Abstract
Development of plant organs is a highly organized process. In Arabidopsis, proper root development requires that distinct cell types and tissue layers are specified and formed in a restricted manner in space and over time. Despite its importance, genetic controls underlying such regularity remain elusive. Here we found that WIP genes expressed in the embryo and suspensor functionally oppose those expressed in the surrounding maternal tissues to orchestrate cell division orientation and cell fate specification in the embryonic root, thereby promoting regular root formation. The maternal WIPs act non-cell autonomously to repress root cell fate specification through SIMILAR TO RADICAL-INDUCED CELL DEATH ONE (SRO) family members. When losing all WIPs, root cells divide irregularly in the early embryo, but this barely alters their fate specification and the morphology of post-embryonic roots. Our results reveal cross-communication between the embryonic and maternal WIPs in controlling root development.
Similar content being viewed by others
Main
In the absence of cell movement, development of multicellular plant organs strictly relies on the coordination between cell division orientation and cell fate specification. Failures in either of them results in aberrant structures with impaired growth1,2,3,4,5,6. Arabidopsis root structure is highly regular in that distinct cell types and tissue layers are specified in order7,8,9,10,11,12. Root formation initiates from the globular stage when the hypophysis divides transversely to generate two meristematic cell layers, the upper quiescent centre (QC) and the basal columella (COL)-initial-precursor layer7. In the heart stage, a pro-root meristem emerges such that COL initial and COL layers are formed and the initials of the epidermis (Epi)/lateral root cap (LRC), ground tissue and vasculature become morphologically distinguishable7. Thereafter, COL initials produce two more COL layers, leading to a distal root that includes one QC, one COL initial and three COL layers in the mature embryo (Fig. 1a–c). However, genetic evidence on how the embryonic root attains this precise patterning is limited.
a–c,e–g, Images of wild-type and wip123456 embryonic roots at indicated stages. Cyan, hypophysis/QC lineage; yellow, COL initial lineage; orange, COL layers; grey, delayed/failed layer formation; light purple, ground tissue initials; olive green, Epi/LRC initials; pink, vascular initials. Scale bars, 50 μm. d, Wild-type, wip123456 and wip245 seedlings at 3 d.p.g. Scale bar, 1 mm. h, Frequency of wild-type and wip123456 embryonic roots with normal or delayed/failed layer formation at indicated stages. Data represents mean ± s.d.; biological replicates (N) and sample size per replicate (n) are listed in Supplementary Table 5. P values were calculated with two-tailed unpaired Student’s t-test; wip123456 versus wild type: **P < 0.01, ***P < 0.005, P1 = 0.0021, P2 = 0.0018, P3 = 7.4 × 10−5, P4 = 0.0094. The number presented at the bottom of each image in a–g represents the counts of indicated phenotype (left) versus the total counts (right). G1, early-globular stage; G2, late-globular stage; H1, early-heart stage; H2, late-heart stage; ME, mature embryo. See also Extended Data Fig. 1.
A subgroup of A1d C2H2 zinc finger WIP transcription factors, conserved in land plants, are required for the development of various multicellular structures13,14,15,16,17,18,19,20,21,22,23,24. In Marchantia polymorpha, MpWIP promotes the morphogenesis of the air pore complex in the dorsal epidermis14. The periclinal cell divisions that form the tiers of the air pore complex mostly fail to occur in mutants with reduced MpWIP protein activities14. We have previously shown that in the sex determination of Cucumis melo, CmWIP1 inhibits carpel development, causing the formation of unisexual male flowers19. This growth inhibitory function of CmWIP1 is shared or partially shared by the Arabidopsis WIP homologues23. The transgenic plants overexpressing CmWIP1 or AtWIP genes display similar growth defects, including smaller rosettes, reduced statures and fertilities23. There are six AtWIP members, WIP1/TT1 (TRANSPARENT TESTA 1) is phylogenetically closer to WIP3 and WIP6, WIP2/NTT (NO TRANSMITTING TRACT) is clustered with WIP4 and WIP513,23. WIP2, WIP4 and WIP5 expression in the embryonic root is necessary for the distal stem cell fate within the root meristem13.
Here we uncover an embryo-maternal communication in Arabidopsis, which is mediated by the WIP gene family members. Embryo-maternal communication in animals is considered important in early zygote/embryo development and implantation25,26,27,28,29. In Arabidopsis, only a few genetic mutations have been identified to show maternal control of embryo development and these mutations affect the induction of bilateral symmetry or the patterning of the suspensor30,31,32. Our results reveal that WIP1, WIP3 and WIP6 are expressed in the maternal tissues surrounding the embryo and suspensor, where they act non-cell autonomously to repress root cell fate specification through SIMILAR TO RADICAL-INDUCED CELL DEATH ONE (SRO) gene family members. The embryonic WIPs functionally oppose those maternal WIPs to orchestrate cell division orientation and cell fate specification in the embryonic root, thereby promoting regular root formation.
WIP genes regulate root cell division orientation
To study the AtWIP gene family, we combined the mutations in all the family members to generate sextuple wip123456 mutants (Supplementary Fig. 1). In stark contrast to the rootless triple wip245 mutants, post-embryonic root growth of wip123456 mutants was comparable to that of the wild type, indicating that WIP1, WIP3 and/or WIP6 inhibit embryonic root formation in wip245 mutants (Fig. 1d and Extended Data Fig. 1a). Cell divisions in wip245 embryonic roots were abnormal from the globular stage and became progressively more severe such that no tissue layers can be distinguished (Fig. 1h and Extended Data Fig. 1b–d)13. Eventually, a malformed ‘peg-like’ structure lacking a functional meristem was formed (Extended Data Fig. 1d–f)13. The rescued development of wip123456 roots prompted us to trace their formation in the early embryo.
Oriented cell divisions were perturbed in wip123456 embryonic roots at early stages (Fig. 1e,f,h and Extended Data Fig. 1g–i). From the globular to the heart stage, ~50% of the hypophysis divided longitudinally or obliquely and ~83% of the pro-distal root meristems were disorganized with unrecognizable tissue layers (Fig. 1e,f,h, Extended Data Fig. 1g–i and Supplementary Fig. 2). However, the majority of the roots in mature embryos (~72%) showed a well-organized morphology, suggesting that oriented cell divisions in the early embryonic root are not instrumental in bringing about a competent root morphogenesis (Fig. 1g,h and Supplementary Fig. 2). In the other ~28%, one COL layer is missing or incomplete (Extended Data Fig. 1j). Together, WIP genes regulate cell division orientation during embryonic root development. Although it is known that WIP2, WIP4 and WIP5 are required for oriented divisions in root cells13, it is not clear whether the rootless phenotype was caused by these defective divisions. The embryonic root phenotype in wip123456 mutants suggests that root formation relies on the WIP-mediated regulatory mechanisms besides those involved in cell division orientation.
WIP1, WIP3 and/or WIP6 repress root cell fate specification
Because cell divisions remained abnormal in wip123456 embryonic roots, these divisions appear not to be critical for the WIP1-, WIP3- and/or WIP6-induced inhibition of wip245 root formation. We therefore examined another process pivotal to root development—cell fate specification. We followed the expression of relevant markers to indicate root cell fates during embryonic stages: DR5 to monitor cellular auxin response33,34, WUSCHEL-RELATED HOMEOBOX5 (WOX5) to mark the hypophysis and QC cells35, and SOMBRERO (SMB) to indicate the differentiation of distal cell types in the root36.
We first investigated their expression in wild-type and in wip245 embryonic roots. In wild type, DR5::GFP and proWOX5::erCFP expression accumulated in the hypophysis and adjacent suspensor cells in the globular stage, then the DR5 maximum shifted basally and the WOX5 maximum converged to the QC cells (Fig. 2a,c and Extended Data Fig. 2a,b). The initial proSMB::erCFP expression was detected in the uppermost suspensor cell at the transition or the early-heart stage, then its expression expanded to the first COL layer (Fig. 2e and Extended Data Fig. 2c). In wip245 mutants, high DR5 and WOX5 expression was retained in the hypophysis and its derivatives until the early-heart stage, then they were depleted from the root cells (Extended Data Fig. 2d,e)13. The DR5 and WOX5 maxima were detected in the cells apically adjacent to the root cells (Extended Data Fig. 2d,e)13. No SMB expression was detected at any stages. These data indicate that WIP2, WIP4 and WIP5 are required to sustain root cell specification from the heart stage onwards.
a–f, Images of wild-type and wip123456 embryos and suspensors expressing indicated markers. Scale bars, 50 μm. g–j, Images of wild-type siliques and developing seeds expressing indicated WIP reporters. Scale bars for g,i, 1 mm; for h,j, 50 μm. k, Images of targeted proWIP4::cWIP1:VENUS expression in wip245 embryos, suspensors and primary roots. Scale bars, 50 μm. White frames highlight cell outlines of the hypophyseal derivatives. Cyan dots, cells in hypophysis/QC lineage; yellow dots, cells in COL initial lineage; orange dots, cells in COL layers; grey dots, cells in delayed/failed layer formation. White arrow in b, the DR5::GFP expression in basal cells of the suspensor; red frames in g and i, the zoom-in areas; red arrow in i, the proWIP3::GUS expression in the funiculus. PR, primary root at 3 d.p.g.; M, micropylar end; C, chalazal end; PC, placentochalaza; oi1, outer integument 1; oi2, outer integument 2; ii1, inner integument 1 (endothelium). The experiments in a–d and e–k were repeated four and three times, respectively, with similar results. See also Extended Data Figs. 2–4.
