Abstract
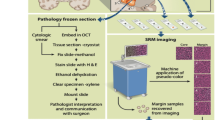
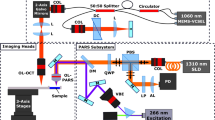
Conventional methods for intraoperative histopathologic diagnosis are labour- and time-intensive, and may delay decision-making during brain-tumour surgery. Stimulated Raman scattering (SRS) microscopy, a label-free optical process, has been shown to rapidly detect brain-tumour infiltration in fresh, unprocessed human tissues. Here, we demonstrate the first application of SRS microscopy in the operating room using a portable fibre-laser-based microscope and unprocessed specimens from 101 neurosurgical patients. We also introduce an image-processing method—stimulated Raman histology (SRH)—that leverages SRS images to create virtual haematoxylin-and-eosin-stained slides, revealing essential diagnostic features. In a simulation of intraoperative pathologic consultation in 30 patients, we found a remarkable concordance of SRH and conventional histology for predicting diagnosis (Cohen’s kappa, κ > 0.89), with accuracy exceeding 92%. We also built and validated a multilayer perceptron based on quantified SRH image attributes that predicts brain-tumour subtype with 90% accuracy. Our findings provide insight into how SRH can now be used to improve the surgical care of brain-tumour patients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Eisenhardt, L. & Cushing, H. Diagnosis of intracranial tumors by supravital technique. Am. J. Pathol. 6, 541–552 (1930).
Somerset, H. L. & Kleinschmidt-DeMasters, B. K. Approach to the intraoperative consultation for neurosurgical specimens. Adv. Anat. Pathol. 18, 446–449 (2011).
Sanai, N., Polley, M.-Y., McDermott, M. W., Parsa, A. T. & Berger, M. S. An extent of resection threshold for newly diagnosed glioblastomas. J. Neurosurg. 115, 3–8 (2011).
Freudiger, C. W. et al. Label-free biomedical imaging with high sensitivity by stimulated Raman scattering microscopy. Science 322, 1857–1861 (2008).
Lu, F. K. et al. Label-free neurosurgical pathology with stimulated Raman imaging. Cancer Res. 76, 3451–3462 (2016).
Ji, M. et al. Detection of human brain tumor infiltration with quantitative stimulated Raman scattering microscopy. Sci. Transl. Med. 7, 309ra163 (2015).
Ji, M. et al. Rapid, label-free detection of brain tumors with stimulated Raman scattering microscopy. Sci. Transl. Med. 5, 201ra119 (2013).
Freudiger, C. W. et al. Stimulated Raman scattering microscopy with a robust fibre laser source. Nat. Photon. 8, 153–159 (2014).
Freudiger, C. W. et al. Multicolored stain-free histopathology with coherent Raman imaging. Lab. Invest. 92, 1492–1502 (2012).
Bini, J. et al. Confocal mosaicing microscopy of human skin ex vivo: spectral analysis for digital staining to simulate histology-like appearance. J. Biomed. Opt. 16, 076008 (2011).
Dobbs, J. et al. Confocal fluorescence microscopy for rapid evaluation of invasive tumor cellularity of inflammatory breast carcinoma core needle biopsies. Breast Cancer Res. Tr. 149, 303–310 (2015).
Ozeki, Y. et al. High-speed molecular spectral imaging of tissue with stimulated Raman scattering. Nat. Photon. 6, 845–851 (2012).
Louis, D. N., Ohgaki, H., Wiestler, O. D. & Cavenee, W. K. (eds) World Health Organization Classification of Tumours of the Central Nervous System 4th edn (World Health Organization and International Agency for Research on Cancer, 2007).
Thompson, E. M. et al. Prognostic value of medulloblastoma extent of resection after accounting for molecular subgroup: a retrospective integrated clinical and molecular analysis. Lancet Oncol. 17, 484–495 (2016).
Orlov, N. et al. WND-CHARM: multi-purpose image classification using compound image transforms. Pattern Recogn. Lett. 29, 1684–1693 (2008).
Stummer, W. et al. Fluorescence-guided surgery with 5-aminolevulinic acid for resection of malignant glioma: a randomised controlled multicentre phase III trial. Lancet Oncol. 7, 392–401 (2006).
Eberlin, L. S. et al. Classifying human brain tumors by lipid imaging with mass spectrometry. Cancer Res. 72, 645–654 (2012).
Jermyn, M. et al. Intraoperative brain cancer detection with Raman spectroscopy in humans. Sci. Transl. Med. 7, 274ra219 (2015).
Camp, C. H. Jr et al. High-speed coherent Raman fingerprint imaging of biological tissues. Nat. Photon. 8, 627–634 (2014).
Evans, C. L. et al. Chemically-selective imaging of brain structures with CARS microscopy. Opt. Express. 15, 12076–12087 (2007).
