Abstract
Although autophagy sequesters Mycobacterium tuberculosis (Mtb) in in vitro cultured macrophages, loss of autophagy in macrophages in vivo does not result in susceptibility to a standard low-dose Mtb infection until late during infection, leaving open questions regarding the protective role of autophagy during Mtb infection. Here we report that loss of autophagy in lung macrophages and dendritic cells results in acute susceptibility of mice to high-dose Mtb infection, a model mimicking active tuberculosis. Rather than observing a role for autophagy in controlling Mtb replication in macrophages, we find that autophagy suppresses macrophage responses to Mtb that otherwise result in accumulation of myeloid-derived suppressor cells and subsequent defects in T cell responses. Our finding that the pathogen-plus-susceptibility gene interaction is dependent on dose has important implications both for understanding how Mtb infections in humans lead to a spectrum of outcomes and for the potential use of autophagy modulators in clinical medicine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
scRNA-seq data have been deposited to the NCBI Gene Expression Omnibus (GEO) database and are accessible through accession number GSE201410. RNA-seq data have been deposited to the NCBI GEO database and are accessible through accession number GSE245206. The raw data are provided in the Supplementary Table 1. The method used for analysing sequencing data is detailed in Methods. Source data are provided with this paper.
Code availability
Standard pipelines were used for the analysis of bulk and scRNA-seq datasets, as indicated in Methods. No custom code was developed for the analysis of data presented in this manuscript.
References
Watson, R. O., Manzanillo, P. S. & Cox, J. S. Extracellular M. tuberculosis DNA targets bacteria for autophagy by activating the host DNA-sensing pathway. Cell 150, 803–815 (2012).
Gutierrez, M. G. et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119, 753–766 (2004).
Castillo, E. F. et al. Autophagy protects against active tuberculosis by suppressing bacterial burden and inflammation. Proc. Natl Acad. Sci. USA 109, E3168–E3176 (2012).
Aylan, B. et al. ATG7 and ATG14 restrict cytosolic and phagosomal Mycobacterium tuberculosis replication in human macrophages. Nat. Microbiol. 8, 803–818 (2023).
Golovkine, G. R. Autophagy restricts Mycobacterium tuberculosis during acute infection in mice. Nat. Microbiol. 8, 819–832 (2023).
Kimmey, J. M. et al. Unique role for ATG5 in neutrophil-mediated immunopathology during M. tuberculosis infection. Nature 528, 565–569 (2015).
Kinsella, R. L. et al. Autophagy prevents early proinflammatory responses and neutrophil recruitment during Mycobacterium tuberculosis infection without affecting pathogen burden in macrophages. PLoS Biol. 21, e3002159 (2023).
Wang, F. et al. ATG5 provides host protection acting as a switch in the atg8ylation cascade between autophagy and secretion. Dev. Cell 58, 866–884.e8 (2023).
Pai, M. et al. Tuberculosis. Nat. Rev. Dis. Prim. 2, 16076 (2016).
Esaulova, E. et al. The immune landscape in tuberculosis reveals populations linked to disease and latency. Cell Host Microbe 29, 165–178.e8 (2021).
Plumlee, C. R. et al. Ultra-low dose aerosol infection of mice with Mycobacterium tuberculosis more closely models human tuberculosis. Cell Host Microbe 29, 68–82.e5 (2021).
Moreira-Teixeira, L. et al. Mouse transcriptome reveals potential signatures of protection and pathogenesis in human tuberculosis. Nat. Immunol. 21, 464–476 (2020).
Scott, H. M. & Flynn, J. L. Mycobacterium tuberculosis in chemokine receptor 2-deficient mice: influence of dose on disease progression. Infect. Immun. 70, 5946–5954 (2002).
Zak, D. E. et al. A blood RNA signature for tuberculosis disease risk: a prospective cohort study. Lancet 387, 2312–2322 (2016).
Berry, M. P. R. et al. An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis. Nature 466, 973–977 (2010).
Wang, Y.-T. et al. Select autophagy genes maintain quiescence of tissue-resident macrophages and increase susceptibility to Listeria monocytogenes. Nat. Microbiol. 5, 272–281 (2020).
Choi, J. et al. The parasitophorous vacuole membrane of Toxoplasma gondii is targeted for disruption by ubiquitin-like conjugation systems of autophagy. Immunity 40, 924–935 (2014).
