol. 15, e2004086 (2017).
Smale, S. T. Transcriptional regulation in the innate immune system. Curr. Opin. Immunol. 24, 51–57 (2012).
Teng, Y. et al. Expression of ETS1 in gastric epithelial cells positively regulate inflammatory response in Helicobacter pylori-associated gastritis. Cell Death Dis. 11, 498 (2020).
Winkler, D. D., Muthurajan, U. M., Hieb, A. R. & Luger, K. Histone chaperone FACT coordinates nucleosome interaction through multiple synergistic binding events. J. Biol. Chem. 286, 41883–41892 (2011).
Hanners, N. W. et al. Western Zika virus in human fetal neural progenitors persists long term with partial cytopathic and limited immunogenic effects. Cell Rep. 15, 2315–2322 (2016).
Ramakrishnan, M. A. Determination of 50% endpoint titer using a simple formula. World J. Virol. 5, 85–86 (2016).
Aboulaich, N. Lentivirus production. Bio-protocol Exchange https://doi.org/10.21769/BioProtoc.39 (2011).
Lourenco, K. L., Chinalia, L. A., Henriques, L. R., Rodrigues, R. A. L. & da Fonseca, F. G. Zoonotic vaccinia virus strains belonging to different genetic clades exhibit immunomodulation abilities that are proportional to their virulence. Virol. J. 18, 124 (2021).
Fromont-Racine, M., Rain, J. C. & Legrain, P. Toward a functional analysis of the yeast genome through exhaustive two-hybrid screens. Nat. Genet. 16, 277–282 (1997).
Bartel, P. L. & Fields, S. Analyzing protein–protein interactions using two-hybrid system. Methods Enzymol. 254, 241–263 (1995).
Kulesh, D. A. et al. Smallpox and pan-orthopox virus detection by real-time 3′-minor groove binder TaqMan assays on the roche LightCycler and the Cepheid smart Cycler platforms. J. Clin. Microbiol. 42, 601–609 (2004).
Wang, T. et al. The histone chaperone FACT modulates nucleosome structure by tethering its components. Life Sci. Alliance 1, e201800107 (2018).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2004).
Liu, Y. et al. FACT caught in the act of manipulating the nucleosome. Nature 577, 426–431 (2020).
Kesten, C., Schneider, R. & Persson, S. In vitro microtubule binding assay and dissociation constant estimation. Bio Protoc. 6, e1759 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
McKinney, W. Data structures for statistical computing in Python. In Proc. 9th Python in Science Conference 51–56 (2010).
Mi, H., Muruganujan, A. & Thomas, P. D. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 41, D377–D386 (2013).
Binns, D. et al. QuickGO: a web-based tool for Gene Ontology searching. Bioinformatics 25, 3045–3046 (2009).
Moutaftsi, M. et al. Correlates of protection efficacy induced by vaccinia virus-specific CD8+ T-cell epitopes in the murine intranasal challenge model. Eur. J. Immunol. 39, 717–722 (2009).
Mahajan, R., Delphin, C., Guan, T., Gerace, L. & Melchior, F. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell 88, 97–107 (1997).
Acknowledgements
This work was supported by: NIH grants 1R21AI144203-01 and 1R35GM137978-01 and Welch Foundation grant I-2062-20210327 (to D.B.G.); NIH grant 1R35GM142689-01 and Cancer Prevention and Research Institute of Texas grant RR 170047 (to D.C.H.); funding from the INC TV Initiative (to F.G.F.); and NIH Training Grant T32 AI007520 (to C.P., E.A.R. and A.E.). We thank the University of Texas Southwestern Medical Center Electron Microscopy Core Facility, funded by NIH grant 1S10OD021685-01A1. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We thank D. Evans (University of Alberta) for the mouse anti-VV F4L and I3L antibodies. We thank C. Mello (University of Massachusetts Medical School) for the ΔA51R, ΔA51RFA51R and VSV-GFP virus strains, as well as the Flag-VV A51R, CPXV FA51R, ECTV FA51R and YLDV FA51R pCDNA3 plasmids. We also thank J. Schoggins (University of Texas Southwestern Medical Center) for the YFV-17D-Venus and HSV-1-GFP virus strains and the MDCK cells. We thank H. W. Virgin (University of Washington School of Medicine) for the IAV strain A/PR/8/34. We thank M. Binder (German Cancer Research Center) for the non-targeting, ΔIRF3 and ΔIFNAR A549 cells. Finally, we thank B. Arif (Natural Resources Canada) for the provision of LD652 cells.
