Abstract
The fall armyworm (FAW) poses a significant threat to global crop production. Here we showed that overexpression of jasmonate ZIM-domain (JAZ) protein GhJAZ24 confers resistance to cotton bollworm and FAW, while also causing sterility in transgenic cotton by recruiting TOPLESS and histone deacetylase 6. We identified the NGR motif of GhJAZ24 that recognizes and binds the aminopeptidase N receptor, enabling GhJAZ24 to enter cells and disrupt histone deacetylase 3, leading to cell death. To overcome plant sterility associated with GhJAZ24 overexpression, we developed iJAZ (i, induced), an approach involving damage-induced expression and a switch from intracellular to extracellular localization of GhJAZ24. iJAZ transgenic cotton maintained fertility and showed insecticidal activity against cotton bollworm and FAW. In addition, iJAZ transgenic rice, maize and tobacco plants showed insecticidal activity against their lepidopteran pests, resulting in an iJAZ-based approach for generating alternative insecticidal proteins with distinctive mechanisms of action, thus holding immense potential for future crop engineering.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available in the article, Extended Data figures and Supplementary Information, or are available from the corresponding author upon reasonable request. The raw sequence data of SfAPN1-13 and SfHDAC3 used in this study are available in the GenBank under the accession codes OM305006, OM305007, OM305008, OM305009, OM305010, OM305011, OM305012, OM305013, OM305014, OM305015, OM305016, OM305017, OM305018 and OM305019. MS proteomics data produced for this study were deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository (dataset identifier PXD039603). Source data are provided with this paper.
References
Gui, F. et al. Genomic and transcriptomic analysis unveils population evolution and development of pesticide resistance in fall armyworm Spodoptera frugiperda. Protein Cell 13, 513–531 (2020).
Li, Y. H., Wang, Z. Y. & Romeis, J. Managing the invasive fall armyworm through biotech crops: a Chinese perspective. Trends Biotechnol. 39, 105–108 (2021).
Lyu, B., Wang, S., Wyckhuys, K. A. G. & Liu, Z. Biological pest control protects pollinators. Science 380, 251 (2023).
Pardo-Lopez, L., Soberon, M. & Bravo, A. Bacillus thuringiensis insecticidal three-domain Cry toxins: mode of action, insect resistance and consequences for crop protection. FEMS Microbiol. Rev. 37, 3–22 (2013).
Byrne, M. J. et al. Cryo-EM structures of an insecticidal Bt toxin reveal its mechanism of action on the membrane. Nat. Commun. 12, 2791 (2021).
Ren, M. Z., Zafar, M. M., Mo, H. J., Yang, Z. E. & Li, F. G. Fighting against fall armyworm by using multiple genes pyramiding and silencing (MGPS) technology. Sci. China Life Sci. 62, 1703–1706 (2019).
Guo, Z. et al. MAPK-dependent hormonal signaling plasticity contributes to overcoming Bacillus thuringiensis toxin action in an insect host. Nat. Commun. 11, 3003 (2020).
Tay, W. T., Meagher, R. L. Jr., Czepak, C. & Groot, A. T. Spodoptera frugiperda: ecology, evolution, and management options of an invasive species. Annu. Rev. Entomol. 68, 299–317 (2023).
Yao, C. et al. Stemborer-induced rice plant volatiles boost direct and indirect resistance in neighboring plants. New Phytol. 237, 2375–2387 (2023).
Zhou, W. et al. A jasmonate signaling network activates root stem cells and promotes regeneration. Cell 177, 942–956 (2019).
Zhang, G. et al. Jasmonate-mediated wound signalling promotes plant regeneration. Nat. Plants 5, 491–497 (2019).
Gao, Y. Q. et al. Ricca’s factors as mobile proteinaceous effectors of electrical signaling. Cell 186, 1337–1351 (2023).
Garrido-Bigotes, A., Valenzuela-Riffo, F. & Figueroa, C. R. Evolutionary analysis of JAZ proteins in plants: an approach in search of the ancestral sequence. Int. J. Mol. Sci. 20, 5060 (2019).
Zhang, B. et al. Gnawing pressure led to the expansion of JAZ genes in angiosperms. Int. J. Biol. Macromol. 230, 123165 (2023).