Martinez, J. et al. Molecular characterization of LC3-associated phagocytosis reveals distinct roles for Rubicon, NOX2 and autophagy proteins. Nat. Cell Biol. 17, 893–906 (2015).
Upadhyay, S. & Philips, J. A. LC3-associated phagocytosis: host defense and microbial response. Curr. Opin. Immunol. 60, 81–90 (2019).
Köster, S. et al. Mycobacterium tuberculosis is protected from NADPH oxidase and LC3-associated phagocytosis by the LCP protein CpsA. Proc. Natl Acad. Sci. USA 114, E8711–E8720 (2017).
Abram, C. L., Roberge, G. L., Hu, Y. & Lowell, C. A. Comparative analysis of the efficiency and specificity of myeloid-Cre deleting strains using ROSA-EYFP reporter mice. J. Immunol. Methods 408, 89–100 (2014).
Schneider, C. et al. Induction of the nuclear receptor PPAR-γ by the cytokine GM-CSF is critical for the differentiation of fetal monocytes into alveolar macrophages. Nat. Immunol. 15, 1026–1037 (2014).
Deretic, V. Autophagy in tuberculosis. Cold Spring Harb. Perspect. Med. 4, a018481 (2014).
Sakowski, E. T. et al. Ubiquilin 1 promotes IFN-γ-induced xenophagy of Mycobacterium tuberculosis. PLoS Pathog. 11, e1005076 (2015).
Ponpuak, M. et al. Delivery of cytosolic components by autophagic adaptor protein p62 endows autophagosomes with unique antimicrobial properties. Immunity 32, 329–341 (2010).
Cohen, S. B. et al. Alveolar macrophages provide an early Mycobacterium tuberculosis niche and initiate dissemination. Cell Host Microbe 24, 439–446.e4 (2018).
Rothchild, A. C. et al. Alveolar macrophages generate a noncanonical NRF2-driven transcriptional response to Mycobacterium tuberculosis in vivo. Sci. Immunol. 4, eaaw6693 (2019).
Sun, D., Wu, R., Zheng, J., Li, P. & Yu, L. Polyubiquitin chain-induced p62 phase separation drives autophagic cargo segregation. Cell Res. 28, 405–415 (2018).
Runwal, G. et al. LC3-positive structures are prominent in autophagy-deficient cells. Sci. Rep. 9, 10147 (2019).
Saitoh, T. et al. Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL-1β production. Nature 456, 264–268 (2008).
Wang, Y.-T., Sansone, A., Smirnov, A., Stallings, C. L. & Orvedahl, A. Myeloid autophagy genes protect mice against fatal TNF- and LPS-induced cytokine storm syndromes. Autophagy 19, 1114–1127 (2023).
Samstein, M. et al. Essential yet limited role for CCR2+ inflammatory monocytes during Mycobacterium tuberculosis-specific T cell priming. eLife 2, e01086 (2013).
Wolf, A. J. et al. Initiation of the adaptive immune response to Mycobacterium tuberculosis depends on antigen production in the local lymph node, not the lungs. J. Exp. Med. 205, 105–115 (2008).
Gabrilovich, D. I. & Nagaraj, S. Myeloid-derived suppressor cells as regulators of the immune system. Nat. Rev. Immunol. 9, 162–174 (2009).
Bronte, V. et al. Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat. Commun. 7, 12150 (2016).
Nair, S. et al. Irg1 expression in myeloid cells prevents immunopathology during M. tuberculosis infection. J. Exp. Med. 215, 1035–1045 (2018).
Singh, B. et al. Myeloid-derived suppressor cells mediate T cell dysfunction in nonhuman primate TB granulomas. mBio 12, e03189-21 (2021).
Lu, Q. et al. Homeostatic control of innate lung inflammation by Vici syndrome gene Epg5 and additional autophagy genes promotes influenza pathogenesis. Cell Host Microbe 19, 102–113 (2016).
Veglia, F. et al. Analysis of classical neutrophils and polymorphonuclear myeloid-derived suppressor cells in cancer patients and tumor-bearing mice. J. Exp. Med. 218, e20201803 (2021).
Magcwebeba, T., Dorhoi, A. & du Plessis, N. The emerging role of myeloid-derived suppressor cells in tuberculosis. Front. Immunol. 10, 917 (2019).
