Abstract
Whether or not autophagy has a role in defence against Mycobacterium tuberculosis infection remains unresolved. Previously, conditional knockdown of the core autophagy component ATG5 in myeloid cells was reported to confer extreme susceptibility to M. tuberculosis in mice, whereas depletion of other autophagy factors had no effect on infection. We show that doubling cre gene dosage to more robustly deplete ATG16L1 or ATG7 resulted in increased M. tuberculosis growth and host susceptibility in mice, although ATG5-depleted mice are more sensitive than ATG16L1- or ATG7-depleted mice. We imaged individual macrophages infected with M. tuberculosis and identified a shift from apoptosis to rapid necrosis in autophagy-depleted cells. This effect was dependent on phagosome permeabilization by M. tuberculosis. We monitored infected cells by electron microscopy, showing that autophagy protects the host macrophage by partially reducing mycobacterial access to the cytosol. We conclude that autophagy has an important role in defence against M. tuberculosis in mammals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
RNA-seq data can be accessed on the GEO database, accession number GSE226503 (https://www.ncbi.nlm.nih.gov/geo/). All other data are available in the main text, source data, Extended Data figures or Supplementary Information. Source data are provided with this paper.
Change history
02 February 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41564-024-01605-0
References
Levine, B. & Kroemer, G. Biological functions of autophagy genes: a disease perspective. Cell 176, 11–42 (2019).
Chang, C., Jensen, L. E. & Hurley, J. H. Autophagosome biogenesis comes out of the black box. Nat. Cell Biol. 23, 450–456 (2021).
Deretic, V. & Lazarou, M. A guide to membrane atg8ylation and autophagy with reflections on immunity. J. Cell Biol. 221, e202203083 (2022).
Lu, Q. et al. Homeostatic control of innate lung inflammation by Vici syndrome gene Epg5 and additional autophagy genes promotes influenza pathogenesis. Cell Host Microbe 19, 102–113 (2016).
Park, S. et al. Autophagy genes enhance murine gammaherpesvirus 68 reactivation from latency by preventing virus-induced systemic inflammation. Cell Host Microbe 19, 91–101 (2016).
Martinez, J. et al. Noncanonical autophagy inhibits the autoinflammatory, lupus-like response to dying cells. Nature 533, 115–119 (2016).
Thurston, T. L. M., Wandel, M. P., Muhlinen, N., von, Foeglein, Á. & Randow, F. Galectin 8 targets damaged vesicles for autophagy to defend cells against bacterial invasion. Nature 482, 414–418 (2012).
Jia, J. et al. AMPK, a regulator of metabolism and autophagy, is activated by lysosomal damage via a novel galectin-directed ubiquitin signal transduction system. Mol. Cell 77, 951–969 (2020).
Global Tuberculosis Report 2020 (World Health Organization, 2020).
Huang, L., Nazarova, E. V. & Russell, D. G. Mycobacterium tuberculosis: bacterial fitness within the host macrophage. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.BAI-0001-2019 (2019).
Mayer-Barber, K. D. et al. Host-directed therapy of tuberculosis based on interleukin-1 and type I interferon crosstalk. Nature 511, 99–103 (2014).
Gutierrez, M. G. et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119, 753–766 (2004).
Kimmey, J. M. et al. Unique role for ATG5 in neutrophil-mediated immunopathology during M. tuberculosis infection. Nature 528, 565–569 (2015).
Behar, S. M. & Baehrecke, E. H. Tuberculosis: autophagy is not the answer. Nature 528, 482–483 (2015).
Watson, R. O., Manzanillo, P. S. & Cox, J. S. Extracellular M. tuberculosis DNA targets bacteria for autophagy by activating the host DNA-sensing pathway. Cell 150, 803–815 (2012).
Castillo, E. F. et al. Autophagy protects against active tuberculosis by suppressing bacterial burden and inflammation. Proc. Natl Acad. Sci. USA 109, E3168–E3176 (2012).