Thireault, C. et al. Repression of jasmonate signaling by a non-TIFY JAZ protein in Arabidopsis. Plant J. 82, 669–679 (2015).
Shyu, C. et al. JAZ8 lacks a canonical degron and has an EAR motif that mediates transcriptional repression of jasmonate responses in Arabidopsis. Plant Cell 24, 536–550 (2012).
Li, Z. et al. JASMONATE-ZIM DOMAIN proteins engage polycomb chromatin modifiers to modulate jasmonate signaling in Arabidopsis. Mol. Plant 14, 732–747 (2021).
He, K. et al. PHOSPHATE STARVATION RESPONSE1 (PHR1) interacts with JASMONATE ZIM-DOMAIN (JAZ) and MYC2 to modulate phosphate deficiency-induced jasmonate signaling in Arabidopsis. Plant Cell 35, 2132–2156 (2023).
Weng, L. et al. TGF-beta1/SMAD3 regulates programmed cell death 5 that suppresses cardiac fibrosis post-myocardial infarction by inhibiting HDAC3. Circ. Res. 133, 1–15 (2023).
Chen, X. et al. A VEL3 histone deacetylase complex establishes a maternal epigenetic state controlling progeny seed dormancy. Nat. Commun. 14, 2220 (2023).
Wu, S. E. et al. Microbiota-derived metabolite promotes HDAC3 activity in the gut. Nature 586, 108–112 (2020).
Gaddelapati, S. C., Albishi, N. M., Dhandapani, R. K. & Palli, S. R. Juvenile hormone-induced histone deacetylase 3 suppresses apoptosis to maintain larval midgut in the yellow fever mosquito. Proc. Natl Acad. Sci. USA 119, e2118871119 (2022).
George, S. & Palli, S. R. Histone deacetylase 3 is required for development and metamorphosis in the red flour beetle, Tribolium castaneum. BMC Genomics 21, 420 (2020).
Lv, W. W., Wei, H. M., Wang, D. L., Ni, J. Q. & Sun, F. L. Depletion of histone deacetylase 3 antagonizes PI3K-mediated overgrowth of Drosophila organs through the acetylation of histone H4 at lysine 16. J. Cell Sci. 125, 5369–5378 (2012).
Chang, H. et al. HDAC3 knockdown dysregulates juvenile hormone and apoptosis-related genes in Helicoverpa armigera. Int. J. Mol. Sci. 23, 14820 (2022).
Song, Y. et al. BIN2 negatively regulates plant defence against Verticillium dahliae in Arabidopsis and cotton. Plant Biotechnol. J. 19, 2097–2112 (2021).
Zhao, G. et al. Gossypium hirsutum salt tolerance is enhanced by overexpression of G. arboreum JAZ1. Front. Bioeng. Biotechnol. 8, 157 (2020).
Zhang, S. et al. The control of carpel determinacy pathway leads to sex determination in cucurbits. Science 378, 543–549 (2022).
An, C. et al. Regulation of jasmonate signaling by reversible acetylation of TOPLESS in Arabidopsis. Mol. Plant 15, 1329–1346 (2022).
Pauwels, L. & Goossens, A. The JAZ proteins: a crucial interface in the jasmonate signaling cascade. Plant Cell 23, 3089–3100 (2011).
Borg, M. et al. An EAR-dependent regulatory module promotes male germ cell division and sperm fertility in Arabidopsis. Plant Cell 26, 2098–2113 (2014).
Wei, B. et al. The molecular mechanism of sporocyteless/nozzle in controlling Arabidopsis ovule development. Cell Res. 25, 121–134 (2015).
Ren, L. et al. OsSPL regulates meiotic fate acquisition in rice. New Phytol. 218, 789–803 (2018).
Krogan, N. T., Hogan, K. & Long, J. A. APETALA2 negatively regulates multiple floral organ identity genes in Arabidopsis by recruiting the co-repressor TOPLESS and the histone deacetylase HDA19. Development 139, 4180–4190 (2012).
Gryffroy, L. et al. Rhizogenic Agrobacterium protein RolB interacts with the TOPLESS repressor proteins to reprogram plant immunity and development. Proc. Natl Acad. Sci. USA 120, e2210300120 (2023).
