Abstract
The water-selective channel protein aquaporin-4 (AQP4) contributes to the migration and proliferation of gliomas, and to their resistance to therapy. Here we show, in glioma cell cultures, in subcutaneous and orthotopic gliomas in rats, and in glioma tumours in patients, that transmembrane water-efflux rate is a sensitive biomarker of AQP4 expression and can be measured via conventional dynamic-contrast-enhanced magnetic resonance imaging. Water-efflux rates correlated with stages of glioma proliferation as well as with changes in the heterogeneity of intra-tumoural and inter-tumoural AQP4 in rodent and human gliomas following treatment with temozolomide and with the AQP4 inhibitor TGN020. Regions with low water-efflux rates contained higher fractions of stem-like slow-cycling cells and therapy-resistant cells, suggesting that maps of water-efflux rates could be used to identify gliomas that are resistant to therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. The raw and analysed datasets generated during the study are too large to be publicly shared, yet they are available for research purposes from the corresponding authors on reasonable request. The raw patient data are available from the authors, subject to approval from the IRB of the Shandong Provincial Hospital affiliated to Shandong First Medical University. Source data are provided with this paper.
References
Omuro, A. & DeAngelis, L. M. Glioblastoma and other malignant gliomas. JAMA 310, 1842–1850 (2013).
Milano, M. T. et al. Patterns and timing of recurrence after temozolomide-based chemoradiation for glioblastoma. Int. J. Radiat. Oncol. Biol. Phys. 78, 1147–1155 (2010).
Sun, D.-P., Lee, Y.-W., Chen, J.-T., Lin, Y.-W. & Chen, R.-M. The bradykinin–BDKRB1 axis regulates aquaporin 4 gene expression and consequential migration and invasion of malignant glioblastoma cells via a Ca2+–MEK1–ERK1/2–NF-κB mechanism. Cancers 12, 667 (2020).
Nico, B. et al. Aquaporin-4 contributes to the resolution of peritumoural brain oedema in human glioblastoma multiforme after combined chemotherapy and radiotherapy. Eur. J. Cancer 45, 3315–3325 (2009).
Papadopoulos, M. C. & Verkman, A. S. Aquaporin water channels in the nervous system. Nat. Rev. Neurosci. 14, 265–277 (2013).
Maugeri, R. et al. Aquaporins and brain tumors. Int. J. Mol. Sci. 17, 1029 (2016).
Warth, A., Mittelbronn, M. & Wolburg, H. Redistribution of the water channel protein aquaporin-4 and the K+ channel protein Kir4.1 differs in low- and high-grade human brain tumors. Acta Neuropathol. 109, 418–426 (2005).
Montgomery, M. K. et al. Glioma-induced alterations in neuronal activity and neurovascular coupling during disease progression. Cell Rep. 31, 107500 (2020).
Noell, S. et al. Dynamics of expression patterns of AQP4, dystroglycan, agrin and matrix metalloproteinases in human glioblastoma. Cell Tissue Res. 347, 429–441 (2012).
Tome-Garcia, J. et al. Analysis of chromatin accessibility uncovers TEAD1 as a regulator of migration in human glioblastoma. Nat. Commun. 9, 1–13 (2018).
Lan, Y. L. et al. Gamabufotalin induces a negative feedback loop connecting ATP1A3 expression and the AQP4 pathway to promote temozolomide sensitivity in glioblastoma cells by targeting the amino acid Thr794. Cell Prolif. 53, 1–21 (2020).
Ruggieri, S. et al. Dp71 expression in human glioblastoma. Int. J. Mol. Sci. 20, 7–10 (2019).
Ruiz-Ontañon, P. et al. Cellular plasticity confers migratory and invasive advantages to a population of glioblastoma-initiating cells that infiltrate peritumoral tissue. Stem Cells 31, 1075–1085 (2013).
Kitchen, P. et al. Targeting aquaporin-4 subcellular localization to treat central nervous system edema. Cell 181, 784–799 (2020).