Author information
Authors and Affiliations
Contributions
D.B.G. and E.A.R. conceived of the experiment, devised the methodology, curated the data, wrote the original draft of the manuscript and visualized the data. D.B.G., E.A.R., D.S., S.C., C.P., S.B.O., A.E., D.H., Y.L. and M.M. performed the investigation. D.B.G., E.A.R., K.L., N.M.A., F.G.F., R.O. and D.C.H. reviewed and edited the manuscript. D.B.G., K.L., N.M.A., F.G.F., R.O. and D.C.H. supervised the experiments. D.B.G., F.G.F. and D.C.H. acquired the funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Guanqun Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The ΔA51R strain exhibits normal viral gene expression and genome replication but displays defective virion assembly in the presence of hSpt16.
a, IB of early (F4L, I3L) and late (F9L, A27L) in A549 WCE at indicated times post-infection (MOI = 10). IB is representative of at least two independent experiments. b, VV genome quantification by qPCR in A549 cultures at indicated times post-infection (MOI = 10). Data are means ± SD for n = 3 experiments. Statistical significance was determined by unpaired two-tailed Student’s t-test comparing genome copy number between ΔA51RFA51R and ΔA51R treatments within each time point. Only statistical comparisons with P < 0.05 are shown. c,d Analysis of total mature virus (MV) and immature virus (IV) frequencies (expressed as a percentage of total particles counted) using electron microscopy in A549 cells 24 hpi (MOI = 10) under scrambled (scram.) or hSpt16 RNAi conditions. c, Representative images of MV particles and various IV forms (crescents, spheres, spheres with nucleoid DNA) typically found in ΔA51RFA51R (top image) and ΔA51R-infected (bottom image) cells, respectively under control RNAi conditions. Selected examples of different types of MV and IV particles are indicated. Scale bar, 800 nm. d, Percentage of MV particles counted among total (MV + IV particles) particles in cells counted from each infection treatment. Data are means ± SD for n = 3 biologically independent experiments.
Extended Data Fig. 2 Conservation of Spt16 SUMOylation among eukaryotes.
a-c, IB of endogenous Spt16 proteins from indicated human (a) and mammalian animal cell lines (b) and from invertebrate cell lines and animal tissues (C. elegans) (c) showing SUMOylated (upper bands) and non-SUMOylated (lower bands) Spt16 forms. Non-human cell lines derive from the following: BSC-40 (Chlorocebus sabaeus); BHK-21 (Mesocricetus auratus); L929 (Mus musculus); SIRC (Oryctolagus cuniculus); R06E (Rousettus aegyptiacus); and LD652 (Lymantria dispar). Images in a-c are representative of at least two independent experiments.
Extended Data Fig. 3 VV A51R associates with microtubules (MTs) and tethers hSpt16SUMO to MTs.
a, Alignment of VV A51R with Proline Rich (PR) Region and Region 1 of human Tau40 MT-binding domains (MTBD)25. “KXGS” motif conserved in all four MTBDs in Tau is boxed. Residues in red indicate those converted to alanine in the triple A51R mutant. b, immunofluorescence staining of U2OS cells transfected with wild-type or triple mutant FA51R expression constructs after 24 h. Where indicated, 40 μM nocodazole was added to depolymerize MTs 6 h post-transfection MTs. Scale bar, 5 μm. c, Overview of in vitro MT co-sedimentation assay. d-e, Stain-free protein gels showing in vitro MT co-sedimentation assays with purified wild-type and mutant His-A51R. S, supernatant; P, pellet. f, Co-IP of transfected wild-type and mutant FA51R constructs with hSpt16 and tubulin in 293 T WCE. g, Overview of in vitro Flag-hSpt16 Co-IP with tubulin in the absence/presence of His-A51R. h, in vitro Co-IP of purified Flag-hSpt16, His-A51R and tubulin. Images in b, d-f, and h are representative of at least two independent experiments. Images in c and g were created with biorender.com.