Next, we analysed the marker expression in wip123456 embryos. In the globular stage, DR5 and WOX5 were highly expressed in the hypophysis and its derivatives (Fig. 2b,d and Extended Data Fig. 2f). Unlike the depleted or inactivated expression of these markers in wip245 roots, their expression in wip123456 roots from the heart stage appeared as in the wild type. The DR5 maximum shifted basally, the WOX5 maximum resided in the cells located at the centre of the root meristem and the initial SMB expression was detected in the uppermost suspensor cell (Fig. 2b,d,f and Extended Data Fig. 2f,g). These results show that the root cells are properly specified in wip123456 embryos. This was further strengthened by the comparison of transcriptional profiles in the young primary root meristems between wild type and wip123456 mutants. Only 68 differentially expressed genes were identified, and among them, no major root patterning genes were present, such as PLETHORA, SHORTROOT and SCARECROW (Supplementary Table 4)37,38.
Additionally, in wip245 and wip123456 mutants, we observed frequent mis-expression of DR5 in basal cells of the suspensor from early developmental stages, indicating that WIP genes are required to repress auxin response in the suspensor (Fig. 2b and Extended Data Fig. 2d,f,h).
WIP1, WIP3 and WIP6 act maternally
To unravel how WIP1, WIP3 and/or WIP6 fulfill the repression of root cell fate specification, we determined their spatiotemporal expression pattern. We generated a set of WIP reporters and first examined their expression patterns in wild-type plants. Expression of a VENUS-tagged WIP1 protein fusion in wip1 mutants complemented the yellow-seed defect (Extended Data Fig. 3a–d)20. WIP1 promoter and WIP1 protein activities were detected in the integument, with the highest level in the endothelium and weaker levels in the outer integument layers (Fig. 2g,h and Extended Data Fig. 3e). WIP3 promoter and WIP3 protein activities were observed in the placentochalaza, funiculus and all silique tissues (Fig. 2i,j and Extended Data Fig. 3g). WIP6 promoter activities resided in the floral organ abscission zone (Extended Data Fig. 3i). In the embryo and suspensor at and before the late-heart stage, we did not detect WIP1, WIP3 and WIP6 protein activities (Extended Data Fig. 3f,h,j). To summarize, WIP1, WIP3 and WIP6 are expressed in the maternal tissues surrounding the embryo and suspensor.
We then assessed whether the WIP1-, WIP3- and/or WIP6-mediated repression of root cell fate specification in wip245 mutants could be due to their altered expression. Because we maintained wip245 homozygous embryos in a wip2+/−45 line with a segregating wip2 allele, we observed WIP1, WIP3 and WIP6 expression in the surrounding maternal tissues of this line. These genes were expressed as in the wild type, with a slight downregulation of WIP1 and upregulation of WIP3 (Extended Data Fig. 3k–n). In wip245 embryos and suspensors, no WIP1, WIP3 and WIP6 expression was detected, suggesting that the repression is not caused by the embryonic activation of these genes (Extended Data Fig. 3o–q).
To further validate this, we forced the accumulation of WIP1 proteins in wip245 embryos and suspensors by using a 4.1 kb WIP4 promoter, whose expression was excluded from the maternal tissues (Extended Data Fig. 4a,b). Targeted WIP1 expression in wip245 mutants rescued their embryonic root phenotypes (including oriented cell divisions) and post-embryonic root growth to a level similar to those of WIP4- or WIP5-complemented wip245 plants (Fig. 2k, Extended Data Fig. 4c–e and Supplementary Fig. 3a–d), indicating that WIP proteins share common functions in the embryo and suspensor.
Taken together, these data suggest that WIP1, WIP3 and WIP6 act non-cell autonomously from the maternal tissues to repress root cell fate specification in the embryonic root. Hence, the regulatory roles of WIP genes during embryonic root development are spatially distributed in the embryonic and surrounding maternal tissues. The permissive role of the embryonic WIPs dominantly oppose the repressive role of the maternal WIP1, WIP3 and/or WIP6 to promote regular root formation.
SRO members are required for WIP1-induced growth arrests
We next explored the molecular mechanisms underlying the maternal WIP-mediated repression of root formation. We have shown previously that overexpression of WIP1 (one of the maternal WIPs) by using the LhGR/pOp6 dexamethasone (DEX)-inducible system strongly inhibits plant growth (DEX:WIP1)23,39; we therefore subjected this line to ethyl methanesulfonate (EMS) mutagenesis.
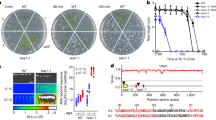
We isolated a suppressor line, q195, that rescued plant/root growth upon DEX induction and identified a non-synonymous cytosine-to-thymine mutation in the C-terminal RST (RCD1-SRO-TAF4)-domain of RADICAL-INDUCED CELL DEATH1 (RCD1) gene (Fig. 3a–c, Extended Data Fig. 5a,b and Supplementary Fig. 4)40,41. This point mutation caused an amino acid change from proline to leucine at position 511 (Pro511Leu) and did not interfere with the transcription of the mutated RCD1 (Fig. 3b, and Supplementary Figs. 4 and 5b). Unlike null rcd1-4 mutants that showed pleiotropic developmental defects, including compact rosettes, malformed leaves and reduced statures41,42, q195 mutants resembled the wild type, indicating that the Pro511Leu is a hypomorphic mutation (Supplementary Fig. 5).
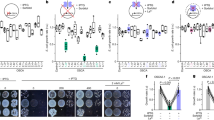
a, RCD1 gene sequence. Positions of the T-DNA insertion (black triangle) and the causal mutation of q195 (blue asterisk) are shown. Coloured boxes indicate the coding sequences: green, WWE domain; pink, PARP domain; blue, RST domain. Blue asterisk: the Pro511Leu mutation (amino acid); C>T: the cytosine-to-thymine mutation (nucleotide). b, Alignment of protein sequences in the RST domain among q195 and SRO family members. Black areas indicate conserved amino acids, cyan area highlights the Pro511Leu mutation. c, Overview of DEX:WIP1 and q195 seedlings germinated on 1/2 MS medium supplemented with 30 nM dexamethasone at 7 d.p.g. Two biological replicates were performed. Scale bars, 1 cm. d, Images of wild-type embryos/suspensors (right panel) and their surrounding maternal tissues (left panel) expressing proRCD1::erCFP. Scale bar, 50 μm. e, Image of proRCD1::gRCD1:VENUS-complemented rcd1-4 embryos and suspensors. Scale bar, 50 μm. f–h, Images of rcd1-4sro1 embryonic roots at indicated stages. The number presented at the bottom of each image represents the counts of indicated phenotype (left) versus the total counts (right). Coloured parts are as described in Fig. 1. Scale bars, 50 μm. i, Frequency of rcd1-4sro1 embryonic roots with normal or delayed/failed layer formation at indicated stages. Data represents mean ± s.d.; biological replicates (N) and sample size per replicate (n) are listed in Supplementary Table 5. P values were calculated using two-tailed unpaired Student’s t-test; rcd1-4sro1 versus wild type: *P < 0.05, ***P < 0.005, P1 = 0.0021, P2 = 0.029, P3 = 0.0045, P4 = 0.0007. The experiments in d and e were repeated three times, with similar results. See also Extended Data Figs. 5 and 6.
Because the Pro511 amino acid in RCD1 is conserved in its closest homologue (Fig. 3b), SIMILAR TO RCD ONE1 (SRO1), to further validate that genes in the SRO family are responsible for the inhibition, we crossed loss-of-function rcd1-4 and sro1 mutants to DEX:WIP1 plants (Supplementary Fig. 5a,b). Similar to q195 mutants, plant/root growth was recovered in both rcd1-4 DEX:WIP1 and sro1 DEX:WIP1 seedlings upon DEX induction (Extended Data Fig. 5b,c), indicating that overexpression of WIP1 acts through RCD1 and SRO1 to induce growth arrests.
SRO members are required for embryonic root development
Since RCD1 and SRO1 were identified from an overexpression system, we assessed whether they were biologically relevant to embryonic root development by investigating their expression and function. RCD1 and SRO1 were expressed in all cells of embryos, suspensors, maternal tissues and primary roots, with a higher transcription level of RCD1 than SRO1 (Fig. 3d and Extended Data Fig. 6a–e)42. In these cells, the C-terminal tagged VENUS protein fusions of RCD1 and SRO1 were mainly nuclear localized (Fig. 3e and Extended Data Fig. 6f,g). Expression of proRCD1-driven genomic RCD1 and proSRO1-driven genomic SRO1 constructs complemented rcd1-4 and rcd1-4sro1 mutants, respectively (Supplementary Fig. 6a–c)41,43.
Post-embryonic root growth of rcd1-4 and sro1 single mutants was similar to that of wild type, but was severely retarded in rcd1-4sro1 double mutants (Supplementary Fig. 6d), suggesting that RCD1 and SRO1 are functionally redundant41,43. We therefore investigated embryonic root development in rcd1-4sro1 mutants. Low frequencies of mis-oriented cell divisions occurred in the hypophysis and COL initial precursors, resulting in ~24% and ~27% of the root meristems being disorganized at the late-globular and late-heart stage respectively (Fig. 3f,g,i and Supplementary Fig. 2). Around 84% of the roots lacked one COL layer in mature embryos (Fig. 3h,i and Supplementary Fig. 2). In addition, QC cells divided periclinally, with the frequency increasing progressively from the transition/early-heart stage onwards (Extended Data Fig. 6h). These phenotypes imply that RCD1 and SRO1 are required for embryonic root development.