Fukuda, A. M. et al. Posttraumatic reduction of edema with aquaporin-4 RNA interference improves acute and chronic functional recovery. J. Cereb. Blood Flow. Metab. 33, 1621–1632 (2013).
Hoang-Minh, L. B. et al. Infiltrative and drug‐resistant slow‐cycling cells support metabolic heterogeneity in glioblastoma. EMBO J. 37, 1–21 (2018).
Suvà, M. L. & Tirosh, I. The glioma stem cell model in the era of single-cell genomics. Cancer Cell 37, 630–636 (2020).
Klughammer, J. et al. The DNA methylation landscape of glioblastoma disease progression shows extensive heterogeneity in time and space. Nat. Med. 24, 1611–1624 (2018).
Blocher, J. et al. Aquaporins AQP1 and AQP4 in the cerebrospinal fluid of bacterial meningitis patients. Neurosci. Lett. 504, 23–27 (2011).
Castañeyra-Ruiz, L. et al. Aquaporin-4 expression in the cerebrospinal fluid in congenital human hydrocephalus. Fluids Barriers CNS 10, 18 (2013).
Mukherjee, A., Wu, D., Davis, H. C. & Shapiro, M. G. Non-invasive imaging using reporter genes altering cellular water permeability. Nat. Commun. 7, 13891 (2016).
Okuchi, S. et al. Diagnostic accuracy of dynamic contrast-enhanced perfusion MRI in stratifying gliomas: a systematic review and meta-analysis. Cancer Med. 8, 5564–5573 (2019).
Bai, R. et al. Shutter-speed DCE-MRI analyses of human glioblastoma multiforme (GBM) data. J. Magn. Reson. Imaging 52, 850–863 (2020).
Li, X., Mangia, S., Lee, J.-H., Bai, R. & Springer, C. S. J. NMR shutter-speed elucidates apparent population inversion of 1H2O signals due to active transmembrane water cycling. Magn. Reson. Med. 82, 411–424 (2019).
Yang, B. & Verkman, A. S. Water and glycerol permeabilities of aquaporins 1–5 and MIP determined quantitatively by expression of epitope-tagged constructs in Xenopus oocytes. J. Biol. Chem. 272, 16140–16146 (1997).
Huber, V. J., Tsujita, M. & Nakada, T. Identification of aquaporin 4 inhibitors using in vitro and in silico methods. Bioorg. Med. Chem. 17, 411–417 (2009).
Chen, Y. et al. Down-regulation of AQP4 expression via p38 MAPK signaling in temozolomide-induced glioma cells growth inhibition and invasion impairment. J. Cell. Biochem. 118, 4905–4913 (2017).
Jovanović, M. et al. Novel TrxR1 inhibitors show potential for glioma treatment by suppressing the invasion and sensitizing glioma cells to chemotherapy. Front. Mol. Biosci. 7, 1–17 (2020).
Ding, T. et al. Knockdown a water channel protein, aquaporin-4, induced glioblastoma cell apoptosis. PLoS ONE 8, 1–9 (2013).
Mou, K. et al. AQP-4 in peritumoral edematous tissue is correlated with the degree of glioma and with expression of VEGF and HIF-α. J. Neurooncol. 100, 375–383 (2010).
Kong, H. et al. AQP4 knockout impairs proliferation, migration and neuronal differentiation of adult neural stem cells. J. Cell Sci. 121, 4029–4036 (2008).
Küppers, E. et al. AQP4 expression in striatal primary cultures is regulated by dopamine—implications for proliferation of astrocytes. Eur. J. Neurosci. 28, 2173–2182 (2008).
Suzuki, Y. et al. Aquaporin-4 positron emission tomography imaging of the human brain: first report. J. Neuroimaging 23, 219–223 (2013).
Harrison, I. F. et al. Impaired glymphatic function and clearance of tau in an Alzheimer’s disease model. Brain 143, 2576–2593 (2020).