Wallis, R. S. & Hafner, R. Advancing host-directed therapy for tuberculosis. Nat. Rev. Immunol. 15, 255–263 (2015).
McCubbrey, A. L., Allison, K. C., Lee-Sherick, A. B., Jakubzick, C. V. & Janssen, W. J. Promoter specificity and efficacy in conditional and inducible transgenic targeting of lung macrophages. Front. Immunol. 8, 1618 (2017).
Vooijs, M., Jonkers, J. & Berns, A. A highly efficient ligand‐regulated Cre recombinase mouse line shows that LoxP recombination is position dependent. EMBO Rep. 2, 292–297 (2001).
Heffner, C. S. et al. Supporting conditional mouse mutagenesis with a comprehensive cre characterization resource. Nat. Commun. 3, 1218 (2012).
Thomson, J. G., Rucker, E. B. & Piedrahita, J. A. Mutational analysis of loxP sites for efficient Cre‐mediated insertion into genomic DNA. Genesis 36, 162–167 (2003).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition). Autophagy 17, 1–382 (2021).
Ji, D. X. et al. Type I interferon-driven susceptibility to Mycobacterium tuberculosis is mediated by IL-1Ra. Nat. Microbiol. 4, 2128–2135 (2019).
Moreira-Teixeira, L. et al. Type I IFN exacerbates disease in tuberculosis-susceptible mice by inducing neutrophil-mediated lung inflammation and NETosis. Nat. Commun. 11, 5566 (2020).
Schmidt-Supprian, M. & Rajewsky, K. Vagaries of conditional gene targeting. Nat. Immunol. 8, 665–668 (2007).
Pépin, G. et al. Cre-dependent DNA recombination activates a STING-dependent innate immune response. Nucleic Acids Res. 44, 5356–5364 (2016).
Queval, C. J., Brosch, R. & Simeone, R. The macrophage: a disputed fortress in the battle against Mycobacterium tuberculosis. Front. Microbiol. 8, 2284 (2017).
Mahamed, D. et al. Intracellular growth of Mycobacterium tuberculosis after macrophage cell death leads to serial killing of host cells. eLife 6, e22028 (2017).
Lerner, T. R. et al. Mycobacterium tuberculosis replicates within necrotic human macrophages. J. Cell Biol. 216, 583–594 (2017).
Tan, J. M. J. et al. An ATG16L1-dependent pathway promotes plasma membrane repair and limits Listeria monocytogenes cell-to-cell spread. Nat. Microbiol. 3, 1472–1485 (2018).
Jia, J. et al. Galectin-3 coordinates a cellular system for lysosomal repair and removal. Dev. Cell 52, 69–87 (2019).
López-Jiménez, A. T. et al. The ESCRT and autophagy machineries cooperate to repair ESX-1-dependent damage at the Mycobacterium-containing vacuole but have opposite impact on containing the infection. PLoS Pathog. 14, e1007501 (2018).
Houben, D. et al. ESX-1-mediated translocation to the cytosol controls virulence of mycobacteria. Cell Microbiol. 14, 1287–1298 (2012).
Simeone, R. et al. Phagosomal rupture by Mycobacterium tuberculosis results in toxicity and host cell death. PLoS Pathog. 8, e1002507 (2012).
Smith, J. et al. Evidence for pore formation in host cell membranes by ESX-1-secreted ESAT-6 and its role in Mycobacterium marinum escape from the vacuole. Infect. Immun. 76, 5478–5487 (2008).
Conrad, W. H. et al. Mycobacterial ESX-1 secretion system mediates host cell lysis through bacterium contact-dependent gross membrane disruptions. Proc. Natl Acad. Sci. USA 114, 1371–1376 (2017).
Bell, S. L., Lopez, K. L., Cox, J. S., Patrick, K. L. & Watson, R. O. Galectin-8 senses phagosomal damage and recruits selective autophagy adapter TAX1BP1 to control Mycobacterium tuberculosis infection in macrophages. Mbio 12, e01871-20 (2021).