Maternal WIPs act through SRO members
Given that RCD1 and SRO1 are required for the WIP1-induced root growth arrest and are biologically relevant to embryonic root development, we asked whether they enabled WIP1, WIP3 and/or WIP6 to inhibit wip245 root formation. To assess this, we generated quadruple rcd1-4wip245 and sro1wip245 mutants. The primary root growth of the mutants was recovered partially or totally, indicating that RCD1 and SRO1 are required for root growth inhibition (Fig. 4a).
a, Overview of rcd1-4wip245 and sro1wip245 seedlings at 7 d.p.g. Scale bar, 1 cm. b–d, Images of rcd1-4wip245 embryonic roots at indicated stages. The number presented at the bottom of each image represents the counts of indicated phenotype (left) versus the total counts (right). Coloured parts are as described in Fig. 1. Scale bars, 50 μm. e, Frequency of rcd1-4wip245 and sro1wip245 embryonic roots with normal or delayed/failed layer formation at indicated stages. Data represents mean ± s.d.; biological replicates (N) and sample size per replicate (n) are listed in Supplementary Table 5. P values were calculated using two-tailed unpaired Student’s t-test; mutant versus wild type: *P < 0.05, **P < 0.01, ***P < 0.005, P1 = 0.0099, P2 = 0.028, P3 = 3.4×10−5, P4 = 2.1 × 10−5, P5 = 0.0002, P6 = 0.0011, P7 = 0.0002, P8 = 0.0043. f–j, Images of rcd1-4wip245 and sro1wip245 embryos, suspensors and primary roots at indicated stages expressing indicated markers. Scale bars, 50 μm. White frames highlight cell outlines of hypophyseal derivatives. Cyan dots, cells in hypophysis/QC lineage; yellow dots, cells in COL initial lineage; grey dots, cells in delayed/failed layer formation. The experiments in f–j were repeated three times, with similar results. See also Extended Data Fig. 7.
We next assessed whether these inhibitory functions of RCD1 and SRO1 could be attributed to their regulation of WIP1, WIP3 and WIP6 expression in rcd1-4wip245 and sro1wip245 embryonic and/or surrounding maternal tissues. In these mutants, we found no change in WIP expression patterns compared with the wild type (Supplementary Fig. 7), suggesting that neither individual RCD1 nor SRO1 regulates the transcription of WIP1, WIP3 and WIP6. Vice versa, RCD1 and SRO1 expression in wip2+/−45, wip245 and wip123456 embryonic and surrounding maternal tissues were comparable to that of the wild type, with a slight upregulation of RCD1 in wip2+/−45 siliques (Supplementary Fig. 8a–m). This suggests that WIPs do not regulate the transcription of RCD1 and SRO1. We then assessed whether WIP1 could physically bind to RCD1 by using the yeast-two-hybrid assay, and detected no interaction (Supplementary Fig. 8n).
To further dissect the cause of the rescue, we traced rcd1-4wip245 and sro1wip245 embryonic root morphologies. In rcd1-4wip245 mutants, although ~90% of the hypophysis underwent transverse cell division as in the wild type, mis-oriented cell divisions still frequently occurred in COL initial precursors, leading to ~78% of the roots lacking separated COL initial and COL tissue layers at the late-heart stage (Fig. 4b,c,e and Supplementary Fig. 2). Mature rcd1-4wip245 roots possessed less COL layers than the wild type, resulting in reduced or loss of amyloplast-containing cells that caused the agravitropism (Fig. 4d,e, Extended Data Fig. 7a,b and Supplementary Fig. 2). The morphology of sro1wip245 embryonic roots phenocopied the wip sextuple mutants, in which the root meristematic cells often divided irregularly at the early stages (Fig. 4e, Extended Data Fig. 7c–e and Supplementary Fig. 2). In summary, oriented cell divisions are not fully rescued during early rcd1-4wip245 and sro1wip245 embryonic root development.
Finally, we followed the expression of the DR5, WOX5 and SMB markers during rcd1-4wip245 and sro1wip245 embryonic root development. Their expression was not affected in general (Fig. 4f–j and Extended Data Fig. 7f–h), indicating that RCD1 and SRO1 are required for the maternal WIP-induced repression of root cell fate specification.
Discussion
In this study, we reveal that WIP genes regulate Arabidopsis embryonic root development by orchestrating cell division orientation and cell fate specification (Fig. 5). WIP genes are spatially expressed in the embryo/suspensor (WIP2, WIP4 and WIP5) and their surrounding maternal tissues (WIP1, WIP2, WIP3 and WIP6) (Fig. 5). WIP functions in the embryo and suspensor are common and dominant, promoting regular root formation. Impairing them causes aborted roots with mis-oriented cell divisions and mis-specified cell types (Fig. 5). The maternal WIPs act non-cell autonomously to disable root cell fate specification in the absence of the embryonic WIPs. When losing them in wip245 mutants, root formation is rescued with properly specified cells, but root cell divisions remain disordered in the early embryo (Fig. 5). WIP2 is expressed both embryonically and maternally (Supplementary Fig. 3e)17, suggesting that it may have dual inputs to embryonic root development. Moreover, we find that RCD1 and SRO1 are responsible for the maternal WIP-induced repression of root formation (Fig. 5). Removal of either of them from wip245 embryonic roots rescues cell fate specification.
WIP genes expressed in the embryo and suspensor functionally oppose those expressed in the surrounding maternal tissues to orchestrate cell division orientation and cell fate specification in the embryonic root, thereby promoting regular root formation. Impairing the embryonic WIPs results in aborted roots with mis-oriented cell divisions and mis-specified cell types. The maternal WIPs act non-cell autonomously to repress root cell fate specification through RCD1 and SRO1. When losing all WIPs, root cells are properly specified, but they divide irregularly in the early embryo. However, these defective divisions barely alter the root morphology in mature embryos.
Spatial expression of WIP genes determines their roles
In Arabidopsis, WIP1 regulates the accumulation of pro-anthocyanins in the integument20. The expression of CmWIP1 and other AtWIP genes in the integument of wip1 mutants is able to complement or partially complement the yellow-seed defect22,23. The ectopic expression of WIP1 in the carpel primordium and pedicel represses carpel development, leading to the transition from hermaphrodite flower to male flower in Arabidopsis23. This echoes the CmWIP1 function in the melon carpel primordium19. Here we show that the ectopic expression of WIP1 in the embryo and suspensor fulfills the permissive role of WIP2, WIP4 and WIP5 (the embryonic WIPs) in root formation. Thus, WIP proteins share or partially share common functions. Together, these data suggest that the spatial expression pattern of WIP genes probably determines their roles.
SRO family members are hub proteins
SRO family members are land-plant specific, containing a poly(ADP-ribose)polymerase (PARP) domain, a C-terminal RST domain and some have an N-terminal WWE domain, a conserved domain found in several PARP, deltex and ubiquitination-related proteins (Fig. 3a)44,45,46. Bona fide PARP proteins catalyse poly(ADP-ribosyl)ation—the synthesis of poly(ADP-ribose) chains by post-translationally transferring ADP-ribose molecules onto targeted proteins. These chains constitute an interaction platform for recruiting binding partners47. RCD1 and SRO1, however, harbour a non-canonical catalytic motif in their PARP domain and show no enzymatic activity45,48. The helical RST domain, a member of αα-hub family, is specialized in interacting with transcription factors41,44,45,49. The WWE domain is predicted to mediate specific protein interactions in ubiquitination and ADP-ribose conjugation46. With these structural signatures, it is conceivable that RCD1 and SRO1 act as non-enzymatic scaffolding proteins to modulate cellular responses, such as DNA repair, protein degradation and cell death. The Pro511 amino acid in RCD1, which is mutated in q195 mutants, resides in the RST domain but not in the α-helix positions—the core components to form an αα-hairpin super-secondary structure required for protein–protein interactions44,49. Thus, the Pro511Leu mutation is unlikely to disrupt the folding of the RST domain. We therefore expect a considerable overlap between the interactome of RCD1 and the mutated RCD1, which may explain the wild-type-like phenotype of q195 mutants. Despite being a hub protein, RCD1 does not bind to WIP1.
Maternal control of embryonic root development
In Drosophila, positional cues provided maternally to eggs are intensively studied. They have been shown to be critical in determining the growth axis and cell fate in early embryogenesis25,26. In plants, it has been shown that nutrients and signalling molecules, including sucrose, polyamine and auxin, can move from the silique (pod) and integument to the developing embryo via the suspensor50,51,52,53,54,55. These lead to the hypothesis that there might be certain mobile molecules travelling between the maternal tissues and the embryo that function as positional cues. The embryonic and the maternal WIPs may modulate the cellular concentration of these molecules to promote or repress root cell fate specification.
Genome-wide analysis in Arabidopsis reveals that the growth inhibitory role of WIPs is executed by upregulating stress-inducible genes and downregulating development-promoting genes23. RCD1 and SRO1 have been shown to be involved in oxidative stress, pathogen defence, hormone signalling and plant development. The phenotypic defects displayed in rcd1 and rcd1sro1 mutants are similar to stress-induced morphogenic response, known to be associated with change in reactive oxygen species status41,42,43,48,56,57,58,59. Thus, WIP and SRO family members modulate growth–defence trade-offs, which are vital for plant survival and reproduction. Their integration into embryonic root development might allow embryos to sense and/or respond to the growth condition of mother plants, potentially leading to competition between the embryonic and maternal WIP-mediated inputs to determine root formation, thereby affecting seed viability. Since three embryonic WIPs act redundantly and dominantly, Arabidopsis has evolved a robust rooting system. It is unlikely that the maternal WIPs can outcompete the embryonic WIPs to induce a rootless phenotype in wild plants challenged by stresses. Instead, post-embryonic root viability and/or root system architecture could be ‘primed’ at the embryonic stage by the cross-communication between these two WIP groups.