Deng, H. et al. SlERF.F12 modulates the transition to ripening in tomato fruit by recruiting the co-repressor TOPLESS and histone deacetylases to repress key ripening genes. Plant Cell 34, 1250–1272 (2022).
Liu, C., Yang, Y., Chen, L., Lin, Y. L. & Li, F. A unified mechanism for aminopeptidase N-based tumor cell motility and tumor-homing therapy. J. Biol. Chem. 289, 34520–34529 (2014).
Curnis, F. et al. Enhancement of tumor necrosis factor alpha antitumor immunotherapeutic properties by targeted delivery to aminopeptidase N (CD13). Nat. Biotechnol. 18, 1185–1190 (2000).
Sun, D. et al. A versatile contribution of both aminopeptidases N and ABC transporters to Bt Cry1Ac toxicity in the diamondback moth. BMC Biol. 20, 33 (2022).
Cooper, M. A., Carroll, J., Travis, E. R., Williams, D. H. & Ellar, D. J. Bacillus thuringiensis Cry1Ac toxin interaction with Manduca sexta aminopeptidase N in a model membrane environment. Biochem. J. 333, 677–683 (1998).
Zhang, S. P. et al. Mutation of an aminopeptidase N gene is associated with Helicoverpa armigera resistance to Bacillus thuringiensis Cry1Ac toxin. Insect Biochem. Mol. Biol. 39, 421–429 (2009).
Emmett, M. J. & Lazar, M. A. Integrative regulation of physiology by histone deacetylase 3. Nat. Rev. Mol. Cell Biol. 20, 102–115 (2019).
Tsai, K. K. et al. Screening of organoids derived from patients with breast cancer implicates the repressor NCOR2 in cytotoxic stress response and antitumor immunity. Nat. Cancer 3, 734–752 (2022).
Martin-Arevalillo, R. et al. Structure of the Arabidopsis TOPLESS corepressor provides insight into the evolution of transcriptional repression. Proc. Natl Acad. Sci. USA 114, 8107–8112 (2017).
Li, X. F. et al. Two ubiquitin-associated ER proteins interact with COPT copper transporters and modulate their accumulation. Plant Physiol. 188, 673 (2022). -673.
Thagun, C., Chuah, J. A. & Numata, K. Targeted gene delivery into various plastids mediated by clustered cell-penetrating and chloroplast-targeting peptides. Adv. Sci. 6, 1902064 (2019).
Ge, X. et al. Efficient genotype-independent cotton genetic transformation and genome editing. J. Integr. Plant Biol. 65, 907–917 (2023).
Gassmann, A. J. & Reisig, D. D. Management of insect pests with Bt crops in the United States. Annu. Rev. Entomol. 68, 31–49 (2023).
Alenghat, T. et al. Histone deacetylase 3 coordinates commensal-bacteria-dependent intestinal homeostasis. Nature 504, 153–157 (2013).
Kuang, Z. et al. The intestinal microbiota programs diurnal rhythms in host metabolism through histone deacetylase 3. Science 365, 1428–1434 (2019).
Ahmed, S. M. H. et al. Fitness trade-offs incurred by ovary-to-gut steroid signalling in Drosophila. Nature 584, 415–419 (2020).
Lin, Y. et al. Characterization of the physiological, histopathological, and gene expression alterations in Spodoptera frugiperda larval midguts affected by toosendanin exposure. Pestic. Biochem. Physiol. 195, 105537 (2023).
Yang, Y., Chaffin, T. A., Ahkami, A. H., Blumwald, E. & Stewart, C. N. Jr. Plant synthetic biology innovations for biofuels and bioproducts. Trends Biotechnol. 40, 1454–1468 (2022).
Mao, Y. B. et al. Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nat. Biotechnol. 25, 1307–1313 (2007).
Wen, N. et al. The overexpression of insect endogenous microRNA in transgenic rice inhibits the pupation of Chilo suppressalis and Cnaphalocrocis medinalis. Pest Manag. Sci. 77, 3990–3999 (2021).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Watson, P. J., Fairall, L., Santos, G. M. & Schwabe, J. W. Structure of HDAC3 bound to co-repressor and inositol tetraphosphate. Nature 481, 335–340 (2012).
Xu, D. et al. Export of defensive glucosinolates is key for their accumulation in seeds. Nature 617, 132–138 (2023).