Rosenberg, O. S. et al. Substrates control multimerization and activation of the multi-domain ATPase motor of type VII secretion. Cell 161, 501–512 (2015).
Stanley, S. A., Johndrow, J. E., Manzanillo, P. & Cox, J. S. The Type I IFN response to infection with Mycobacterium tuberculosis requires ESX-1-mediated secretion and contributes to pathogenesis. J. Immunol. 178, 3143–3152 (2007).
Watson, R. O. et al. The cytosolic sensor cGAS detects Mycobacterium tuberculosis DNA to induce type I interferons and activate autophagy. Cell Host Microbe 17, 811–819 (2015).
Beckwith, K. S. et al. Plasma membrane damage causes NLRP3 activation and pyroptosis during Mycobacterium tuberculosis infection. Nat. Commun. 11, 2270 (2020).
Lim, J.-A., Zare, H., Puertollano, R. & Raben, N. Atg5flox-derived autophagy-deficient model of Pompe disease: does it tell the whole story? Mol. Ther. Methods Clin. Dev. 7, 11–14 (2017).
Koo, I. C. et al. ESX‐1‐dependent cytolysis in lysosome secretion and inflammasome activation during mycobacterial infection. Cell Microbiol. 10, 1866–1878 (2008).
Wassermann, R. et al. Mycobacterium tuberculosis differentially activates cGAS- and inflammasome-dependent intracellular immune responses through ESX-1. Cell Host Microbe 17, 799–810 (2015).
Mayer-Barber, K. D. et al. Cutting edge: caspase-1 independent IL-1β production is critical for host resistance to Mycobacterium tuberculosis and does not require TLR signaling in vivo. J. Immunol. 184, 3326–3330 (2010).
Yousefi, S. et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat. Cell Biol. 8, 1124–1132 (2006).
Zhao, X. et al. Bcl-xL mediates RIPK3-dependent necrosis in M. tuberculosis-infected macrophages. Mucosal Immunol. 10, 1553–1568 (2017).
Behar, S. M. et al. Apoptosis is an innate defense function of macrophages against Mycobacterium tuberculosis. Mucosal Immunol. 4, 279–287 (2011).
Repasy, T. et al. Bacillary replication and macrophage necrosis are determinants of neutrophil recruitment in tuberculosis. Microbes Infect. 17, 564–574 (2015).
Paik, S., Kim, J. K., Chung, C. & Jo, E.-K. Autophagy: a new strategy for host-directed therapy of tuberculosis. Virulence 10, 448–459 (2018).
Flores-Valdez, M. A., Segura-Cerda, C. A. & Gaona-Bernal, J. Modulation of autophagy as a strategy for development of new vaccine candidates against tuberculosis. Mol. Immunol. 97, 16–19 (2018).
Maskey, D. et al. ATG5 is induced by DNA-damaging agents and promotes mitotic catastrophe independent of autophagy. Nat. Commun. 4, 2130 (2013).
Abram, C. L., Roberge, G. L., Hu, Y. & Lowell, C. A. Comparative analysis of the efficiency and specificity of myeloid-Cre deleting strains using ROSA-EYFP reporter mice. J. Immunol. Methods 408, 89–100 (2014).
Yu, Y.-R. A. et al. A protocol for the comprehensive flow cytometric analysis of immune cells in normal and inflamed murine non-lymphoid tissues. PLoS ONE 11, e0150606 (2016).
Chen, M. et al. Lipid mediators in innate immunity against tuberculosis: opposing roles of PGE2 and LXA4 in the induction of macrophage death. J. Exp. Med. 205, 2791–2801 (2008).
Park, J. S., Tamayo, M. H., Gonzalez‐Juarrero, M., Orme, I. M. & Ordway, D. J. Virulent clinical isolates of Mycobacterium tuberculosis grow rapidly and induce cellular necrosis but minimal apoptosis in murine macrophages. J. Leukoc. Biol. 79, 80–86 (2006).