Methods
Plant materials and growth conditions
Arabidopsis thaliana plants, Columbia ecotype Col-0, were used for all experiments and transgenic lines in this study. Transferred DNA (T-DNA) insertion mutants of wip1 (tt1-3; SALK_026171), wip2 (ntt-2; SALK_007406), wip2-3 (SM_3_16705), wip2-4 (SM_3_23211), wip3 (SALK_072471), wip4 (SALK_014672), wip5 (SALK_114838), wip6 (SALK_148869), rcd1-4 (GABI_229D11) and sro1 (sro1-1; SALK_074525) were obtained from the Eurasian Arabidopsis Stock Centre (uNASC). The T-DNA insertion was genotyped by PCR-based genotyping (M0267S, Taq polymerase, NEB or 9PIM300, GoTaq DNA polymerase, Promega). The transcription of the inserted gene was quantified by reverse transcription quantitative PCR (RT–qPCR). Primers used in genotyping and RT–qPCR are listed in Supplementary Table 1. Triple wip245 mutant is also termed nww13. The two-component DEX:WIP1 (35::LhGR/pOp6::cWIP1) overexpression system was generated as previously described23.
Seeds were fume sterilized in a sealed container with 30 ml bleach (9.6% sodium hypochlorite) supplemented with 1.2 ml 37% hydrochloric acid for 4–6 h, then suspended in 0.1% agarose and soaked at 4 °C in the dark for 2 d. Seeds were plated on 1/2x MS growth medium consisting of half-strength Murashige Skoog salts (including vitamins), 0.8% plant agar and MES buffer (pH 5.8). Seedlings and plants were grown at 22 °C in a 16 h light/8 h dark cycle.
Cloning, construction and transgenic plants
All primers used for cloning are listed in Supplementary Table 1. Promoter, coding sequences (CDs) and genomic fragments of proSMB4.1kb, proWIP14.1kb, proWIP24.6kb, proWIP34.5kb, proWIP44.1kb, proWIP54.9kb, proWIP65.3kb, proRCD12.7kb, proSRO12.3kb, cWIP1, cWIP6, gWIP3, gWIP4, gWIP5, gRCD1, cRCD1, gSRO1, RCD16.7kb (promoter to 3’UTR), SRO14.8kb (promoter to 3’UTR) and cANAC013 were cloned from Col-0 genomic or Col-0 derived complementary DNA templates using Thermo Phusion high-fidelity DNA polymerase (F530S, NEB). Promoter fragments of proSMB4.1kb, proWIP14.1kb, proWIP24.6kb, proWIP34.5kb, proWIP44.1kb, proWIP54.9kb, proWIP65.3kb and proSRO12.3kb were inserted into a synthetic MultiSite Gateway-compatible entry vector pENTRY57_L4R1 (synthesized by ProteoGenix) containing a pair of BsaI cutting site directly flanked by the attL4 and attR1 sites. In brief, pENTRY57_L4R1 was cut and linearized by BsaI (R0535, NEB), the attL4/attR1-containing part was separated by agarose gel electrophoresis and purified by NucleoSpin gel and PCR clean-up kit (740609, Macherey-Nagel). proSMB4.1kb, proWIP14.1kb, proWIP24.6kb, proWIP34.5kb, proWIP44.1kb, proWIP54.9kb, proWIP65.3kb and proSRO12.3kb were inserted into the purified/linearized pENTR57_L4R1, using ClonExpress II one-step cloning kit (C112-02, Vazyme). proRCD12.7kb was inserted into a MultiSite Gateway entry vector pDONR P4_P1r using Gateway BP Clonase II enzyme (11789020, Invitrogen). cWIP1, cWIP6, gWIP3, gWIP4, gWIP5, gRCD1, cRCD1, gSRO1, RCD16.7kb, SRO14.8kb and cANAC013 were inserted into a MultiSite Gateway entry vector pDONR 221, using Gateway BP Clonase II enzyme (11789020, Invitrogen).
All constructs used in this study are listed in Supplementary Table 2. Destination vector of pH7m34GW60, pDEST22 (Invitrogen) and pDEST32 (Invitrogen) were used to generate final constructs using Gateway LR Clonase II enzyme (11791020, Invitrogen). proWOX5::erCFP, proSMB::erCFP, proWIP1::erCFP, proWIP3::erCFP, proWIP4::erCFP, proWIP5::erCFP, proRCD1::erCFP and proSRO1::erCFP were generated by fusing the proWOX54.5kb61, proSMB4.1kb, proWIP14.1kb, proWIP34.5kb, proWIP44.1kb, proWIP54.9kb, proRCD12.7kb and proSRO12.3kb promoter in front of an endoplasmic reticulum-tagged cyan fluorescent protein coding region (erCFP), respectively. proWIP1::GUS, proWIP2::GUS, proWIP3::GUS, proWIP4::GUS, proWIP5::GUS, proWIP6::GUS, proRCD1::GUS and proSRO1::GUS were generated by fusing the proWIP14.1kb, proWIP24.6kb, proWIP34.5kb, proWIP44.1kb, proWIP54.9kb, proWIP65.3kb, proRCD12.7kb and proSRO12.3kb promoter in front of the β-glucuronidase (GUS) coding sequence, respectively. proWIP1::cWIP1:VENUS, proWIP3::gWIP3:VENUS, proWIP4::gWIP4:VENUS, proWIP5::gWIP5:VENUS, proWIP6::cWIP6:VENUS, proRCD1::gRCD1:VENUS and proSRO1::gSRO1:VENUS were generated by fusing the 3’ end of cWIP1, gWIP3, gWIP4, gWIP5, cWIP6, gRCD1 and gSRO1 CDs or genomic sequence to a C-terminal coding region of a VENUS yellow fluorescent protein62, respectively, and placing them under the proWIP14.1kb, proWIP34.5kb, proWIP44.1kb, proWIP54.9kb, proWIP65.3kb, proRCD12.7kb and proSRO12.3kb promoter, respectively. proWIP4::cWIP1:VENUS was constructed by fusing the 3’ end of cWIP1 CDs sequence to the C-terminal coding region of the VENUS and placing them under the proWIP44.1kb promoter. RCD1:NosT and SRO1:NosT were generated by putting RCD16.7kb and SRO14.8kb genomic sequences (from promoter to 3’UTR) in front of a nopaline synthase terminator (NosT), respectively.
Transformation was performed according to the floral dip method63. Transformants were selected on the basis of their resistance. Transformants in wip2+/−45, rcd1-4+/−sro1 or rcd1-4sro1+/− segregating populations were additionally selected (if applicable) for the homozygous wip245 and rcd1-4sro1 lines by genotyping and the no-transmitting-tract phenotype of wip217.
Microscopy and histology
To visualize and quantify morphologies of embryos and suspensors at and before the heart stage, young siliques were first collected for tissue clearance, followed by SCRI Renaissance 2200 (SR2200) staining64. To clear the tissue, young siliques were immersed in an NaOH-SDS solution (200 mM NaOH + 1.0% SDS) for 2 h at 60 °C, rinsed two times with water, treated with 5% sodium hypochlorite solution (bleach), vacuum infiltrated for 2 h and left at 4 °C overnight. To stain the tissue, cleared siliques were rinsed two times with water and once with PBS buffer (pH 8.0; 6603369, Beckman Coulter), stained with SR2200 solution (0.1% v/v SR2200, 1.0% v/v dimethyl sulfoxide (DMSO), 0.05% w/v Triton X-100 and 5.0% glycerol in PBS buffer pH 8.0), vacuum infiltrated for 30 min, left at 4 °C overnight and up to several weeks. Before imaging embryos and suspensors, silique valves were peeled off manually.
To visualize and quantify root morphologies in mature embryos, aniline blue staining was performed65. Seeds were soaked overnight in water at 4 °C and the seed coats were removed. The embryos were dehydrated through 15, 50, 70 and 96% (v/v) ethanol, two changes of 100% (v/v) ethanol at 15 min each, and left in 100% (v/v) ethanol at 4 °C overnight. Then, the embryos were rehydrated through 96, 70, 50 and 15% (v/v) ethanol and two changes of water at 15 min in each. The embryos were stained in a 1:20 dilution of an aniline blue stock solution (0.5% w/v aniline blue (28631-66-5, Acros Organics), 0.2 M phosphate buffer, pH 6.5) for 30 min and rinsed with water for 15 min. The embryos were dehydrated and then rehydrated again through the ethanol series as described above. Stained embryos were transferred to microscope slides, mounted in Hoyer’s solution (30 g gum arabic (G9752, Sigma-Aldrich), 200 g chloral hydrate (23100, Sigma-Aldrich), 20 g glycerol and 50 ml Milli Q water). Samples were covered with cover slides and left undisturbed for 3 d. Images were recorded using a Zeiss LSM 880 laser scanning confocal microscope. SR2200 and aniline blue were excited with a diode 405 nm laser line, and the emission was measured at 425–570 nm.