Wang, C. et al. Apolipoprotein C-II induces EMT to promote gastric cancer peritoneal metastasis via PI3K/AKT/mTOR pathway. Clin. Transl. Med. 11, e522 (2021).
He, J. et al. An R2R3 MYB transcription factor confers brown planthopper resistance by regulating the phenylalanine ammonia-lyase pathway in rice. Proc. Natl Acad. Sci. USA 117, 271–277 (2020).
Ruan, W. et al. Two RING-finger ubiquitin E3 ligases regulate the degradation of SPX4, an internal phosphate sensor, for phosphate homeostasis and signaling in rice. Mol. Plant 12, 1060–1074 (2019).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Li, W. et al. NADPH levels affect cellular epigenetic state by inhibiting HDAC3-Ncor complex. Nat. Metab. 3, 75–89 (2021).
Van der Does, D. et al. Salicylic acid suppresses jasmonic acid signaling downstream of SCFCOI1-JAZ by targeting GCC promoter motifs via transcription factor ORA59. Plant Cell 25, 744–761 (2013).
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572 (2007).
Zhao, F. F. et al. Visualizing the essential role of complete virion assembly machinery in efficient hepatitis C virus cell-to-cell transmission by a viral infection-activated split-intein-mediated reporter system. J. Virol. 91, e01720–16 (2017).
Kurusu, T. et al. Plasma membrane protein OsMCA1 is involved in regulation of hypo-osmotic shock-induced Ca2+ influx and modulates generation of reactive oxygen species in cultured rice cells. BMC Plant Biol. 12, 1–15 (2012).
Zhao, W. N., Zheng, S. S. & Ling, H. Q. An efficient regeneration system and Agrobacterium-mediated transformation of Chinese upland rice cultivar Handao297. Plant Cell Tiss. Org. 106, 475–483 (2011).
Sunilkumar, G., Vijayachandra, K. & Veluthambi, K. Preincubation of cut tobacco leaf explants promotes mediated transformation by increasing gene induction. Plant Sci. 141, 51–58 (1999).
Du, D. X., Jin, R. C., Guo, J. J. & Zhang, F. D. Infection of embryonic callus with Agrobacterium enables high-speed transformation of maize. Int. J. Mol. Sci. 20, 279 (2019).
Zhang, J. et al. Pest control. Full crop protection from an insect pest by expression of long double-stranded RNAs in plastids. Science 347, 991–994 (2015).
Jiang, Y. D. & Hsieh, J. HDAC3 controls gap 2/mitosis progression in adult neural stem/progenitor cells by regulating CDK1 levels. Proc. Natl Acad. Sci. USA 111, 13541–13546 (2014).
Ishii, S., Kurasawa, Y., Wong, J. M. & Yu-Lee, L. Y. Histone deacetylase 3 localizes to the mitotic spindle and is required for kinetochore–microtubule attachment. Proc. Natl Acad. Sci. USA 105, 4179–4184 (2008).
Acknowledgements
This research was supported by the National Key R&D Program of China number 2020YFA0908000 (M.R. and H.M.) and number 2023YFE0199400 (M.R.); the National Science Foundation of China number 31621005 (F.L.), number U23A20182 (F.L.), number U1804231 (F.L.) and number 31972469 (M.R.); the Key project at central government level number 2060302 (M.R.); the Sichuan Science and Technology Program number 2023YFQ0100 (M.R.), number 2023ZYD0089 (M.R.), number 2022YFH0054 (M.R.) and number 2023JDG0028 (M.R.); the Local Financial Funds of Chengdu National Agricultural Science and Technology Center number NASC2023TD08 (M.R.), number NASC2021ST08 (M.R.), number NASC2021PC04 (M.R.) and number NASC2022KR07 (M.R.); the Central Public-interest Scientific Institution Basal Research Fund number S2023011 (F.L.); the Science and Technology Innovation Project of the Chinese Academy of Agricultural Sciences number 34-IUA-02 (M.R.); and the Innovation Program of Chinese Academy of Agricultural Sciences (F.L.).