Extended Data Fig. 4 VV A51R does not affect total hSpt16 protein levels.
a, Representative IB of hSpt16 in A549 WCE at indicated times post-infection (MOI = 10). b-c, Densitometric quantification of IB experiments represented in a for total hSpt16SUMO (b) or non-SUMOylated hSpt16 (c) protein levels plotted as fold change relative to levels in mock-infected WCE at each respective time point. Data in b-c are means ± SD for n = 3 biologically independent experiments. Statistical significance was determined by unpaired two-tailed Student’s t-test comparing hSpt16/hSpt16SUMO levels in infected versus mock-infected WCE at each time point. However, no significant differences were detected at any time point in either cell type (not shown). d, Representative IB of hSpt16 in U2OS WCE at indicated times post-infection (MOI = 10). e-f, Densitometric quantification of IB experiments represented in d for total hSpt16SUMO (e) or non-SUMOylated hSpt16 (f) protein levels plotted as fold change relative to levels in mock-infected WCE at each respective time point. Data in e-f are means ± SD for n = 3 biologically independent experiments. Statistical significance was determined by unpaired two-tailed Student’s t-test comparing hSpt16/hSpt16SUMO levels in infected versus mock-infected WCE at each time point. However, no significant differences were detected at any time point in either cell type (not shown). g, Quantification of fractionation experiments of U2OS cells mock- or ΔA51R-infected for 18 h performed as in Fig. 3h,i. Where indicated, proteasome inhibitor [bortezomib, (5 nM)] was added to media 2 hpi. Data are means ± SD; n = 3 biologically independent experiments.
Extended Data Fig. 5 hSpt16 SUMOylation does not affect histone or nucleosome interactions in vitro.
a, in vitro nickel bead pulldown of His-tagged H3/H4 and H2A/H2B complexes with purified Flag-hSpt16. b, in vitro FACT binding assay with reconstituted (H3-H4)2 tetrasomes or (H2A-H2B)-(H3-H4)2 hexasomes50. Where indicated, purified FACT complexes were treated with ULP-1 to remove hSpt16 SUMOylation prior to being added to binding assays. IBs of FACT treatments are shown below indicating hSpt16 and SSRP1 levels. Images in a,b are representative of two independent experiments.
Extended Data Fig. 6 ETS-1 induction by VV infection occurs by 4 hpi and requires early VV gene expression but not viral DNA replication.
a, Representative IB of ETS-1 levels in WCE from A549 cells infected with indicated strains for indicated times (MOI = 10). b, Densitometric quantification of n = 3 biologically independent IB experiments represented in a. Fold change in ETS-1 protein levels are plotted relative to levels in mock-infected WCE harvested 1 hpi. Data are means ± SD. Statistical significance was determined by unpaired two-tailed Student’s t-test. Only statistical comparisons where ETS-1 levels in infected WCE were significantly different (P < 0.05) from levels in mock-infected WCE are shown. c, IB of ETS-1 in WCE from A549 cells under indicated infection conditions 4 hpi. Where indicated, inoculum was heat-inactivated prior to infection to prevent VV gene expression. d, IB of ETS-1 in WCE from A549 cells under indicated infection conditions. Where indicated, AraC was added to media 1 hpi. VV A27L is a late VV protein, and its expression requires VV DNA replication and thus is inhibited by AraC treatment. Images in c-d are representative of two independent experiments.
Extended Data Fig. 7 The Type I IFN and FACT-ETS-1-depedent antiviral responses are independent pathways.
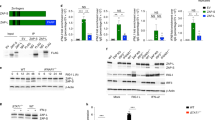
a-d, Representative IB (a,c) and quantification (b,d) of ETS-1 expression using IB of WCE from control or IRF3 knockout A549 cells infected with the indicated strains for 4 h (a,b) or 8 h (c,d). e-h, Representative IB (e,g) and quantification (f,h) of ETS-1 expression using IB of WCE from control or IFNAR knockout A549 cells infected with the indicated strains for 4 h (e,f) or 8 h (g,h). i-l, Representative IB (i,k) and quantification (j,l) of ETS-1 expression using IB of WCE from control or STAT1 knockout A549 cells infected with the indicated strains for 4 h (i,j) or 8 h (k,l). m, VV titers 48 hpi (MOI = 0.01) in IRF3 knockout A549 cells after treatment with indicated RNAi. Scram., scrambled. n-o, Representative IB (n) and quantification (o) of ETS-1 expression using IB of WCE from A549 cells treated with indicated doses of recombinant IFN-β for 24 h. IB of two ISGs (IFIT3 and STAT1) is also shown to illustrate ISG expression increases after IFN-β treatment. p-s, Representative IB (p,r) and quantification (q,s) of IFIT3 (ISG) expression using IB of WCE from A549 cells treated with scrambled or ETS-1 RNAi and then mock- or IFN-β (500 U/mL)-treated for 8 h (p-q) or 24 h (r-s). Data in b,d,f,h,j,l,m,o,q,s are means ± SD for n = 3 biologically independent experiments. Statistical significance was determined by unpaired two-tailed Student’s t-test. For brevity, only the most relevant statistical comparisons are shown. ns, not significant.