To visualize fluorescent markers in living embryos and suspensors, developing seeds were dissected into several droplets of a digestive enzyme solution (1 mg ml−1 cellulase onozuka R-10, 0.8 mg ml−1 macerozyme R-10, 80 mM d-sorbitol, 10% glycerol and 0.058% MES pH 5.8)66 on a microscope slide for ~1.5 h in a sealed container, then the digestive enzyme solution was replaced by a staining solution (0.1% v/v SR2200 and 20% glycerol in PBS buffer, pH 8.0). Samples were covered with cover slides, and embryos and suspensors were squeezed out by gently pressing the cover slide. Images were recorded using the Zeiss LSM 880. SR2200 was excited with the 405 nm line, and the emission was measured at 410–530 nm when combined with green fluorescent protein (GFP), 410–470 nm when combined with cyan fluorescent protein (CFP) and VENUS; GFP was excited with a 488 nm argon laser and the emission was measured at 490–595 nm; CFP was excited with a 458 nm argon laser and the emission was measured at 465–580 nm; VENUS was excited with a 514 nm argon laser and the emission was measured at 515–625 nm.
To visualize fluorescent markers in developing seeds, samples were mounted in the 0.1 mg ml−1 propidium iodide (PI; P4170, Sigma-Aldrich) solution supplemented with 7% sucrose67. To visualize fluorescent markers in primary roots, samples were stained with PI. To visualize amyloplasts in primary roots, the modified pseudo-Schiff propidium iodide (mPS-PI) staining method was performed as previously described68. Images were recorded using the Zeiss LSM 880. PI was excited with a 561 nm diode laser, or the 488 nm argon laser (when combined with GFP) or the 514 nm argon laser (when combined with VENUS) and the emission was measured at 585–720 nm. GFP was excited with a 488 nm argon laser and the emission was measured at 490–545 nm; CFP was excited with a 458 nm argon laser and the emission was measured at 455–535 nm; VENUS was excited with a 514 nm argon laser and the emission was measured at 515–590 nm.
Histostaining of promoter-driven GUS activities was visualized by staining siliques and developing seeds for 24 h at 37 °C in a GUS solution: 1 mM X-gluc (R0851, Thermo Fisher) dissolved in dimethylformamide, 10 mM EDTA, 0.1% w/v Triton X-100, 1 mM potassium ferrocyanide K4Fe(CN)6 (P3289, Sigma-Aldrich), 1 mM potassium ferricyanide K3Fe(CN)6 (P8131, Sigma-Aldrich) and 100 mM sodium phosphate buffer, pH 7.0. Before imaging, stained siliques were incubated in 70% ethanol for 3–7 d until chlorophyll was removed. Stained developing seeds were cleared with a chloral-hydrate solution (7 g chloral hydrate, 2 ml Milli Q water and 1 ml glycerol). Images of siliques were recorded using a ZEISS Stemi 305 microscope. Images of developing seeds and primary roots were recorded using an Olympus BX53 (Nomarski/DIC) microscope.
Images were processed using ZENblack_2-3SP1 and Adobe Photoshop CS4. For the confocal images showing the early embryonic morphologies and amyloplasts in primary roots, colours were inverted using Photoshop to have a white background and a black cell outline for better visualization. For the confocal images showing the marker/reporter expression, individual colour channels reflecting the SR2200 or PI staining were sometimes adjusted using ZEN black edition to highlight the cell wall. Images were rotated and cropped using Photoshop.
RT–qPCR analysis
All primer sets used for RT–qPCR are listed in Supplementary Table 1. Young siliques encompassing embryos from approximately the globular to the heart stage were sampled. Total RNAs were extracted by RNeasy plant mini kit (74904, Qiagen), treated with DNase I solution (89836, Thermo Fisher) and subjected to first-strand cDNA synthesis using SuperScript II reverse transcriptase (18064022, Invitrogen). RT products were used as templates for PCR reactions using MESA GREEN qPCR MasterMix Plus for SYBR Assay dTTP 7.5 ml (RT-SY2X-03+WOUN, Eurogentec). All PCR reactions were performed in a CFX96 real-time PCR system (Bio-Rad). Gene expression was calculated relative to ACTIN2 (AT3G18780) using the comparative Ct (2–ΔΔCt) method (ABI PRISM 7700 Sequence Detection System, Applied Biosystems User Bulletin #2, 2001; https://assets.thermofisher.com/TFS-Assets/LSG/manuals/cms_040980.pdf). To detect WIP1 and WIP3 expression levels in wild-type, wip2+/−45, rcd1-4wip245 and sro1wip245 young siliques (Extended Data Fig. 3n and Supplementary Fig. 7i), ‘WIP1_PrimerSet 2’ and ‘WIP3_PrimerSet 1’ were used (Supplementary Fig. 1).
RNA-seq analysis
To determine the genes that are differentially expressed in the primary root meristem of wild-type and wip123456 plants, their seeds were germinated on 1/2x MS growth medium covered with a mesh. Root tips of 5-day-old seedlings were cut with a razor and immediately frozen in liquid nitrogen. Total RNAs were extracted by PicoPure RNA isolation kit (KIT0204, Applied Biosystems) and treated with DNase I (89836, Thermo Fisher). RNA-seq libraries were prepared using NEBNext Poly(A) mRNA Magnetic Isolation Module (E7490, NEB), followed by NEBNext Ultra II RNA library prep kit for Illumina (E7770, NEB) with NEBNext Multiplex Oligos for Illumina (E7500, NEB). RNAs and libraries were quantified using Agilent RNA 6000 Pico kit (41105500, Life Technologies) and Agilent DNA 1000 kit (41106100, Life Technologies), performed on a 2100 Bioanalyzer instrument (Agilent). Two libraries were generated for each genotype. Multiplex-sequencing was conducted on a NextSeq500 platform (Illumina), and between 30 and 35 million read pairs per sample were obtained.
RNA-seq data were analysed using an in-house Snakemake (v5.31.1) pipeline69. Reads were trimmed with Cutadapt (v2.10), and their quality was controlled by FastQC (v0.11.9). Then the reads were aligned against TAIR10 reference genome (https://www.arabidopsis.org/) by STAR (v2.7.5c) software and the alignments were filtered by SAMtools (v1.10). Counts per gene were computed with featureCount (v2.0.1). Differential expression analysis was performed with R script tool DiCoExpress (docker://registry.forgemia.inra.fr/gnet/dicoexpress:latest). The RNA-seq data have been deposited to Sequence Read Archive (PRJNA774717).
The genome sequence can be downloaded (need a subscription to access) at https://www.arabidopsis.org/download_files/Genes/TAIR10_genome_release/TAIR10_chromosome_files/TAIR10_chr_all.fas.
The annotation can be downloaded at https://www.arabidopsis.org/download_files/Genes/Araport11_genome_release/archived/Araport11_GTF_genes_transposons.201606.gtf.gz.
Yeast-two hybrid assay
CDs of RCD1 were cloned into the bait vector pDEST 32 in-frame fused with the Gal4-DNA-binding domain. CDs of ANAC013 (AT1G32870) and WIP1 were cloned into the prey vector pDEST 22. As it has been shown that ANAC013 interacts with RCD141, this pair was used as the positive control. Yeast strain MaV203 was used for the transformation employing the LiAc method70. Briefly, yeast competent cells were prepared and resuspended in TEL solution (10 mM Tris-HCl, pH 7.5, 1 mM ethylenedinitrilotetraacetic acid (EDTA) and 0.1 M LiAc (L6883, Sigma-Aldrich)). For the transformation, 0.1 mg salmon DNA (D9156, Sigma-Aldrich) and 100 ng plasmid DNA (the one to be tested) were added to 50 µl of the competent cells. Then, the cells were gently resuspended in 300 µl plate solution (50% poly(ethylene glycol) (PEG3350; P3640, Sigma-Aldrich) in TEL solution), incubated at 30 °C for 30 min, added 40 µl DMSO, heat shocked at 42 °C for 15 min, placed on ice for 2 min, centrifuged at 14,000 rpm for 10 sec, resuspended in TE solution (10 mM Tris-HCl, pH 7.5 and 1 mM EDTA) and plated on synthetic defined (SD) medium without Trp and/or Leu to select the transformants. The yeast colonies transformed with cRCD1-pDEST32 (cRCD1 in pDEST32) and pDEST22 (an empty vector control) plasmids were selected for the autoactivation test on the SD medium without His, Trp and Leu. A 30 mM 3-amino-1,2,4-triazole (A8056, Sigma-Aldrich) concentration was used to repress the autoactivation of RCD1. Two independent transformations were performed, with three colonies tested per transformation.
EMS suppressor screen
Ethyl methanesulphonate (EMS; M0880, Sigma-Aldrich) suppressor screen was applied to DEX:WIP1 seeds. Around 10,000 seeds were incubated for 17 h at room temperature with gentle agitation in 10 ml 0.3% (v/v) EMS. Then 10 ml Na2S2O3 (1 M) was added to the mix, followed by rotation for 5 min. The mutagenized seeds were washed with water six times (20 min per washing) and sown on soil. A total of 3,500 M1 plants were grown and self-pollinated to produce M2 seeds. To screen the mutant collection, M2 seeds were sown on 1/2x MS medium supplemented with 30 nM dexamethasone (DEX; D4902, Sigma-Aldrich), which was sufficient to inhibit the growth of DEX:WIP1 plants.