Author information
Authors and Affiliations
Contributions
F.L., M.R., X.C. and H.M. conceived and designed the project. H.M., H.C., X.J., Z.X., G.R. and G.Z. performed experiments. L.F., M.R., H.M., X.J., J.F.W., G.H., X.L. and H.C. analysed data. F.L., M.R., G.H., J.F.W., L.F. and H.M. wrote the manuscript, with contributions from all authors. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks José-Antonio Daròs and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The conservatism of JAZ families is diverse.
a-c, 35 S::GhJAZ24 transgenic cotton plants were identified in the genomes (a), transcription at the mRNA level (b), and expression at the protein level (c). n = 3 biologically independent repeats (b). d, Subcellular localisation of GhJAZ24 protein in tobacco leaves. e, Multiple sequence alignment of the Jas degron (loop region and α-helix region), EAR motif (LxLxL peptide, indicated by blue boxes) and TIFYNGR region within the ZIM motif (indicated by red boxes) from JAZs of cotton and the canonical AtJAZ1 and AtJAZ7. Red lines indicate the structural features within the Jas motif that physically associate with COI1 and JA-Ile. f, GhJAZ24 lacks LPIAR motif and thus is unable to associate strongly with COI1 in the presence of JA-Ile (20 μM). g, Sequence alignment of HDA6 in Gossypium hirsutum (GhHDA6) and A. thaliana (AtHDA6). H134, H135, G143, F144, D170, H172, D259, L266, G296, and Y298 sites of HsHDAC3 (Homo sapiens, Gene ID: NP_001341968.1) were active sites, which included a Zn binding site, lipophilic tube, and foot pocket. D170, H172, and D259 sites of HsHDAC3 included Zn binding sites (ion binding sites), three residue positions, and an active center containing catalytic Zn ion that coordinated two aspartates and one histidine. h, GhJAZ24 lacks LPIAR motif and thus is stabilized against jasmonate (JA)-mediated degradation. In resting state, JAZ proteins mediate repression of JA responses by recruiting the transcription corepressor TPL either directly (EAR-containing JAZ proteins) or via EAR motif of NINJA (canonical JAZ proteins). In activation state, JA-Ile initiates the signalling cascade by promoting interaction of LPIAR motif of canonical JAZ proteins with the SCFCOI1, canonical JAZ proteins are degraded subsequently via the 26 S proteasome pathway, and canonical JAZ-mediated repression is relieved (Actication). GhJAZ24 is stabilized against JA-mediated degradation due to the absence of LPIAR motif, and thus maintains its transcriptional repression. Data are mean ± s.e.m. *P < 0.05; **P < 0.01 (two-tailed Student’s t-test). Scale bars, 20 μm (d).
Extended Data Fig. 2 Phylogenetic tree with 404 orthologs from 25 species identified by PhyloGenes to show divergence times.
Divergence times were estimated using Timetree and are indicated by blue dots at the internodes with 95% highest posterior density (HPD). *, the following species substitutions were used: Marchantia polymorpha (replaced with Marchantia paleacea), Ananas comosus (replaced with Brocchinia acuminata), Oryza sativa (replaced with Oryza longistaminata), Zea mays (replaced with Zea diploperennis). Abbreviations: EDI, Ediacaran; C, Cambrian; O, Ordovician; D, Devonian; MIS, Mississippian; P, Permian; T, Triassic; J, Jurassic; K, Cretaceous; Pg, Paleogene; MYA, million years ago. The number of NGR-containing JAZ genes and the total number of JAZ genes in 25 species are shown in parentheses.
Extended Data Fig. 3 NGR-containing JAZ proteins exhibit cytotoxicity to Sf9 cells.
a-i, SDS-PAGE (left) and western blot (right) analysis of the heterologous expression of recombinant GhJAZ24:His, GhJAZ24LGK:His, GhJAZ24:GFP, GhJAZ8:His, GhJAZ13:His, GhJAZ14:His, GhJAZ27:His, GhJAZ28:His, DzJAZ (Durio zibethinus JAZ, XP_022720271.1) proteins in E. coli cells. BSA, 2.00 µg. j, Fluorescence microscopy images 24 h after treatment of Sf9 cells with 1 μg/mL JAZ proteins. k, NGR-containing JAZ protein (1 μg/mL) from Durio zibethinus exhibited cytotoxicity to Sf9 cells (n = 3 biologically independent repeats). l, Apoptosis of Sf9 cells treated with NGR-containing JAZ proteins at IC50 (n = 3 biologically independent repeats). m, Cell cycle data of Sf9 cells treated with JAZ proteins at IC50 (n = 3 biologically independent repeats). n-r, IC50 at 24 h after treatment of Sf9 cells with NGR-containing JAZ proteins (n = 3 biologically independent repeats). Data are mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 (k,l, two-tailed Student’s t-test; m, one-way ANOVA, Tukey’s HSD test). Scale bars, 100 μm (j).