Tsiganov, E. N. et al. Gr-1dim CD11b+ immature myeloid-derived suppressor cells but not neutrophils are markers of lethal tuberculosis infection in mice. J. Immunol. 192, 4718–4727 (2014).
du Plessis, N. et al. Increased frequency of myeloid-derived suppressor cells during active tuberculosis and after recent Mycobacterium tuberculosis infection suppresses T-cell function. Am. J. Respir. Crit. Care Med. 188, 724–732 (2013).
Obregón-Henao, A., Henao-Tamayo, M., Orme, I. M. & Ordway, D. J. Gr1intCD11b+ myeloid-derived suppressor cells in Mycobacterium tuberculosis infection. PLoS ONE 8, e80669 (2013).
Andrade, B. B. et al. Plasma heme oxygenase-1 levels distinguish latent or successfully treated human tuberculosis from active disease. PLoS ONE 8, e62618 (2013).
Qiu, L. et al. Severe tuberculosis induces unbalanced up‐regulation of gene networks and overexpression of IL‐22, MIP‐1α, CCL27, IP‐10, CCR4, CCR5, CXCR3, PD1, PDL2, IL‐3, IFN‐β, TIM1, and TLR2 but low antigen‐specific cellular responses. J. Infect. Dis. 198, 1514–1519 (2008).
Barber, D. L., Mayer-Barber, K. D., Feng, C. G., Sharpe, A. H. & Sher, A. CD4 T cells promote rather than control tuberculosis in the absence of PD-1-mediated inhibition. J. Immunol. 186, 1598–1607 (2011).
Grieshaber-Bouyer, R. et al. The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments. Nat. Commun. 12, 2856 (2021).
Evrard, M. et al. Developmental analysis of bone marrow neutrophils reveals populations specialized in expansion, trafficking, and effector functions. Immunity 48, 364–379.e8 (2018).
Davies, L. C., Jenkins, S. J., Allen, J. E. & Taylor, P. R. Tissue-resident macrophages. Nat. Immunol. 14, 986–995 (2013).
Bellamy, R. Susceptibility to mycobacterial infections: the importance of host genetics. Genes Immun. 4, 4–11 (2003).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005).
Hwang, S. et al. Nondegradative role of Atg5-Atg12/ Atg16L1 autophagy protein complex in antiviral activity of interferon gamma. Cell Host Microbe 11, 397–409 (2012).
Matsunaga, K. et al. Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat. Cell Biol. 11, 385–396 (2009).
Murphy, J. M. et al. The pseudokinase MLKL mediates necroptosis via a molecular switch mechanism. Immunity 39, 443–453 (2013).
Huynh, J. P. et al. Bhlhe40 is an essential repressor of IL-10 during Mycobacterium tuberculosis infection. J. Exp. Med. 215, 1823–1838 (2018).
Tan, S., Sukumar, N., Abramovitch, R. B., Parish, T. & Russell, D. G. Mycobacterium tuberculosis responds to chloride and pH as synergistic cues to the immune status of its host Cell. PLoS Pathog. 9, e1003282 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Germain, P.-L., Lun, A., Meixide, C. G., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data. F1000Research 10, 979 (2022).
Acknowledgements
We thank J. Ernst at UCSF for providing P25/Nur77-GFP/CD45.1 mice; J. Murphy at WEHI and D. Lenschow at WashU for sharing Mlkl−/− mice, and S. Tan at Tufts for sharing pCherry3 plasmid; S. Li and L. Ge at Tsinghua for discussion on autophagy analysis; and S. Andhey at WashU for helpful discussion about scRNA-seq data. The research was supported by the National Key R&D Program of China (2023YFC2306300 to Y.-T.W.), NIH grant R01 (AI132697 to C.L.S.), Burroughs Wellcome Fund Investigators in the Pathogenesis of Infectious Disease (to C.L.S.), the Philip and Sima Needleman Center for Autophagy Therapeutics and Research (to C.L.S.), NIH grant U19 (AI142784 to C.L.S. and H.W.V.), National Natural Science Foundation of China (32370804 to Y.-T.W.), Tsinghua-Peking Joint Center for Life Sciences (to Y.-T.W), Tsinghua University Dushi Program (20231080015 to Y.-T.W.), and Tsinghua University School of Medicine (to Y.-T.W.). Authors also received support from NIH grant T32 AI007172 (to M.E.M.), T32 AI007172 (to J.A.V.W.) and T32 GM007067 to (S.V.H.); and a Potts Memorial Foundation postdoctoral fellowship (to R.L.K.), Stephen I. Morse Fellowship (to S.K.N.), and Alexander & Gertrude Berg Fellowship (to N.D.). This work was supported in part by the Bursky Center for Human Immunology and Immunotherapy Programs at Washington University Immunomonitoring Laboratory. We also thank the Tsinghua University Branch of the China National Center for Protein Sciences (Beijing) and Tsinghua University Core Facilities of the Center for Biomedical Analysis, Technology Center for Protein Research, and the Cell Function Analysing Facility for technical support; and the Genome Technology Access Center at the McDonnell Genome Institute at Washington University School of Medicine.