Bulk genomic DNA sequencing and analysis
To identify the causal mutation in q195, the M2 q195 revertant was backcrossed to DEX:WIP1 (as a paternal pollen donor) plants, and the F1 plants self-pollinated to produce the F2 segregating population. Genomic DNAs were collected from 20 F2 revertant plants (survivals on 1/2x MS medium supplemented with 30 nM DEX) and 20 randomly selected F2 plants. Next-generation sequencing DNA libraries were constructed following the standard Illumina protocol via TruSeq SBS kit v3-HS (2 × 100 bp; FC-401-3001, Illumina). The libraries were sequenced on Illumina-HiSeq2500. Sequences were trimmed using Trimmomatic (v0.39) and paired-end reads were mapped to the Col-0 reference genome using CLC-Genomics workbench 9.0 software with the following parameters: no_masking; match_score, 1; mismatch_cost, 2; insertion_cost, 3; deletion_cost, 3; length_fraction, 1; similarity_fraction, 0.987.
Mapped single-nucleotide polymorphisms were further filtered according to the character of EMS-induced mutations (mainly G/C to A/T transitions) and the monomorphism in the revertant (expected as a recessive mutation). To pinpoint the causal mutation, a cleaved amplified polymorphic sequences-based mapping was applied to the F2 and F3 segregating populations of more than 1,000 plants, and the mapping delimited the causal mutation to a single gene (AT1G32230, Supplementary Fig. 4). To genotype the causal mutation (q195; the primer set is listed in Supplementary Table 1), the PCR products were digested by DdeI (R0175L, NEB) before running an agarose electrophoresis gel. Wild-type fragments displayed a 220 bp band, while q195 fragments were digested and displayed a 190 bp band.
Protein sequences of the RST domain from the mutated RCD1 (q195 mutants) and Arabidopsis SRO family members were aligned using MEGA-X (v10.0.5; MUSCLE algorithm, default setting). The RST sequences used in the alignment are listed in Supplementary Table 3.
Quantification of embryonic root morphology
To statistically quantify embryonic root morphologies at different developmental stages, young siliques were randomly collected from wild-type (Col-0), wip2+/−45, wip24+/−5, wip245+/−, wip123456, rcd1-4wip245, sro1wip245, rcd1-4+/−sro1 and rcd1-4sro1+/− plants. We maintained wip245 and rcd1-4sro1 embryos in the line with a segregating wip or rcd1-4/sro1 allele, respectively; their morphology at the early-globular stage (G1; before the first cell division of the hypophysis) was indistinguishable. Therefore, the numbers indicated in the bottom of Extended Data Fig. 1b (left panel) and Fig. 3f (left panel) are the counts from all the genetic backgrounds. From the late-globular stage (G2) onwards, wip245 and rcd1-4sro1 embryos were morphologically recognizable.
In mature embryonic roots, the formation of the third columella cell layer (the youngest one) in the wild type or wild-type-like mutants was sometimes incomplete, but always with the central two-cell files accomplished. This was used as a criterion to classify columella layer numbers. If only one of the central columella initials has transversely divided, the new layer was not considered to be formed (Extended Data Fig. 1j).
Root growth measurement
To measure primary root growth, 4- or 5-day-post-germination (d.p.g.) seedlings grown on 1/2x MS medium were transferred to 1/2x MS medium supplemented with either mock or 30 nM DEX for 48 h. Root tip positions were marked by black dots when the seedlings were freshly transferred. The 48 h root growth was measured from the black dot to the root tip by using Fiji-Image J (https://imagej.net/software/fiji/downloads).
Statistics and reproducibility
Biological replicates (N), sample size per biological replicate (n) and P values (two-tailed unpaired Student’s t-test, Microsoft Excel 2016) can be found in the figures, figure legends, Supplementary Tables 5–8 or Source Data. Error bars are s.d. or s.e.m. Bar graphs overlaid with dot plots were generated using Graphpad Prism (v9.3.1) and edited using Adobe Illustrator CS4. For RNA-seq data, differentially expressed genes were defined with the false discovery rate (FDR) < 0.01 and log2 fold change >0.5 and <−0.5.
To determine the DR5 expression pattern in rcd1-4wip245 mutants, DR5::GFP wild-type plants from a transgenic line with a strong expression were crossed with rcd1-4wip245 mutants. To determine the proWIP3::GUS and proWIP6::GUS expression pattern in rcd1-4wip245 mutants, two independent transgenic lines were examined per biological replicate. To determine the expression pattern and complementation of other markers or reporters in each genetic background, three independent transgenic lines were examined per biological replicate. All experiments were biologically repeated at least two times, with similar results.
Because the DR5 expression in examined transgenic lines per genotype showed a similar pattern, to quantify the DR5 expression in basal cells of the suspensor, data from three independent transgenic lines of the wild type, and two independent transgenic lines of wip245 and wip123456 were blindly collected per genotype per biological replicate.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The RNA-seq data of wild-type and wip123456 primary root meristems have been deposited to Sequence Read Archive (PRJNA774717). All data supporting the findings of this study are available in this Article and its Supplementary Information, or from A. Bendahmane upon reasonable request. Source data are provided with this paper.
References
Scheres, B. & Benfey, P. N. Asymmetric cell division in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 505–537 (1999).
Abrash, E. B. & Bergmann, D. C. Asymmetric cell divisions: a view from plant development. Dev. Cell 16, 783–796 (2009).
De Smet, I. & Beeckman, T. Asymmetric cell division in land plants and algae: the driving force for differentiation. Nat. Rev. Mol. Cell Biol. 12, 177–188 (2011).
Petricka, J. J., Van Norman, J. M. & Benfey, P. N. Symmetry breaking in plants: molecular mechanisms regulating asymmetric cell divisions in Arabidopsis. Cold Spring Harb. Perspect. Biol. 1, a000497 (2009).
Pillitteri, L. J., Guo, X. & Dong, J. Asymmetric cell division in plants: mechanisms of symmetry breaking and cell fate determination. Cell. Mol. Life Sci. 73, 4213–4229 (2016).
Heidstra, R. Asymmetric cell division in plant development. Prog. Mol. Subcell. Biol. 45, 1–37 (2007).
Scheres, B. et al. Embryonic origin of the Arabidopsis primary root and root meristem initials. Development 120, 2475–2487 (1994).
Capron, A., Chatfield, S., Provart, N. & Berleth, T. Embryogenesis: pattern formation from a single cell. Arabidopsis Book 7, e0126 (2009).
Jenik, P. D., Gillmor, C. S. & Lukowitz, W. Embryonic patterning in Arabidopsis thaliana. Annu. Rev. Cell Dev. Biol. 23, 207–236 (2007).
Lau, S., Slane, D., Herud, O., Kong, J. & Jurgens, G. Early embryogenesis in flowering plants: setting up the basic body pattern. Annu. Rev. Plant Biol. 63, 483–506 (2012).
ten Hove, C. A., Lu, K. J. & Weijers, D. Building a plant: cell fate specification in the early Arabidopsis embryo. Development 142, 420–430 (2015).
Palovaara, J., de Zeeuw, T. & Weijers, D. Tissue and organ initiation in the plant embryo: a first time for everything. Annu. Rev. Cell Dev. Biol. 32, 47–75 (2016).
Crawford, B. C. W. et al. Genetic control of distal stem cell fate within root and embryonic meristems. Science 347, 655–659 (2015).
Jones, V. A. & Dolan, L. MpWIP regulates air pore complex development in the liverwort Marchantia polymorpha. Development 144, 1472–1476 (2017).
Englbrecht, C. C., Schoof, H. & Bohm, S. Conservation, diversification and expansion of C2H2 zinc finger proteins in the Arabidopsis thaliana genome. BMC Genomics 5, 39 (2004).
Marsch-Martinez, N. et al. The NTT transcription factor promotes replum development in Arabidopsis fruits. Plant J. 80, 69–81 (2014).
Crawford, B. C. W., Ditta, G. & Yanofsky, M. F. The NTT gene is required for transmitting-tract development in carpels of Arabidopsis thaliana. Curr. Biol. 17, 1101–1108 (2007).
Petricka, J. J., Clay, N. K. & Nelson, T. M. Vein patterning screens and the defectively organized tributaries mutants in Arabidopsis thaliana. Plant J. 56, 251–263 (2008).
Martin, A. et al. A transposon-induced epigenetic change leads to sex determination in melon. Nature 461, 1135–1138 (2009).
Sagasser, M., Lu, G. H., Hahlbrock, K. & Weisshaar, B. A. thaliana TRANSPARENT TESTA 1 is involved in seed coat development and defines the WIP subfamily of plant zinc finger proteins. Genes Dev. 16, 138–149 (2002).
Coen, O. et al. A TRANSPARENT TESTA transcriptional module regulates endothelium polarity. Front. Plant Sci. 10, 1801 (2019).
Appelhagen, I. et al. Weird fingers: functional analysis of WIP domain proteins. FEBS Lett. 584, 3116–3122 (2010).
Roldan, M. V. G. et al. Integrative genome-wide analysis reveals the role of WIP proteins in inhibition of growth and development. Commun. Biol. 3, 239 (2020).
Appelhagen, I. et al. TRANSPARENT TESTA1 interacts with R2R3-MYB factors and affects early and late steps of flavonoid biosynthesis in the endothelium of Arabidopsis thaliana seeds. Plant J. 67, 406–419 (2011).
Wieschaus, E. Positional information and cell fate determination in the early Drosophila embryo. Curr. Top. Dev. Biol. 117, 567–579 (2016).
Lynch, J. A. Evolution of maternal control of axial patterning in insects. Curr. Opin. Insect Sci. 31, 37–42 (2019).
Kölle, S., Hughes, B. & Steele, H. Early embryo-maternal communication in the oviduct: a review. Mol. Reprod. Dev. 87, 650–662 (2020).
Fazeli, A. Maternal communication with gametes and embryos. Theriogenology 70, 1182–1187 (2008).