Extended Data Fig. 4 Expression profile of SfAPNs genes and integrity analysis of GhJAZ24 protein.
a, SfAPNs show high expression in different developmental stages and tissues. L1-6, first- to sixth-instar larvae; PP, prepupae; P, pupae; MA, male adults; FA, female adults; EG, eggs. H, head; I, integument; M, midgut; T, Malpighian tubules. For each gene, the expression fold changes are color-coded according to the gradient red indicate high expression. The relative expression of each gene in the developmental stage L1 was set to 1. Data are mean ± s.e.m. n = 3 biologically independent repeats. b, His and GFP tags were arranged respectively on the N- and C- terminal of GhJAZ24 to detect the integrity of GhJAZ24 after entry into Sf9 cells. Cells treated with 0.5 μmol/L proteins were detected by western blot after 24 h treatment.
Extended Data Fig. 5 GhJAZ24 interacts with candidate nuclear targets.
a-d, Y2H (a), CoIP (b,c, b for SfSTK and GhJAZ24, c for SfLamin-C and GhJAZ24), and LCI (d) experiments verified binding of GhJAZ24 to the candidate targets. e, Tandem mass spectra of peptides identified by mass spectrometry. The peptide can be specifically matched to SfHDAC3 (Sfru02874) in the FAW database (ZJU_Sfru_1.0). f, Spatiotemporal expression of SfHDAC3 in different tissues and developmental stages of FAW (n = 3 biologically independent repeats). Data are mean ± s.e.m. g, Sequence alignment of HDAC3/HDA6 proteins in FAW, Homo sapiens (HsHDAC3, Gene ID: NP_001341968.1), G. hirsutum (GhHDA6, Gene ID: Gh_D03G0986.1), Zea mays (ZmHDA102, Gene ID: NP_001105077.1), and Oryza sativa (OsHDA9, Gene ID: XP_015635867.1). H134, H135, G143, F144, D170, H172, D259, L266, G296, and Y298 sites of HsHDAC3 were active sites, which included a Zn binding site, lipophilic tube, and foot pocket. D170, H172, and D259 sites of HsHDAC3 included Zn binding sites (ion binding sites), three residue positions, and an active center containing catalytic Zn ion that coordinated two aspartates and one histidine. Gene ID represents the gene accession number of the GenBank database (http://www.ncbi.nlm.nih.gov/).
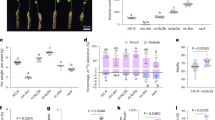
Extended Data Fig. 6 iJAZ module confers localization switch, inducible expression of GhJAZ24 and plants resistance to insects.
a, Intracellular-to-extracellular localization switch of GhJAZ24 in cotton protoplasts (row 1) and rice protoplasts (rows 2 to 4). b, Puncture damage to iJAZ-cotton plants induced strong expression of GhJAZ24 proteins by fluorescence detection based on the GFP fusion. c, Fluorescence imaging of the GhJAZ24: GFP and GhJAZ24: mCherry fusion proteins in iJAZ rice leaves. d,e,g, Bioassay for iJAZ- rice challenged by FAW (d,e) or rice leaffolder (g) pests. WT or 35 S::Cry1Ac transgenic rice set as control. f, Relative weight of rice plants with FAW larvae infection (n = 3 independent experiments). The relative weight of rice plants was calculated by Image J software. h, Confocal microscopy images of the midgut tissue of FAW fed with iJAZ-rice leaves. TUNEL: in light microscope images, nuclei were stained bluish-purple and apoptotic cells were stained brown. mCherry: in fluorescence microscope images, nuclear signals appeared blue and mCherry fluorescence signals appeared red. i, HDAC3 enzyme activity decreased after feeding on iJAZ rice in FAW larvae (n = 3 biologically independent repeats). Data are mean ± s.e.m. *P < 0.05 (two-tailed Student’s t-test). Scale bars, 10 μm (a), 10 μm (c), 1 mm (g), 300 μm (h, HE, mCherry), 200 μm (h, TUNEL, TUNEL-IF).