Author information
Authors and Affiliations
Contributions
Y.-T.W. and C.L.S. designed the project, analysed the data and wrote the manuscript. S.F. performed experiments, analysed the data, made figures and assisted with manuscript writing. Y.-T.W., M.E.M., R.L.K., C.S.C., S.K.N., S.R.M., N.D. and A. Samuels performed experiments. A. Swain and M.N.A helped design and perform the scRNA-seq experiment. J.A.V.W., S.M.C., X.C., S.V.H., R.W., D.K. and A. Smirnov assisted with experiments and data analysis. H.W.V advised on project design. All authors read and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.W.V. is a founder of Casma Therapeutics and the Vaccine Company. The work reported here was not funded by either company. H.W.V. also holds shares in Vir Biotechnology, which did not fund this work. All other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Maziar Divangahi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
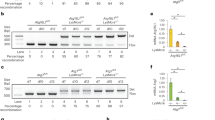
Extended Data Fig. 1 Analysis of Atg16l1 gene deletion efficiency from primary cells and weight loss of mice infected with high-dose Mtb.
a, PCR assay to confirm deletion of Atg16l1 in myeloid cells from the lung of mice. 2 representative samples of n = 4. b-c, Weight loss of mice after aerosol infection with high-dose Mtb. Data are presented as mean ± s.e.m.
Extended Data Fig. 2 Ex vivo and in vivo analysis of autophagy during Mtb infection in alveolar macrophages.
a, Representative images of LC3-stained alveolar macrophages isolated from naïve mice and then infected ex vivo with mCherry-Mtb for 48 h. LC3 (green), nuclear staining (blue) and Mtb (red). Scale bars, 2 or 3 μm. b,c, Quantitative analysis of the area of LC3 puncta (b) and % of Mtb colocalized with LC3 (c) in mCherry-Mtb+ CD11c+ alveolar macrophages infected ex vivo with mCherry-Mtb. Atg16l1f/f, n = 14 cells for area of LC3 puncta, n = 15 cells for % of LC3+ puncta Mtb area; Atg16l1f/f-CD11c-cre, n = 21 cells for area of LC3 puncta, n = 26 cells for % of LC3+ puncta Mtb area. d, Representative images of LC3-stained alveolar macrophages from mice at 14 dpi with high-dose mCherry-Mtb. LC3 (green), nuclear staining (blue) and Mtb (red). Scale bars, 2 μm. e,f, Quantitative analysis of the area of LC3 puncta (e) and % of Mtb colocalized with LC3 (f) in mCherry-Mtb+ CD11c+ alveolar macrophages from BALF of mice at 14 dpi of high-dose Mtb infection. Atg16l1f/f, n = 27 cells for area of LC3 puncta, n = 31 cells for % of LC3+ puncta Mtb area; Atg16l1f/f-CD11c-cre, n = 29 cells for area of LC3 puncta, n = 31 cells for % of LC3+ puncta Mtb area. Data representative (Means ± s.e.m.) of n = 3 biological repeats. P values calculated by two-tailed Mann-Whitney tests. * for P < 0.05, and **** P < 0.0001. ns = not significant.