Idelevich, A. & Vilella, F. Mother and embryo cross-communication. Genes https://doi.org/10.3390/genes11040376 (2020).
Ray, S., Golden, T. & Ray, A. Maternal effects of the short integument mutation on embryo development in Arabidopsis. Dev. Biol. 180, 365–369 (1996).
Costa, L. M. et al. Central cell-derived peptides regulate early embryo patterning in flowering plants. Science 344, 168–172 (2014).
Prigge, M. J. & Wagner, D. R. The Arabidopsis serrate gene encodes a zinc-finger protein required for normal shoot development. Plant Cell 13, 1263–1279 (2001).
Ottenschlager, I. et al. Gravity-regulated differential auxin transport from columella to lateral root cap cells. Proc. Natl Acad. Sci USA 100, 2987–2991 (2003).
Friml, J. et al. Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 426, 147–153 (2003).
Sarkar, A. K. et al. Conserved factors regulate signalling in Arabidopsis thaliana shoot and root stem cell organizers. Nature 446, 811–814 (2007).
Willemsen, V. et al. The NAC domain transcription factors FEZ and SOMBRERO control the orientation of cell division plane in Arabidopsis root stem cells. Dev. Cell 15, 913–922 (2008).
Petricka, J. J., Winter, C. M. & Benfey, P. N. Control of Arabidopsis root development. Annu. Rev. Plant Biol. 63, 563–590 (2012).
Scheres, B. Stem-cell niches: nursery rhymes across kingdoms. Nat. Rev. Mol. Cell Biol. 8, 345–354 (2007).
Craft, J. et al. New pOp/LhG4 vectors for stringent glucocorticoid-dependent transgene expression in Arabidopsis. Plant J. 41, 899–918 (2005).
Belles-Boix, E., Babiychuk, E., Van Montagu, M., Inze, D. & Kushnir, S. CEO1, a new protein from Arabidopsis thaliana, protects yeast against oxidative damage. FEBS Lett. 482, 19–24 (2000).
Jaspers, P. et al. Unequally redundant RCD1 and SRO1 mediate stress and developmental responses and interact with transcription factors. Plant J. 60, 268–279 (2009).
Teotia, S. & Lamb, R. S. The paralogous genes RADICAL-INDUCED CELL DEATH1 and SIMILAR TO RCD ONE1 have partially redundant functions during Arabidopsis development. Plant Physiol. 151, 180–198 (2009).
Teotia, S. & Lamb, R. S. RCD1 and SRO1 are necessary to maintain meristematic fate in Arabidopsis thaliana. J. Exp. Bot. 62, 1271–1284 (2011).
Christensen, L. F. & Staby, L. Evolutionary conservation of the intrinsic disorder-based Radical-Induced Cell Death1 hub interactome. Sci. Rep. 9, 18927 (2019).
Jaspers, P. et al. The RST and PARP-like domain containing SRO protein family: analysis of protein structure, function and conservation in land plants. BMC Genomics 11, 170 (2010).
Aravind, L. The WWE domain: a common interaction module in protein ubiquitination and ADP ribosylation. Trends Biochem. Sci. 26, 273–275 (2001).
Rissel, D. & Peiter, E. Poly(ADP-Ribose) polymerases in plants and their human counterparts: parallels and peculiarities. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20071638 (2019).
Wirthmueller, L. et al. Arabidopsis downy mildew effector HaRxL106 suppresses plant immunity by binding to RADICAL-INDUCED CELL DEATH1. New Phytol. 220, 232–248 (2018).
Bugge, K. et al. Structure of radical-induced cell death1 hub domain reveals a common αα-scaffold for disorder in transcriptional networks. Structure 26, 734–746.e7 (2018).
Stadler, R., Lauterbach, C. & Sauer, N. Cell-to-cell movement of green fluorescent protein reveals post-phloem transport in the outer integument and identifies symplastic domains in Arabidopsis seeds and embryos. Plant Physiol. 139, 701–712 (2005).
Kawashima, T. & Goldberg, R. B. The suspensor: not just suspending the embryo. Trends Plant Sci. 15, 23–30 (2010).
Yeung, E. C. Embryogeny of Phaseolus: the role of the suspensor. Z. Pflanzenphysiol. 96, 17–28 (1980).
Schulz, P. & Jensen, W. A. Capsella embryogenesis: the suspensor and the basal cell. Protoplasma 67, 139–163 (1969).
Robert, H. S. et al. Maternal auxin supply contributes to early embryo patterning in Arabidopsis. Nat. Plants 4, 548–553 (2018).
Nagl, W. Translocation of putrescine in the ovule, suspensor and embryo of Phaseolus coccineus. J. Plant Physiol. 136, 587–591 (1990).
Ahlfors, R. et al. Arabidopsis RADICAL-INDUCED CELL DEATH1 belongs to the WWE protein-protein interaction domain protein family and modulates abscisic acid, ethylene, and methyl jasmonate responses. Plant Cell 16, 1925–1937 (2004).
Brosche, M. et al. Transcriptomics and functional genomics of ROS-induced cell death regulation by RADICAL-INDUCED CELL DEATH1. PLoS Genet. 10, e1004112 (2014).
Potters, G., Pasternak, T. P., Guisez, Y. & Jansen, M. A. Different stresses, similar morphogenic responses: integrating a plethora of pathways. Plant Cell Environ. 32, 158–169 (2009).
Blomster, T. et al. Apoplastic reactive oxygen species transiently decrease auxin signaling and cause stress-induced morphogenic response in Arabidopsis. Plant Physiol. 157, 1866–1883 (2011).
Karimi, M., De Meyer, B. & Hilson, P. Modular cloning in plant cells. Trends Plant Sci. 10, 103–105 (2005).
Blilou, I. et al. The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433, 39–44 (2005).
Siligato, R. et al. MultiSite gateway-compatible cell type-specific gene-inducible system for plants. Plant Physiol. 170, 627–641 (2016).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Musielak, T. J., Schenkel, L., Kolb, M., Henschen, A. & Bayer, M. A simple and versatile cell wall staining protocol to study plant reproduction. Plant Reprod. 28, 161–169 (2015).
Bougourd, S., Marrison, J. & Haseloff, J. Technical advance: an aniline blue staining procedure for confocal microscopy and 3D imaging of normal and perturbed cellular phenotypes in mature Arabidopsis embryos. Plant J. 24, 543–550 (2000).
Zhou, X., Shi, C., Zhao, P. & Sun, M. Isolation of living apical and basal cell lineages of early proembryos for transcriptome analysis. Plant Reprod. 32, 105–111 (2019).
Figueiredo, D. D., Batista, R. A., Roszak, P. J., Hennig, L. & Kohler, C. Auxin production in the endosperm drives seed coat development in Arabidopsis. eLife https://doi.org/10.7554/eLife.20542 (2016).
Truernit, E. et al. High-resolution whole-mount imaging of three-dimensional tissue organization and gene expression enables the study of phloem development and structure in Arabidopsis. Plant Cell 20, 1494–1503 (2008).
Molder, F. et al. Sustainable data analysis with Snakemake. F1000Research 10, 33 (2021).
Gietz, R. D. & Woods, R. A. Yeast transformation by the LiAc/SS carrier DNA/PEG method. Methods Mol. Biol. 313, 107–120 (2006).
Acknowledgements
We thank M. Crespi and T. Blein for helpful discussions; B. Scheres for critical reading of the manuscript; H. Morin, C. Troadec, A. d. B. d. Granrut, all FLOCAD team members, the imaging platform and greenhouse teams at the Institute of Plant Sciences Paris-Saclay (IPS2) for technical support; and the Eurasian Arabidopsis Stock Centre (uNASC) for sharing research materials. This work was supported by the European Research Council (ERC-SEXYPARTH, 341076), the ANR (EPISEX, ANR-17-CE20-0019), and the LabEx Saclay Plant Sciences-SPS (ANR-10-LABX-40-SPS). M.V.G.R. was supported by the Intra-European Fellowships for Career Development (IEF) (Grant PIEF-GA-2012-330908).
Author information
Authors and Affiliations
Contributions
Y.D. and A. Bendahmane conceptualized the project; Y.D. and A. Bendahmane developed the methodology; A. Boualem and M.V. conducted formal analysis; Y.D., M.V.G.R., A.H., N.H. and F.I. conducted the investigations; Y.D., A.H. and F.I. procured resources; Y.D. wrote the original draft; Y.D., M.V.G.R., A. Boualem, M.V. and A. Bendahmane reviewed and edited the draft; Y.D. and A. Bendahmane supervised the project; A. Bendahmane acquired funding and administered the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 WIP genes regulate root cell division orientation.
a, Overview of wild type, wip245 (nww), wip2-4wip45 and wip2-3wip45 seedlings at 7 day-post-germination (d.p.g.). wip2-3 allele: SM_3_16705; wip2-4 allele: SM_3_23211. Scale bar: 1 cm. b-j, Images of wip245 and wip123456 embryonic roots at indicated stages. The number presented at the bottom of each image represents the counts of indicated phenotype (left) versus the total counts (right). G1: early-globular stage; G2: late-globular stage; H1: early-heart stage; H2: late-heart stage; ME: matured embryo. Magenta and blue frames: the zoom-in areas. White arrows in j indicate COL layer; white asterisks in j mark the newly formed COL cells; colored asterisks in h-i indicate possible cell division patterns from H1 to H2. Cyan: hypophysis/QC lineage; yellow: COL initial lineage; orange: COL layers; grey: delayed/failed layer formation; light purple: ground tissue initials; olive green: Epi/LRC initials; pink: vascular initials. QC: quiescent center; COL: columella; Epi: epidermis; LRC: lateral root cap. Scale bars: 50 μm. Related to Fig. 1.