Extended Data Fig. 7 iJAZ module confers maize and tobacco plants resistance to FAW.
a,b, Puncture damage to iJAZ- maize and tobacco plants induced strong expression of GhJAZ24 proteins by fluorescence detection based on the GFP fusion. c,f, Bioassay for T2 generation iJAZ- maize (c) and tobacco (f) challenged by FAW pests. Nine second-instar FAW larvae per replicate were fed with freshly cut maize and tobacco leaves for 48 h and monitored daily. d, Representative FAW larval morphologies at 48 h after feeding with iJAZ maize leaves. e, The mortality of FAW larvae (n = 3 independent experiments). g, The weight of FAW larvae at 48 h after feeding with iJAZ tobacco leaves (n = 3 independent experiments). h, The weight of tobacco leaves at 48 h after feeding by FAW larvae (n = 3 independent experiments). Data are mean ± s.e.m. *P < 0.05, **P < 0.01, n.s., no significant difference (two-tailed Student’s t-test). Scale bars, 1 cm (c,d,f).
Extended Data Fig. 8 The entry receptor APN4 of GhJAZ24 typically exists in lepidopteran pests.
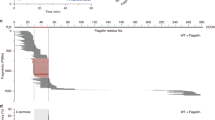
a, Maximum-likelihood inferred tree (s.d. <0.01, optimal log-likelihood value (-112,454.229) from full-length APN protein sequences found in the NCBI database. The tree was constructed with 153 APN sequences. The scale bar corresponds to 0.6 substitutions per nucleotide position. Purple-, pink-, orange-, and blue-colored branches indicate sequences of Bombycoidea, Noctuoidea, Pyraloidea, and Yponomeutoidea, respectively. All sequences are labelled by species name and accession number. b, The number of 153 APN sequences was summarized for each genus. c, WebLogo plots highlight amino acid conservation in the NHP motif in APN proteins. The 11 amino acid residues of the NHP motif in human APN are shown in pink. These orthologous proteins typically exist in lepidopteran pests, as follows: PxAPN4a (Plutella xylostella, Gene ID: MG873050), PxAPN4b (Plutella xylostella, Gene ID: MG873051), AjAPN4 (Achaea Janata, Gene ID: ABH07377), BmAPN4 (Bombyx mori, Gene ID: XP_012552709), CmAPN4 (Cnaphalocrocis medinalis, Gene ID: ADZ05468), CsAPN4 (Chilo suppressalis, Gene ID: ADZ57273), HaAPN4 (Helicoverpa armigera, Gene ID: AAP37950), HpAPN4 (Helicoverpa punctigera, Gene ID: AAF37559), HvAPN4 (Heliothis virescens, Gene ID: AAK58066), LdAPN4 (Lymantria dispar, Gene ID: AAL26894), MsAPN4 (Manduca sexta, Gene ID: AAM18718), OfAPN4 (Ostrinia furnacalis, Gene ID: ACB87202), OnAPN4 (Ostrinia nubilalis, Gene ID: ACV74256), PmAPN4 (Papilio machaon, Gene ID: XP_014370422), PpAPN4 (Papilio polytes, Gene ID: XP_013142032), PrAPN4 (Pieris rapae, Gene ID: XP_022128349), PxuAPN4 (Papilio xuthus, Gene ID: XP_013171158), SeAPN4 (Spodoptera exigua, Gene ID: AAP44967), SlAPN4 (Spodoptera litura, Gene ID: AAK69605), TnAPN4 (Trichoplusia ni, Gene ID: AAX39866). Gene ID represents the gene accession number of the GenBank database (http://www.ncbi.nlm.nih.gov/).