Kut, C. et al. Detection of human brain cancer infiltration ex vivo and in vivo using quantitative optical coherence tomography. Sci. Transl. Med. 7, 292ra100 (2015).
Yang, Y. et al. Differential diagnosis of breast cancer using quantitative, label-free and molecular vibrational imaging. Biomed. Opt. Express 2, 2160–2174 (2011).
Gao, L. et al. Differential diagnosis of lung carcinoma with coherent anti-Stokes Raman scattering imaging. Arch. Pathol. Lab. Med. 136, 1502–1510 (2012).
Sanai, N. et al. Intraoperative confocal microscopy for brain tumors: a feasibility analysis in humans. Neurosurgery 68, ons282–ons290 (2011).
Barker, J., Hoogi, A., Depeursinge, A. & Rubin, D. L. Automated classification of brain tumor type in whole-slide digital pathology images using local representative tiles. Med. Image Anal. 30, 60–71 (2016).
Mousavi, H. S., Monga, V., Rao, G. & Rao, A. U. Automated discrimination of lower and higher grade gliomas based on histopathological image analysis. J. Pathol. Inform. 6, 15 (2015).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 20, 37–46 (1960).
Fleiss, J. L. & Cohen, J. The equivalence of weighted kappa and the intraclass correlation coefficient as measures of reliability. Educ. Psychol. Meas. 33, 613–619 (1973).
Curry, W. T. Jr, Carter, B. S. & Barker, F. G. II Racial, ethnic, and socioeconomic disparities in patient outcomes after craniotomy for tumor in adult patients in the United States, 1988–2004. Neurosurgery 66, 427–437; discussion 437–438 (2010).
Smith, E. R., Butler, W. E. & Barker, F. G. II Is there a ‘July phenomenon’ in pediatric neurosurgery at teaching hospitals? J. Neurosurg. 105, 169–176 (2006).
Brinjikji, W., Lanzino, G. & Cloft, H. J. Cerebrovascular complications and utilization of endovascular techniques following transsphenoidal resection of pituitary adenomas: a study of the Nationwide Inpatient Sample 2001–2010. Pituitary 17, 430–435 (2014).
Acknowledgements
The authors would like to thank H. Wagner for manuscript editing. Research reported in this publication was supported by the National Institute of Biomedical Imaging and Bioengineering (R01EB017254 to X.S.X. and D.A.O.), National Institute of Neurologic Disorders and Stroke (K08NS087118 to S.H.R.), the National Institutes of Health Director’s Transformative Research Award Program T-R01 (R01EB010244-01 to X.S.X.) and the National Cancer Institute of the National Institutes of Health (P30CA046592). This work was also supported by Fast Forward Medical Innovation, the University of Michigan—Michigan Translational Research and Commercialization for Life Sciences Program (U-M MTRAC) and the Michigan Institute for Clinical and Health Research (2UL1TR000433).
Author information
Authors and Affiliations
Contributions
D.A.O., B.P., Y.S.N., C.W.F., J.K.T., T.C.H. and S.C.-P. conceived the study, designed the experiments and wrote the paper; they were assisted by M.G. and X.S.X., who provided guidance on the study design. D.A.O., S.L. and M.G. performed the SRH imaging of all specimens. C.W.F. and J.K.T. built the SRS microscope. B.P., Y.S.N., J.B. and T.D.J. analysed the data. S.C.-P., K.A.M., S.H.R., M.S., S.V., A.P.L. and A.F.-H. interpreted microscopic images and revised the manuscript. T.D.J., D.A.W. and Y.S.N. performed the statistical analyses. D.A.O., S.L.H.-J., H.J.L.G., J.A.H., C.O.M. and O.S. provided surgical specimens for imaging. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
X.S.X. and D.A.O. are advisers and shareholders of Invenio Imaging, Inc., a company developing SRS microscopy systems. C.W.F. and J.K.T. are employees and shareholders of Invenio Imaging, Inc.
Supplementary information
Supplementary Information
Supplementary figures and tables. (PDF 9394 kb)
Rights and permissions
About this article
Cite this article
Orringer, D., Pandian, B., Niknafs, Y. et al. Rapid intraoperative histology of unprocessed surgical specimens via fibre-laser-based stimulated Raman scattering microscopy. Nat Biomed Eng 1, 0027 (2017). https://doi.org/10.1038/s41551-016-0027
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41551-016-0027
This article is cited by
-
Characterization of lamina propria remodeling in pediatric eosinophilic esophagitis using second harmonic generation microscopy
Translational Medicine Communications (2024)
-
Multi-molecular hyperspectral PRM-SRS microscopy
Nature Communications (2024)
-
Virtual histological staining of unlabeled autopsy tissue
Nature Communications (2024)
-
Development and prospective validation of an artificial intelligence-based smartphone app for rapid intraoperative pituitary adenoma identification
Communications Medicine (2024)
-
Mechanical characteristics of glioblastoma and peritumoral tumor-free human brain tissue
Acta Neurochirurgica (2024)