Extended Data Fig. 2 Cell fate specification in wip123456 and wip245 embryonic roots.
a-e, Images of embryos, suspensors and primary roots expressing indicated reporters in wild type, wip245 and wip123456 mutants. Scale bars: 50 μm. h, Frequency and counts of the suspensors with or without DR5::GFP expression in their basal cells. Wild type, wip245, wip136 and wip123456 suspensors between globular and heart stage were sampled. Data in the frequency (the upper panel) represents mean ± s.d. from four biological replicates; sample size per replicate (n) =20. P values were calculated with two-tailed unpaired Student’s t test, mutant versus wild type: ***P < 0.005. Data in the suspensor counts (the lower panel) represents total number of examined suspensors, including but not restricting to the one used in the frequency experiments. Red lines in d,e highlight cell outlines of the wip245 hypophyseal derivatives; white arrows indicate the DR5::GFP expression in basal cells of the suspensor; white frames highlight cell outlines of the hypophyseal derivatives. Cyan dots: cells in hypophysis/QC lineage; yellow dots: cells in COL initial lineage; orange dots: cells in COL layers; grey dots: cells in delayed/failed layer formation. PR: primary root at 3 d.p.g.; QC: quiescent center; COL: columella. The experiments in a,b,f,d,e and c,g were repeated four and three times respectively, with similar results. Related to Fig. 2.
Extended Data Fig. 3 WIP1, WIP3 and WIP6 are maternally expressed.
a-d, Images of wild type, wip1 and proWIP1::cWIP1:VENUS complemented wip1 (L-12 and L-14) seeds. Scale bars: 1 mm. e-j, Images of wild type embryos, suspensors and their surrounding maternal tissues expressing indicated WIP reporters. Scale bars for e-h,j: 50 μm; scale bar for i: 1 mm. k-m, Images of wip2+/−45 siliques and developing seeds expressing indicated WIP reporters. Scale bar for k: 50 μm; scale bars for the silique panel in l,m: 1 mm; scale bars for the seed panel in l: 100 μm. n, RT-qPCR analysis of WIP1 and WIP3 transcription in wild type and wip2+/−45 siliques containing embryos between globular and heart stage. Data represents mean ± s.e.m. from three biological replicates, within each three technical repeats were included. P values were calculated with two-tailed unpaired Student’s t test, mutant versus wild type: *P < 0.05, **P < 0.01. o-q, Images of wip245 embryos and suspensors expressing indicated WIP reporters. Scale bars: 50 μm. White arrow in g: the proWIP3::gWIP3:VENUS signal. Cyan dots: cells in hypophysis/QC lineage; yellow dots: cells in COL initial lineage; orange dots: cells in COL layers. M: micropylar end; C: chalazal end; oi2: outer integument 2; oi1: outer integument 1 and ii1: inner integument 1 (endothelium); QC: quiescent center; COL: columella. The experiments in e-m and o-q were repeated three times, with similar results. Related to Fig. 2.
Extended Data Fig. 4 Embryonic expression of WIP genes promotes root formation.
a, GUS-staining of proWIP4::GUS in wild type siliques and developing seeds. Scale bar for the silique panel: 1 mm; scale bar for the seed panel: 100 μm. b, Images of wild type embryos, suspensors and primary roots expressing proWIP4::erCFP. Scale bars: 50 μm. c, Images of proWIP4::gWIP4:VENUS complemented wip245 embryonic and primary roots. Scale bar: 50 μm. d, Overview of wild type, wip245 and proWIP4::gWIP4:VENUS complemented wip245 (L-1 and L-5) seedlings at 7d.p.g.. Scale bars: 1 cm. e, Overview of wild type, wip245, proWIP4::cWIP1:VENUS complemented wip245 (L-5 and L-4) seedings at 7d.p.g.. Scale bars: 1 cm. Yellow frame and arrows: the proWIP4::gWIP4:VENUS signal in the uppermost suspensor cell; white frames highlight cell outlines of the hypophyseal derivatives. Cyan dots: cells in hypophysis/QC lineage; yellow dots: cells in COL initial lineage; orange dots: cells in COL layers. M: micropylar end; C: chalazal end; PR: primary root at 3 d.p.g.; QC: quiescent center; COL: columella. The experiments in a and b-e were repeated two and three times respectively, with similar results. Related to Fig. 2.
Extended Data Fig. 5 WIP1 inhibits plant growth via SRO family members.
a, Overview of wild type, rcd1-4, pOp6::cWIP1, 35::LhGR, DEX:WIP1, q195 and rcd1-4 DEX:WIP1 seedlings germinated on 1/2 MS medium supplemented with 30 nM DEX at 7 d.p.g.. Two biological replicates were performed. Scale bar: 1 cm. b, Left panel: overview of 35 S::LhGR, DEX:WIP1, q195 and rcd1-4 DEX:WIP1 seedlings grown on 1/2 MS medium supplemented with 30 nM DEX for 48 h. Right panel: quantification of the 48h-root growth. c, Left panel: overview of 35 S::LhGR, DEX:WIP1 and sro1 DEX:WIP1 seedlings grown on 1/2 MS medium supplemented with 30 nM DEX for 48 h. Right panel: quantification of the 48h-root growth. Black dots in b,c mark the root tip positions when the seedlings were freshly transferred, the 48-root growth was measured from the black dot to the root tip. Data represents mean ± s.e.m. from four biological replicates; sample size per replicate (n) =15. Mean value of the 48h-root growth on the mock medium is set to 100%. P values were calculated with two-tailed unpaired Student’s t test, 35 S::LhGR, q195, rcd1-4 DEX:WIP1 and sro1 DEX:WIP1 versus DEX:WIP1 respectively: ***P < 0.005. Scale bar: 1 cm. Related to Fig. 3.
Extended Data Fig. 6 RCD1 and SRO1 expression.
a, RT-qPCR analysis of RCD1 and SRO1 transcription in wild type siliques containing embryos between globular and heart stage. Data represents mean ± s.e.m. from two biological replicates, within each three technical repeats were included. P values were calculated with two-tailed unpaired Student’s t test, RCD1 versus SRO1: ***P < 0.005. b-e, Images of wild type siliques, developing seeds, embryos, suspensors and primary roots expressing indicated RCD1 and SRO1 reporters. Scale bars for b,d: 1 mm; scale bars for c,e: 50 μm. f-g, Images of proRCD1::gRCD1:VENUS complemented rcd1-4 and proSRO1::gSRO1:VENUS complemented sro1 embryos and primary roots. Scale bars: 50 μm. h, Frequency of wild type and rcd1-4sro1 roots with or without periclinally divided QC cells at indicated stages. Data represents mean ± s.d.; biological replicates (N) and sample size per replicate (n) are listed in Supplementary Table 6. P values were calculated with two-tailed unpaired Student’s t test, rcd1-4sro1 versus wild type: ***P < 0.005, P1 = 0.00098, P2 = 7.67E-05, P3 = 8.05E-05. H1: early-heart stage; H2: late-heart stage; ME: mature embryo. The experiments in b-g were repeated three times, with similar results. Related to Fig. 3.
Extended Data Fig. 7 The maternal WIPs act through SRO members to inhibit embryonic root formation.
a, mPS-PI staining of amyloplasts in wild type and rcd1-4wip245 primary roots. Scale bar: 100 μm. b, Quantification of COL layer numbers in wild type, rcd1-4wip245 and sro1wip245 mature embryos and primary roots. Data represents mean ± s.e.m.; biological replicates (N) and sample size per replicate (n) are listed in Supplementary Table 7. P values were calculated with two-tailed unpaired Student’s t test, mutant versus wild type: ***P < 0.005. c-e, sro1wip245 embryonic roots at indicated stages. The number presented at the bottom of each image represents the counts of indicated phenotype (left) versus the total counts (right). G1: early-globular stage; G2: late-globular stage; H1: early-heart stage; H2: late-heart stage; ME: matured embryo. Scale bars: 50 μm. f-h, Images of sro1wip245 embryos, suspensors and primary roots expressing indicated markers. White frames highlight cell outlines of the hypophyseal derivatives. Cyan dots: cells in hypophysis/QC lineage; yellow dots: cells in COL initial lineage; grey dots: cells in delayed/failed layer formation. Scale bars: 50 μm. Cyan: hypophysis/QC lineage; yellow: COL lineage; orange: COL layers; grey: delayed/failed layer formation; light purple: ground tissue initials; olive green: Epi/LRC initials. PR: primary root at 3 d.p.g.; QC: quiescent center; COL: columella; Epi: epidermis; LRC: lateral root cap. The experiments in a, f-g and h were repeated two, four and three times respectively, with similar results. Related to Fig. 4.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 1–8.
Supplementary Table
Supplementary Table 4 RNA-seq analysis of wild type and wip123456 primary root meristems. Supplementary Table 5 Frequency of embryonic roots with normal or delayed/failed layer formation. Supplementary Table 6 Frequency of wild type and rcd1-4sro1 roots with or without periclinally divided QC cells. Supplementary Table 7 Quantification of COL layer numbers. Supplementary Table 8 Source data for supplementary figures.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Du, Y., Roldan, M.V.G., Haraghi, A. et al. Spatially expressed WIP genes control Arabidopsis embryonic root development. Nat. Plants 8, 635–645 (2022). https://doi.org/10.1038/s41477-022-01172-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-022-01172-4
This article is cited by
-
A dialogue between generations
Nature Plants (2022)