Extended Data Fig. 9 The nuclear target HDAC3 of GhJAZ24 is conserved in insect species.
a, Maximum-likelihood inferred tree (s.d. <0.01, optimal log-likelihood value (-35,556.894) from full-length HDAC3 protein sequences found in NCBI database. The tree was constructed with 465 HDAC3 sequences from the NCBI database. The scale bar corresponds to 1.0 substitutions per nucleotide position. All sequences are labelled by species name and accession number. b, WebLogo plots highlight amino acid conservation in HDAC3 proteins in other insect pest species. The 10 amino acid residues in HDAC3 were shown in pink word. D171, H173, and D260 sites of SfHDAC3 were the Zn binding sites. These orthologous proteins typically exist in lepidopteran pests, as follows: SlHDAC3 (Spodoptera litura, Gene ID: XP_022831573.1), HaHDAC3 (Helicoverpa armigera, Gene ID: XP_021195587.1), TnHDAC3 (Trichoplusia ni, Gene ID: XP_026740888.1), HkHDAC3 (Hyposmocoma kahamanoa, Gene ID: XP_026333196.1), GmHDAC3 (Galleria mellonella, Gene ID: XP_026754881.1), MhHDAC3 (Maniola hyperantus, Gene ID: XP_034827651.1), AaHDAC3 (Aricia agestis, Gene ID: XP_041972205.1), MsHDAC3 (Manduca sexta, Gene ID: KAG6455343.1), BiHDAC3 (Brenthis ino, Gene ID: CAH0713927.1), ApHDAC3 (Arctia plantaginis, Gene ID: CAB3219866.1), CsHDAC3 (Chilo suppressalis, Gene ID: RVE44174.1), PaHDAC3 (Pararge aegeria, Gene ID: XP_039752448.1), VtHDAC3 (Vanessa tameamea, Gene ID: XP_026489054.1), BaHDAC3 (Bicyclus anynana, Gene ID: XP_023947594.1), AtHDAC3 (Amyelois transitella, Gene ID: XP_013193311.1), McHDAC3 (Melitaea cinxia, Gene ID: XP_045454150.1), LsHDAC3 (Leptidea sinapis, Gene ID: VVD00622.1), PrHDAC3 (Pieris rapae, Gene ID: XP_022129510.1), PbHDAC3 (Pieris brassicae, Gene ID: XP_045518296.1). Gene ID represents the gene accession number of the GenBank database (http://www.ncbi.nlm.nih.gov/).
Extended Data Fig. 10 Transcript profile of GhJAZs in cotton and feeding preference of FAW on different crop leaves.
a, Expression profiles of JAZ genes under FAW stress in cotton (n = 3 biologically independent repeats). The green arrow indicates the identity of the Jas degron (loop region and α-helix region, Extended Data Fig. 1e) from high to low among the 30 JAZ members. Gene expression in the wounded plants (Ctrl) was set to 1. b, Feeding choice of FAW on maize, rice, and cotton leaves. Biological parameters of FAW on three different hosts, maize (the primary host plants), rice (the alternate host plants) and cotton (a non-preferred host plants) were evaluated. Ten second-instar FAW larvae per replicate were fed with 1 g of maize, rice, or cotton leaves for 24 h. Scale bars, 1 cm. c,d, Fresh weight of remaining maize, rice, and cotton leaves fed by FAW mixedly (c) or separately (d) (n = 3 independent experiments). e, The activities of digestive enzymes in the FAW larval midguts when fed with WT, or iJAZ tobacco leaves for 48 h (n = 9 biologically independent repeats). Data are mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; n.s., no significant difference (a,c,d, one-way ANOVA, Tukey’s HSD test; e, two-tailed Student’s t-test).
Supplementary information
Supplementary Information
Supplementary Figures 1–5, Supplementary Tables 1–9 and Supplementary Sequence 1.
Supplementary Data
Source data for Supplementary Figs. 2 and 3.
Source data
Source Data Fig. 1
Unprocessed gels and statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Unprocessed gels and statistical source data.
Source Data Fig. 4
Unprocessed gels and statistical source data.
Source Data Fig. 5
Unprocessed gels and statistical source data.
Source Data Extended Data Figs. 1, 3, 4 and 5
Unprocessed gels and statistical source data.
Source Data Extended Data Fig. 6, 7 and 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mo, H., Chang, H., Zhao, G. et al. iJAZ-based approach to engineer lepidopteran pest resistance in multiple crop species. Nat. Plants (2024). https://doi.org/10.1038/s41477-024-01682-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41477-024-01682-3