Bai, R., Springer, C. S., Plenz, D. & Basser, P. J. Fast, Na+/K+ pump driven, steady-state transcytolemmal water exchange in neuronal tissue: a study of rat brain cortical cultures. Magn. Reson. Med. 79, 3207–3217 (2018).
Gates, E. D. H. et al. Guiding the first biopsy in glioma patients using estimated Ki-67 maps derived from MRI: conventional versus advanced imaging. Neuro. Oncol. 21, 527–536 (2019).
Kowalski-Chauvel, A. et al. Alpha-6 integrin promotes radioresistance of glioblastoma by modulating DNA damage response and the transcription factor Zeb1. Cell Death Dis. 9, 872 (2018).
Kawanishi, M. et al. Effect of delayed mild brain hypothermia on edema formation after intracerebral hemorrhage in rats. J. Stroke Cerebrovasc. Dis. 17, 187–195 (2008).
Salman, M. M. et al. Hypothermia increases aquaporin 4 (AQP4) plasma membrane abundance in human primary cortical astrocytes via a calcium/transient receptor potential vanilloid 4 (TRPV4)- and calmodulin-mediated mechanism. Eur. J. Neurosci. 46, 2542–2547 (2017).
Springer, C. S. Using 1H2O MR to measure and map sodium pump activity in vivo. J. Magn. Reson. 291, 110–126 (2018).
Kanai, R. et al. Oncolytic virus-mediated manipulation of DNA damage responses: synergy with chemotherapy in killing glioblastoma stem cells. J. Natl Cancer Inst. 104, 42–55 (2012).
Visvader, J. E. & Lindeman, G. J. Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat. Rev. Cancer 8, 755–768 (2008).
Seo, Y. S. et al. Radiation-induced changes in tumor vessels and microenvironment contribute to therapeutic resistance in glioblastoma. Front. Oncol. 9, 1259 (2019).
Jablonski, E. M., Webb, A. N., McConnell, N. A., Riley, M. C. & Hughes, F. M. Plasma membrane aquaporin activity can affect the rate of apoptosis but is inhibited after apoptotic volume decrease. Am. J. Physiol. Cell Physiol. 286, C975–C985 (2004).
Soeda, A. et al. The evidence of glioblastoma heterogeneity. Sci. Rep. 5, 7979 (2015).
Sottoriva, A. et al. Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc. Natl Acad. Sci. USA 110, 4009–4014 (2013).
Gallaher, J. A. et al. From cells to tissue: how cell scale heterogeneity impacts glioblastoma growth and treatment response. PLoS Comput. Biol. 16, 1–27 (2020).
Warth, A., Kröger, S. & Wolburg, H. Redistribution of aquaporin-4 in human glioblastoma correlates with loss of agrin immunoreactivity from brain capillary basal laminae. Acta Neuropathol. 107, 311–318 (2004).
Vodnala, S. K. et al. T cell stemness and dysfunction in tumors are triggered by a common mechanism. Science 363, eaau0135 (2019).
Papadopoulos, M. C. & Saadoun, S. Key roles of aquaporins in tumor biology. Biochim. Biophys. Acta Biomembr. 1848, 2576–2583 (2015).
Smith, A. J., Jin, B. J., Ratelade, J. & Verkman, A. S. Aggregation state determines the localization and function of M1- and M23-aquaporin-4 in astrocytes. J. Cell Biol. 204, 559–573 (2014).
Peng, Y., Wu, W., Shang, Z., Li, W. & Chen, S. Inhibition of lncRNA LINC00461/miR-216a/aquaporin 4 pathway suppresses cell proliferation, migration, invasion, and chemoresistance in glioma. Open Life Sci. 15, 532–543 (2020).
Badaut, J. et al. Brain water mobility decreases after astrocytic aquaporin-4 inhibition using RNA interference. J. Cereb. Blood Flow. Metab. 31, 819–831 (2011).