Divangahi, M., Desjardins, D., Nunes-Alves, C., Remold, H. G. & Behar, S. M. Eicosanoid pathways regulate adaptive immunity to Mycobacterium tuberculosis. Nat. Immunol. 11, 751–758 (2010).
Martin, C. J. et al. Efferocytosis is an innate antibacterial mechanism. Cell Host Microbe 12, 289–300 (2012).
Westman, J., Grinstein, S. & Marques, P. E. Phagocytosis of necrotic debris at sites of injury and inflammation. Front. Immunol. 10, 3030 (2020).
Dallenga, T. et al. M. tuberculosis-induced necrosis of infected neutrophils promotes bacterial growth following phagocytosis by macrophages. Cell Host Microbe 22, 519–530 (2017).
Sogi, K. M., Lien, K. A., Johnson, J. R., Krogan, N. J. & Stanley, S. A. The tyrosine kinase inhibitor Gefitinib restricts Mycobacterium tuberculosis growth through increased lysosomal biogenesis and modulation of cytokine signaling. ACS Infect. Dis. 3, 564–574 (2017).
Penn, B. H. et al. An Mtb-human protein–protein interaction map identifies a switch between host antiviral and antibacterial responses. Mol. Cell 71, 637–648 (2018).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Hwang, S. et al. Nondegradative role of Atg5-Atg12/ Atg16L1 autophagy protein complex in antiviral activity of interferon gamma. Cell Host Microbe. 11, 397–409 (2012).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005).
Acknowledgements
We acknowledge members of the Cox, Stanley, Vance and Portnoy laboratories for helpful discussion. We thank D. A. Portnoy (University of California Berkeley) for graciously letting us use his Keyence microscope. We thank H. W. Virgin (Washington University) for providing all the mouse lines used in this study. We thank the staff of the Office of Laboratory Animal Care of the University of California Berkeley for the care provided to the animals used in this study. We thank the staff at the University of California Berkeley Electron Microscope Laboratory for advice and assistance in electron microscopy sample preparation and data collection. The sequencing for RNA-seq was carried out at the DNA Technologies and Expression Analysis Cores at the University of California Davis Genome Center, supported by NIH Shared Instrumentation grant 1S10OD010786-01. Embedding in wax, sectioning and staining for histology was performed at the University of California Davis Center for Genomic Pathology Laboratory. This work was supported by NIH grants AI162583 and AI063302 and National Institute of Allergy and Infectious Diseases Host Pathogen Map Initiative grant U19AI135990 (J.S.C.).
Author information
Authors and Affiliations
Contributions
J.S.C., G.R.G. and G.M. conceptualized the study. G.R.G. was responsible for conducting experiments, acquisition of data and analysis. G.R.G. and M.C. performed qPCR experiments. N.G. performed RNA-seq data analysis. G.R.G. and L.M.P. performed live imaging experiments. L.M.P., G.M. and J.S.C. provided reagents. G.R.G., A.W.R., R.M.M., H.M., T.R., J.B. and E.V.D. performed in vivo experiments. G.R.G., R.R.-L. and R.M.M. performed M. marinum experiments. G.R.G. and H.M. performed immunofluorescence experiments. G.R.G., D.J. and R.Z. conducted electron microscopy experiments. G.R.G. and H.N. performed western blot experiments. G.R.G. and J.S.C. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Maziar Divangahi, Vojo Deretic and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sequences of LoxP sites, map of LoxP site insertions and measurement of Cre expression levels.
a, Sequences of LoxP sites in Atg5fl/fl, Atg16L1fl/fl and Atg7fl/fl mice. b, Map of LoxP sites and Neomycin resistance cassettes insertions in Atg5fl/fl, Atg16L1fl/fl and Atg7fl/fl mice. Grey boxes denote exons. c, Measurement of mRNA levels of Cre in bone marrow-derived macrophages differentiated for 10 days. Results are the means ± SD from 3 biological replicates. n.d. = not detected. Statistical analysis performed with two-sided t-test.