Extended Data Fig. 8 A51R-hSpt16SUMO interaction suppresses ETS-1 expression and promotes VV virulence in BALB/c mice.
a, Representative IB of A549 WCE from cells infected with indicated VV (MOI = 10) for 4 h. b, Densitometric quantification of ETS-1 protein levels from a. Data are means ± SD; n = 3 biologically independent experiments. c, VV titers 48 hpi (MOI = 0.01) in A549 cell cultures transfected with indicated RNAi treatments. d, Percent change in body weight of BALB/c male mice still alive at each time point (n = 20 animals over two independent experiments) after intranasal VV infection (102 PFU/animal). Data are means ± SE. e, Percent survival of BALB/c mice from d over time. f, Tissue viral loads in BALB/c mice (n = 5 animals) challenged as in d 8 dpi. Horizontal lines indicate means. g, qRT-PCR analysis of relative Ets1 mRNA levels in lung tissues from BALB/c animals (n = 5) infected with 102 PFU at 8 dpi. Ets1 mRNA levels are plotted as relative to levels in mock (PBS)-treated animals. Horizontal lines indicate means. ns, not significant. Image in a is representative of at three independent experiments. Statistical significance was determined by unpaired two-tailed Student’s t-test in b and c, by Log-Rank (Mantel-Cox) tests in e, and by unpaired two-tailed Mann-Whitney U tests in f and g. Only the most relevant statistical comparisons with P < 0.05 are shown for brevity.
Extended Data Fig. 9 Additional analyses of ΔA51RFA51R158-162Ala mutant infection in C57BL/6 mice.
a, Percent change in body weight of C57BL/6 male mice (n = 15 animals over two independent experiments) still alive at each time point after intranasal VV infection (105 PFU/animal). Data are means ± SE. b, Percent survival of C57BL/6 mice from a over time. c, Tissue viral loads from C57BL/6 mice (n = 5 animals) after intranasal challenge with 102 PFU at 8 dpi. Horizontal lines indicate means. Statistical significance was determined by Log-Rank (Mantel-Cox) tests in b and by unpaired two-tailed Mann-Whitney U tests and in c.
Extended Data Fig. 10 Model for FACT-ETS-1 Antiviral Response (FEAR) pathway.
VV early gene expression triggers hSpt16SUMO nuclear accumulation and formation of FACT complexes containing hSpt16SUMO that license FACT to associate with H2BK120ub sites in transcriptionally active chromatin and activate ETS-1 expression. Elevation of ETS-1 expression levels to a “critical threshold” results in ETS-1-dependent antiviral gene expression and VV restriction. VV A51R is expressed early during infection5 and inhibits FEAR pathway by tethering hSpt16SUMO to MTs and blocking its nuclear import. A51R may also prevent FACT complex formation in the cytosol by outcompeting SSRP1 for hSpt16SUMO binding, which would also serve to prevent nuclear accumulation of hSpt16SUMO-containing FACT complexes (not shown). Note: our fraction data in Fig. 3b-c, indicate that there are constant pools of hSpt16SUMO in the nucleus in uninfected cells (not shown) and thus we hypothesize that VV infection triggers hSpt16SUMO nuclear translocation from cytosolic pools to reach a critical level of hSpt16SUMO-containing FACT complexes in the nucleus required to induce ETS-1 levels to a critical threshold required for antiviral gene expression. Image created with biorender.com.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 (with associated source data), source code and Supplementary Results.
Supplementary Tables
Supplementary Tables 1–4.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Unprocessed western blots/gels.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Representative microscopy images.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed western blots/gels.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Representative microscopy images.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Representative microscopy images.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Unprocessed western blots/gels.
Source Data Extended Data Fig. 3
Representative microscopy images.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots/gels.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rex, E.A., Seo, D., Chappidi, S. et al. FEAR antiviral response pathway is independent of interferons and countered by poxvirus proteins. Nat Microbiol 9, 988–1006 (2024). https://doi.org/10.1038/s41564-024-01646-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-024-01646-5
This article is cited by
-
Primal FEAR protects against infection
Nature Microbiology (2024)