Jullienne, A. et al. Modulating the water channel AQP4 alters miRNA expression, astrocyte connectivity and water diffusion in the rodent brain. Sci. Rep. 8, 1–12 (2018).
Bai, R. et al. Feasibility of filter-exchange imaging (FEXI) in measuring different exchange processes in human brain. NeuroImage 219, 117039 (2020).
Le Bihan, D. & Iima, M. Diffusion magnetic resonance imaging: what water tells us about biological tissues. PLoS Biol. 13, 1–13 (2015).
Bai, R., Stewart, C. V., Plenz, D. & Basser, P. J. Assessing the sensitivity of diffusion MRI to detect neuronal activity directly. Proc. Natl Acad. Sci. USA 113, E1728–E1737 (2016).
Gupta, R. K. et al. Relationships between choline magnetic resonance spectroscopy, apparent diffusion coefficient and quantitative histopathology in human glioma. J. Neurooncol. 50, 215–226 (2000).
Smith, A. J. & Verkman, A. S. The ‘glymphatic’ mechanism for solute clearance in Alzheimer’s disease: game changer or unproven speculation? FASEB J. 32, 543–551 (2018).
Callaghan, P. T. & Furó, I. Diffusion–diffusion correlation and exchange as a signature for local order and dynamics. J. Chem. Phys. 120, 4032–4038 (2004).
Lasič, S., Nilsson, M., Lätt, J., Ståhlberg, F. & Topgaard, D. Apparent exchange rate mapping with diffusion MRI. Magn. Reson. Med. 66, 356–365 (2011).
Schilling, F. et al. MRI measurements of reporter-mediated increases in transmembrane water exchange enable detection of a gene reporter. Nat. Biotechnol. 35, 75–80 (2017).
Nedjati-Gilani, G. L. et al. Machine learning based compartment models with permeability for white matter microstructure imaging. NeuroImage 150, 119–135 (2017).
Hill, I. et al. Machine learning based white matter models with permeability: an experimental study in cuprizone treated in-vivo mouse model of axonal demyelination. NeuroImage 224, 117425 (2021).
Bai, R., Springer, C. S., Plenz, D. & Basser, P. J. Brain active transmembrane water cycling measured by MR is associated with neuronal activity. Magn. Reson. Med. 81, 1280–1295 (2019).
Dickie, B. R., Parker, G. J. M. & Parkes, L. M. Measuring water exchange across the blood-brain barrier using MRI. Prog. Nucl. Magn. Reson. Spectrosc. 116, 19–39 (2020).
Ohene, Y. et al. Non-invasive MRI of brain clearance pathways using multiple echo time arterial spin labelling: an aquaporin-4 study. NeuroImage 188, 515–523 (2019).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Atkins, R. J. et al. Cell quiescence correlates with enhanced glioblastoma cell invasion and cytotoxic resistance. Exp. Cell. Res. 374, 353–364 (2019).
Deleyrolle, L. P. et al. Evidence for label-retaining tumour-initiating cells in human glioblastoma. Brain 134, 1331–1343 (2011).
Filby, A., Begum, J., Jalal, M. & Day, W. Appraising the suitability of succinimidyl and lipophilic fluorescent dyes to track proliferation in non-quiescent cells by dye dilution. Methods 82, 29–37 (2015).
Landini, G., Martinelli, G. & Piccinini, F. Colour deconvolution: stain unmixing in histological imaging. Bioinformatics 37, 1485–1487 (2021).
Tanner, V. A., Ploug, T. & Tao-Cheng, J. H. Subcellular localization of SV2 and other secretory vesicle components in PC12 cells by an efficient method of preembedding EM immunocytochemistry for cell cultures. J. Histochem. Cytochem. 44, 1481–1488 (1996).
Du, J., Tao-Cheng, J. H., Zerfas, P. & McBain, C. J. The K+ channel, Kv2.1, is apposed to astrocytic processes and is associated with inhibitory postsynaptic membranes in hippocampal and cortical principal neurons and inhibitory interneurons. Neuroscience 84, 37–48 (1998).