Extended Data Fig. 2 Gating strategy used to isolate lung populations in M. tuberculosis-infected mice.
Lungs from infected mice were harvested after 21 or 70 days of infection. Single cells from lung homogenates were sorted into different immune cell populations according to the strategy outlined in this figure.
Extended Data Fig. 3 Additional in vivo data on autophagy-mutant mice infected with M. tuberculosis.
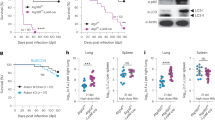
Mice (n = 10-12 per group, except for LysMcre+/+ for which n = 5) were infected with approximately 100 colony-forming units (c.f.u.) of M. tuberculosis and tracked for weight change (a-c). Data is presented as means ± SD. Mice that lost 15% body weight were sacrificed are counted at −15% for subsequent time points. d-e, CFU counts from the spleens of mice (n = 2-6 per group) infected with approximately 100 c.f.u of M. tuberculosis. Spleens were harvested at 21 days postinfection (d) or 70 days postinfection (e). Data is presented as means ± SD. f-g, Proportion of immune cell populations in the lungs of Atg7 (f) and Atg16L1 (g) mutant mice infected for 70 days with M. tuberculosis. Results are the means ± SD from 2–7 mice per condition. Nt = Neutrophils, Tc = T cells, Bc = B cells, Mac = Macrophages, DC = Dendritic cells, Mono = Ly6Chi inflammatory monocytes. For statistical analysis for CFU and histology experiments, LysMcre+/- and LysMcre+/+ conditions were compared to the respective LysMwt/wt control. * P ≤ 0.05, ** P ≤ 0.01 by two-sided Mann–Whitney test.
Extended Data Fig. 4 Statistics of cell death and M. tuberculosis growth during time-course macrophage infection with M. tuberculosis.
a, b, Same data as in Fig. 4b, c but each day is presented as an individual graph. Data represents number of macrophages (a) and Mean Fluorescent Intensity of fluorescent bacteria (b). Results are the means ± SD from 4 technical replicates. For statistical analysis, LysMcre+/- and LysMcre+/+ conditions were compared to the respective LysMwt/wt control. * P ≤ 0.05 by two-sided t-test/.
Extended Data Fig. 5 Growth of M. tuberculosis in autophagy-mutant BMMs and susceptibility of Atg5-flox hemizygous macrophages.
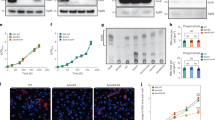
a, BMMs from autophagy-mutant mice were infected with M. tuberculosis at a MOI of 2. CFU were enumerated at day 0, day 2 and day 4 postinfection. Results are the means ± SD from 3 technical replicates per condition. For statistical analysis, LysMcre+/- and LysMcre+/+ conditions were compared to the respective LysMwt/wt control for the day 4 condition. * P ≤ 0.05 by two-sided t-test. Grey star denotes conditions were the macrophage monolayer was disrupted. b, Macrophages from Atg5fl/wt, Atg5fl/wt LysMcre+/-, Atg5fl/wt LysMcre+/+ and Atg5fl/fl LysMcre+/+ were infected with M. tuberculosis at a MOI of 1 and macrophages were enumerated by imaging every day for 7 days. Macrophages carrying one wild-type allele of Atg5 and one recombined allele (Atg5fl/wt LysMcre+/-, Atg5fl/wt LysMcre+/+) were indistinguishable from wild-type cells. Each condition was tested in technical triplicates.
Extended Data Fig. 6 Absence of autophagy promotes phagosome escape of Mycobacteria in macrophages.