Extended Data Fig. 3 Heightened inflammation in lungs of autophagy deficient mice after high-dose Mtb infection.
a, Concentration of cytokines in high-dose Mtb infected lungs as detected by multiplex cytokine panel. Data (Means ± s.e.m.) pooled from 2 independent experiments. Atg16l1f/f, n = 7 mice at 14, 21dpi; Atg16l1f/f-LysM-cre, n = 9 mice at 14 dpi, n = 7 mice at 21 dpi. P values were calculated by two-tailed t-tests. b, The number of total immune cells in the lung of mice during high-dose Mtb infection. Data (Means ± s.e.m.) pooled from 3 independent experiments. Atg16l1f/f, n = 15 mice at naïve condition, n = 7 mice at 14 dpi, n = 12 mice at 21 dpi; Atg16l1f/f-LysM-cre, n = 16 mice at naïve condition, n = 10 mice at 14 dpi, n = 17 mice at 21 dpi. P values were calculated by two-tailed Mann-Whitney tests. * for P < 0.05, ** for P < 0.01, *** P < 0.001, and **** P < 0.0001. ns = not significant.
Extended Data Fig. 4 Histology analysis of naïve mouse lungs.
a, Representative H&E-stained sections of naïve mouse lungs. Data represent n = 3 mice each group.
Extended Data Fig. 5 Analysis of antigen specific T cells and MHC-II level on innate immune cells.
a,b,c Quantification (a,b) and representative flow plot (c) of Ag85a and ESAT6 positive CD4+ T cells from lungs and mLNs of mice at 14 dpi (a) and 21 dpi (b,c) of high-dose Mtb infection. Atg16l1f/f, n = 6; Atg16l1f/f-LysM-cre, n = 6 mice. d, MHC-II mean fluorescent intensity (MFI) in alveolar macrophages, non-alveolar macrophages, DCs, and monocytes from lungs at 14 dpi of high-dose infection with high-dose Mtb. Atg16l1f/f, n = 6; Atg16l1f/f-LysM-cre, n = 5 mice. Data (Means ± s.e.m.) from 2 independent experiments are graphed. P values calculated by two-tailed Mann-Whitney tests. ns = not significant.
Extended Data Fig. 6 Accumulation of Ly6GintGr-1int neutrophils in autophagy deficient mice is associated with susceptibility and high Mtb burden.
a,d,f, The number and percentage of Gr-1 high (Gr-1hi) neutrophils (a), Gr-1 int (Gr-1int) neutrophils(d), and alveolar macrophages and DCs (f) in lungs of high-dose Mtb infected mice treated with neutrophil depletion antibody (1A8) or isotype control immunoglobulin (control) at 21 dpi of high-dose Mtb infection. Atg16l1f/f, n = 9 mice for control treatment, n = 10 mice for 1A8 treatment; Atg16l1f/f-LysM-cre, n = 9 mice for control treatment, n = 10 mice for 1A8 treatment. b, Mtb CFU in lungs at 21 dpi of high-dose Mtb infections with 1A8 or control antibody treatment. Atg16l1f/f, n = 10 mice for control treatment, n = 11 mice for 1A8 treatment; Atg16l1f/f-LysM-cre, n = 10 mice for control treatment, n = 12 mice for 1A8 treatment. c,e, number of Mtb infected cells, measured as GFP+ cells in the lungs at 21 dpi with 1A8 or control antibody treatment. Atg16l1f/f, n = 9 mice for control treatment, n = 10 mice for 1A8 treatment; Atg16l1f/f-LysM-cre, n = 9 mice for control treatment, n = 10 mice for 1A8 treatment. Data (Means ± s.e.m.) pooled from 3 independent experiments. P values calculated by two-tailed Mann-Whitney tests. * for P < 0.05, ** for P < 0.01, *** P < 0.001, and **** P < 0.0001. ns = not significant.
Extended Data Fig. 7 Non-alveolar macrophages in the lungs did not exhibit increased apoptosis at 21 dpi.
a, Representative histogram and quantification of flow cytometry analysis of FLICA+ non-alveolar macrophages from lungs of mice at 21 dpi of high-dose Mtb infection. Grey histogram indicates isotype control. Data are presented as mean ± s.e.m. Atg16l1f/f, n = 6; Atg16l1f/f-LysM-cre, n = 6 mice. P values calculated by two-tailed Mann-Whitney tests. ns = not significant.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Fig. 2
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, S., McNehlan, M.E., Kinsella, R.L. et al. Autophagy promotes efficient T cell responses to restrict high-dose Mycobacterium tuberculosis infection in mice. Nat Microbiol 9, 684–697 (2024). https://doi.org/10.1038/s41564-024-01608-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-024-01608-x