Acknowledgements
The study was supported by the National Key Research and Development Program of China (grant 2022ZD0206000, to R.B.), the National Natural Science Foundation of China (NSFC) (grants 82172050, 81873894, and 82222032 to R.B.; grant 81641176, to Y.C.L.; grant 82202114, to W.B.), the Natural Science Foundation of Zhejiang Province, China (grant LR20H180001, to R.B.), the Taishan Scholars Program (no. tsqn20161070, to Y-C.L.) and the Natural Science Foundation of Shandong Province (grant ZR2019HM067, to Y-C.L.). We appreciate discussions and constructive comments from P. J. Basser at the National Institutes of Health, J. Polimeni at Harvard Medical School and Massachusetts General Hospital, and Z. Chen at the Department of Mathematics, Shandong University. We also thank the support from the MOE Frontier Science Center for Brain Science & Brain-Machine Integration, Zhejiang University.
Author information
Authors and Affiliations
Contributions
R.B., Y-C.L, Y.J., S.X. and G.H. designed the research study and analysed and interpreted the data. Y.J., G.H. and S.X. performed most experiments and analysed the data. Y.J., G.H., W.J., B.Q., R.L. and C.L. performed the cell-culture experiments. Y.J., G.H. and Z.W performed the animal experiments. S.X., Y.Z., P.Z., M.G., Y-C.L. and B.W. performed the human test. Y.-C.H. and Y.S. provided MRI-sequence support. Y.J., S.X. and C-J.L. performed the histology, IHC and flow cytometry. Y.J., S.X., G.H., B.W., Z.W., C-J.L., P.Z., M.G., C.L., Y-C.L. and R.B. critically read the manuscript. Y.J., G.H., Y-C.L., R.B. and S.X. contributed to manuscript writing, and Y.J. and R.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.B., Y.J. and G.H. have filed a patent application that describe aspects of this technology (2022100372579, China, 2022). The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 DCE-MRI measurements of cell cultures in vitro.
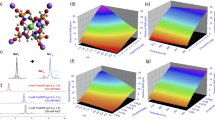
a: the customized chamber to collect cells and perform MRI measurements inside the bench-top MRI system. The cell collection in the MR tube and setup for MRI experiments followed the order from a-1 to a-4. b: An example of the IR-TSE raw images of U87MG at [CA] = 5 mM (upper) and 0 mM (lower). The ROI layer of cell pellet is illustrated with red rectangle. c: The longitudinal relaxation rate constants (R1 ≡ 1/T1) as a function of CA concentration in PBS at room temperature (23.5 ± 1 °C), n = (3, 3, 3, 3, 3, 3), the data is shown as mean (dots) +/- SEM. d, e: an example of the normalized IR-TSE signal (Supplementary Methods Section 1, equation (1)) of the U87MG cells (ROI-averaged) at CA concentration [CA] = 5 mM (d) and 0 mM (e) in which the blue circles and continuous red curves are the normalized IR-TSE data and the model fitting results with SS model, respectively.
Extended data Fig. 2 Typical AQP4 expression and distribution in C6 cell line following TMZ treatment.
a, Typical confocal microscopy images of AQP4 (red) and DAPI (blue) in C6 cell lines. Scale bar, 25 µm. b, c, Fluorescence colocalization analysis between nucleus and AQP4 by line profiles (the dotted, white lines in a) of staining intensity for AQP4 (red line) and nucleus (blue line).