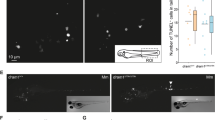
a, Illustrative images of wild-type or ΔeccC M. tuberculosis bacteria in Atg5fl/fl or Atg5fl/fl LysMcre+/+ macrophages after 24 hours of infection. Orange asterisks denote bacteria fully enclosed in phagosomes, blue asterisks show bacteria with poor membrane preservation, red asterisks denote bacteria in the cytosol. Scale bar = 0.4 µm. b, Quantification of bacteria fully enclosed in phagosomes, present directly in the cytosol or with poor membrane preservation. Data is from a single experiment. A minimum of 40 bacteria per condition were quantified. c, Actin tail formation by M. marinum expressing mCherry in Atg5fl/fl or Atg5fl/fl LysMcre+/+ macrophages was measured at 6.5 h, 8 h, 9.5 h and 11 hours postinfection. Results are the means ± SD for 15 images from two technical replicates. * P ≤ 0.05 by two-sided t-test. d, Autophagy-mutant macrophages were infected with M. tuberculosis expressing GFP or ΔeccC M. tuberculosis expressing GFP. Colocalization with Gal-3 was measured at 10 h postinfection. Results are the means ± SD from three technical replicates. For statistical analysis, LysMcre+/- and LysMcre+/+ conditions were compared to the respective LysMwt/wt control. * P ≤ 0.05, ** P ≤ 0.01, *** P ≤ 0.001 by two-sided Mann–Whitney test.
Extended Data Fig. 7 Survival of uninfected autophagy-deficient macrophages and statistics of cell death during infection with ΔeccC M. tuberculosis.
a-b, Quantification of number of uninfected macrophages over time. Data is averaged from two independent experiments with each condition tested in technical duplicates. Data presented as a time course (a) or as selected time points for statistical analysis (b). Data is presented as means ± SD. c, Same data as in Fig. 5c but with selected timepoints presented as an individual graphs. Results are the means ± SD from two independent experiments with each condition tested in technical duplicates. For statistical analysis, number of macrophages in cre-positive conditions were compared to the respective LysMwt/wt control. * P ≤ 0.05 by two-sided t-test.
Extended Data Fig. 8 RNA-seq on autophagy-mutant macrophages infected with M. tuberculosis. RNA was collected from uninfected BMMs and BMMs infected with M. tuberculosis for 48 h.
a-c, Volcano plot showing differentially expressed genes in Atg5fl/fl LysMcre+/+. Grey line demarcates adj.p.value of 0.05. (a), Atg16L1fl/fl LysMcre+/+ (b) and Atg7fl/fl LysMcre+/+ (c) macrophages compared to their respective flox controls. d-f, Venn diagrams showing genes with significant transcriptional changes (adj.p.value ≤ 0.05) for the cre-positive conditions compared to their respective LysMwt/wt control. Venn diagram for all DE genes in uninfected conditions (d), all DE genes in infected conditions (e) and genes exclusively differentially expressed during infection (that is with genes with significant DE in uninfected conditions filtered out) (f). g-i, Gene enrichment analysis showing pathways enriched in M. tuberculosis-infected cells. Genes with significant transcriptional changes (adj.p.value ≤ 0.05) were used for pathway enrichment using the hypergeometric test (HGT). Datasets for enrichment included: KEGG, REACTOME, MSigDB C2_CP and C3, GO, CORUM, and HALLMARK. Top 15 pathways commonly enriched in Atg5fl/fl LysMcre+/+, Atg16L1fl/fl LysMcre+/+ and Atg7fl/fl LysMcre+/+ (g), Top 15 pathways commonly enriched in all 3 genotypes specifically during infection (h) and top 15 pathways enriched exclusively in Atg5fl/fl LysMcre+/+ macrophages (i).