Extended Data Fig. 3 Changes of AQP4, kio and other characters upon TMZ treatment on C6 cell line.
a-f: The changes in a, kio, n = (4, 6, 4, 6), p = 0.0099, p = 0.0006. b, AQP4 (rfu) / DAPI (rfu), n = (2, 5, 3, 3), p = 0.0328, p < 0.0001 c, migration length (normalized by the control groups), n = (5, 4, 8, 8), p = 0.0836, p = 0.0001, d, Ki-67 (rfu) / DAPI (rfu), n = (3, 6, 3, 6), p = 0.1041, p = 0.0133 e, cell proliferation speed, n = (6, 15, 3, 34, 13), p = 0.0505, p = 0.0001, and f, the SCCs fraction (OG+ cells/ total cells), n = (4, 3, 3, 3), p < 0.0001. Here, control group represents C6 cell lines incubated with DMSO only. In e, the results from AQP4 KO group is also shown. The data is shown as mean ± SEM. * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001, ns, Non-significant, From a to f, two-sided unpaired t-test between 3days and 7days in control, p = (0.3612, 0.7030, 0.9999, 0.3611, 0.7981, 0.1001). Here, the data points overlay the corresponding box.
Extended Data Fig. 4 Correlations between kio and proliferation speed (and SCCs) in U87MG cell line experiencing proliferation cycles.
a: The dynamic changes in cell counting (black, dashed circle), proliferation rate (black dots), and kio (red triangles) as a function of cell culture time. The data is shown as mean (dots) ± SEM (error bar), n = (9, 9, 8, 7, 11, 16, 7). Here the three proliferation phases were defined as (I) lag phase (0hr-48hr), (II) logarithmic growth phase (72hr-96hr), and (III) stationary and decline phase (from 120 hr). b: Linear correlation was observed between kio and proliferation rate. n = (9, 8, 7, 11, 16). c: Typical results of cell tracing with OG at different culture time. Scale bar: 50 µm. d: Statistics of OG+ fractions as a function of culture time (mean + /-SEM). From up to down p < 0.0001, p < 0.0001, p = 0.0016, p = 0.0005, p = 0.0004, ** p < 0.01, *** p < 0.001, **** p < 0.0001, n = (4, 4, 4, 5). Here, the data points overlay the corresponding box, two-sided unpaired t-test. e, Linear correlation was also observed between kio and OG+ fraction (that is, SCC fraction) In (b, e), areas between the two dotted lines reflect the 95% confidence interval in linear regression, the data is shown as mean (dots) ± SEM.
Extended Data Fig. 5 Optimization of DCE-MRI for precise kio measurements in vivo.
a: The implementation of MGE sequence in Water-exchange DCE-MRI eliminates potential T2* artifacts caused by the contrast agent at 7 T by fitting the MGE data (Supplementary Methods Section 3, equation 5) to obtain the purely T1-weighted signal S (TE = 0 ms). b: The original data with the shortest TE (2.8 ms) still shows T2*-induced signal attenuation. c: Monto Carlo simulations demonstrate that our optimized protocol (dual-bolus injection and optimized sequence settings) shows one-fold smaller standard deviation of kio estimation than the conventional scanning protocol (single-bolus injection, TR = 10 ms, FA = 10°. Std denotes the standard deviation). Box plot specifications: box bounds mean 25th and 75th percentile, center = 50th percentile, minima/maxima = center ± 1.5 × (75th percentile – 25th percentile), no whiskers shown. d-f: An example of a kio maone-foldone-foldp overlaid on T2-weighted image in the tumor region (d) and the model fittings of the DCE-MRI data with SS model for pixels located in the core (e, low kio = 1.6 s−1) and ring (f, high kio = 10.0 s−1) of the tumor. Here the raw data and the fitting results are shown as dots and continuous curves, respectively. Scale bar, 2 mm.
Extended Data Fig. 6 The kio map precisely reveals the intra-tumoral AQP4 distribution in each rat glioma model.
a-j: significant linear correlation between kio values and AQP4+ fractions was observed in each animal. Here, we used a series of concentric donut-shape ROIs to divide the tumor slice into six zones considering the ring-shape distribution of AQP4, as demonstrated in Fig. 5e. k. In the control group of TGN020 modulation (Fig. 6), the whole-tumor-averaged kio doesn’t show significant changes between the two days with the saline treatment. Paired two-sided, t-test, ns non-significant, p = 0.9588. The bar height and error bar width represent the mean and standard error of the mean, respectively. The data points overlay the corresponding box. n = 4.