Extended Data Fig. 9 M. tuberculosis infection induces necrosis-like cell death and efferocytosis in autophagy-deficient macrophages.
a, Western blot for pRIP3, RIP3, BCL-xL and β-actin on cell lysates from uninfected BMMs and BMMs infected for 48 hours with M. tuberculosis. b, Measurement by ELISA of IL-1β secretion in supernatant of macrophages infected for 48 hours with M. tuberculosis. Results are the means ± SD from two independent experiments, in which each condition was tested in technical triplicates. * P ≤ 0.05 by two-sided t-test. c-d, Atg5fl/fl LysMcre+/+ macrophages were infected with GFP-expressing M. tuberculosis and imaged every 30 minutes over a 6-day period (Supplementary video 3). Time and mode of cell death were recorded for 145 Atg5fl/fl LysMcre+/+ macrophages infected with GFP-expressing M. tuberculosis from 3 independent experiments. The average time of death by necrosis-like cell death (c) and the delay between efferocytosis and subsequent cell death (d) were measured. Center line: median; box limits: upper and lower quartiles; whiskers: 1.5x interquartile range. n = 80 and 65, respectively.
Extended Data Fig. 10 Illustrative images and statistics of PI and CellEvent staining of macrophages infected with M. tuberculosis.
a, Illustrative images of experiment quantified in Fig. 6a, b. Autophagy-deficient macrophages were infected with M. tuberculosis at a MOI of 1. Propidium iodide and CellEvent Caspase-3/7 reagent were added to the media at the beginning of the infection. Cells were imaged every 30 minutes over a period of 8 days. Scale bar denotes 100 µm. b, Same data as in Fig. 6a, b but with selected timepoints presented as an individual graphs. Results are the means ± SD with each condition tested in technical triplicates. For statistical analysis, number of CellEvent (b) and propodium iodide (c) positive macrophages in cre-positive conditions were compared to the respective LysMwt/wt control. * P ≤ 0.05, ** P ≤ 0.01, *** P ≤ 0.001 by two-sided t-test.
Supplementary information
Supplementary Information
Supplementary video legends, methods for Extended Data and references.
Supplementary Video 1
Time-lapse sequence of autophagy-deficient macrophages infected with GFP-expressing M. tuberculosis.
Supplementary Video 2
Time-lapse sequence of autophagy-deficient macrophages infected with M. tuberculosis and incubated with PI and CellEvent.
Supplementary Video 3
Illustrative video of necrosis and efferocytosis of autophagy-deficient macrophages infected with M. tuberculosis.
Source data
Source Data Fig. 1
Unprocessed gels for Fig. 1a–c and unprocessed blots for Fig. 1g,h.
Source Data Fig. 1
Statistical source data for Fig. 1d–f.
Source Data Fig. 2
Statistical source data for Fig. 2 and Extended Data Fig. 3a–c.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 4
Statistical source data for Fig. 4 and Extended Data Fig. 4a,b.
Source Data Fig. 5
Statistical source data for Fig. 5 and Extended Data Fig. 7c.
Source Data Fig. 6
Statistical source data for Fig. 6 and Extended Data Figs. 9c,d and 10b,c.
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1c.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3d–g.
Source Data Extended Data Fig. 5
Statistical source data for Extended Data Fig. 5a,b.
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6a–d.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7a,b.
Source Data Extended Data Fig 9
Statistical source data for Extended Data Fig. 9b.
Source Data Extended Data Fig. 9
Unprocessed blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Golovkine, G.R., Roberts, A.W., Morrison, H.M. et al. Autophagy restricts Mycobacterium tuberculosis during acute infection in mice. Nat Microbiol 8, 819–832 (2023). https://doi.org/10.1038/s41564-023-01354-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01354-6
This article is cited by
-
Autophagy promotes efficient T cell responses to restrict high-dose Mycobacterium tuberculosis infection in mice
Nature Microbiology (2024)
-
Osthole ameliorates myonecrosis caused by Clostridium perfringens type A infection in mice
One Health Advances (2023)
-
Autophagy counteracts Mycobacterium tuberculosis infection at early stages
Nature Structural & Molecular Biology (2023)
-
Autophagy is part of the answer to tuberculosis
Nature Microbiology (2023)
-
The ubiquitin ligase TRIM32 promotes the autophagic response to Mycobacterium tuberculosis infection in macrophages
Cell Death & Disease (2023)