Extended Data Fig. 7 A linear correlation is observed between kio and AQP4 expression in the rat orthotopic model of C6 glioma.
a, b: Typical examples of kio maps and AQP4 IHC results from two animals with small (a) and large kio (b) values. From up to down, they are the contrast-enhanced T1-weighted images (the position of tumor was illustrated with the white dashed circle), the kio maps overlaid on the T1-weighted images, and the typical AQP4 IHC results of the position pointed by the white arrows. MRI Scale bar, 2 mm; IHC Scale bar, 25 µm. c: A linear correlation is observed between the whole-tumor-averaged kio and of AQP4+ fractions in the seven rats of orthotopic glioma. The solid line reflects linear regression analysis and the two dashed curves denote 95% confidence intervals. n = 7.
Extended Data Fig. 8 The procedure of stereotactic biopsy in human glioma and the downstream analysis.
a: The placement of stereotactic frame from the frontal and lateral view. b: The illustration of trajectory for the biopsy entry point and the target on the MRI. c: The stereotactic biopsy platform. d, e: The view of aspiration side window cutting needle (d) and the acquired sample (e). f-i: examples of the downstream analysis for H&E, here, n = 35. Scale bar 50 µm. (f), IEM, here, n = 5. Scale bar 0.1 µm. (g), IHC, here, for AQP4, n = 45, for ZEB1, n = 10. Scale bar 50 µm. (h), and FACS (i) two biopsy points were obtained from one glioma patient, one sample for C6 TMZ 7 day. Scale bar 100 µm.
Extended Data Fig. 9 A special case of glioma patient and the sample-averaged statistics.
This is a recurrent glioblastoma patient who received radiofrequency ablation surgery with multiple needle tracts due to the large size of tumor region. In this special case, 10 biopsy samples were safely collected along the planned trajectory of radiofrequency ablation needles. a: Examples of kio maps and the positions to obtain the biopsy samples (white arrows). b: AQP4 IHC of the three biopsy points illustrated in a. Scale bar, 25 µm. c: A linear correlation is observed between kio and fractions of AQP4+ cells in the 10 stereotactic biopsy points from this patient. d: A linear correlation between sample-averaged kio and fractions of AQP4+ cells is still preserved in the 19 data points from 19 patients. Here, for the patient with multiple biopsies acquired, the averaged results from all biopsies of this patient was used as the representative biopsy result for this patient. The solid line reflects linear regression analysis and the two dashed curves denote 95% confidence intervals.
Extended Data Fig. 10 Comparison between conventional MRI and DCE-MRI-derived kio in predicting AQP4 distribution in human glioma.
Here the results using a random forest model evaluated with a five-fold cross validation method on the 45 stereotactic biopsy points were shown. a: the results of conventional MRI including contrast-enhanced T1-weighted imaging, T2-weighted imaging, apparent diffusion coefficient (ADC) and diffusion weighted imaging (DWI). b: the results of kio. c: the results of the combination of conventional MRI and kio. In a-c, the solid line reflects linear regression analysis between the predicted AQP4 and the observed AQP4. The two dashed curves denote 95% confidence intervals. R2 is the coefficient of determination. d: one example of the predicted AQP4 expression map in one glioma patient using kio only.
Supplementary information
Supplementary Information
Supplementary discussion, methods, figures, tables and references.
Source data
Source Data for Figs. 2–8 and Extended Data Figs. 1–10
Source data, statistics and unprocessed WBs.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jia, Y., Xu, S., Han, G. et al. Transmembrane water-efflux rate measured by magnetic resonance imaging as a biomarker of the expression of aquaporin-4 in gliomas. Nat. Biomed. Eng 7, 236–252 (2023). https://doi.org/10.1038/s41551-022-00960-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-022-00960-9
This article is cited by
-
MRI of transmembrane water efflux in gliomas
Nature Biomedical Engineering (2022)