Abstract
The universally conserved enzyme CTP synthase (CTPS) forms filaments in bacteria and eukaryotes. In bacteria, polymerization inhibits CTPS activity and is required for nucleotide homeostasis. Here we show that for human CTPS, polymerization increases catalytic activity. The cryo-EM structures of bacterial and human CTPS filaments differ considerably in overall architecture and in the conformation of the CTPS protomer, explaining the divergent consequences of polymerization on activity. The structure of human CTPS filament, the first structure of the full-length human enzyme, reveals a novel active conformation. The filament structures elucidate allosteric mechanisms of assembly and regulation that rely on a conserved conformational equilibrium. The findings may provide a mechanism for increasing human CTPS activity in response to metabolic state and challenge the assumption that metabolic filaments are generally storage forms of inactive enzymes. Allosteric regulation of CTPS polymerization by ligands likely represents a fundamental mechanism underlying assembly of other metabolic filaments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Noree, C., Sato, B.K., Broyer, R.M. & Wilhelm, J.E. Identification of novel filament-forming proteins in Saccharomyces cerevisiae and Drosophila melanogaster . J. Cell Biol. 190, 541–551 (2010).
Liu, J.L. Intracellular compartmentation of CTP synthase in Drosophila . J. Genet. Genomics 37, 281–296 (2010).
Ingerson-Mahar, M., Briegel, A., Werner, J.N., Jensen, G.J. & Gitai, Z. The metabolic enzyme CTP synthase forms cytoskeletal filaments. Nat. Cell Biol. 12, 739–746 (2010).
Carcamo, W.C. et al. Induction of cytoplasmic rods and rings structures by inhibition of the CTP and GTP synthetic pathway in mammalian cells. PLoS One 6, e29690 (2011).
Shen, Q.J. et al. Filamentation of metabolic enzymes in Saccharomyces cerevisiae . J. Genet. Genomics 43, 393–404 (2016).
Narayanaswamy, R. et al. Widespread reorganization of metabolic enzymes into reversible assemblies upon nutrient starvation. Proc. Natl. Acad. Sci. USA 106, 10147–10152 (2009).
An, S., Kumar, R., Sheets, E.D. & Benkovic, S.J. Reversible compartmentalization of de novo purine biosynthetic complexes in living cells. Science 320, 103–106 (2008).
Barry, R.M. et al. Large-scale filament formation inhibits the activity of CTP synthetase. eLife 3, e03638 (2014).
Strochlic, T.I. et al. Ack kinase regulates CTP synthase filaments during Drosophila oogenesis. EMBO Rep. 15, 1184–1191 (2014).
Petrovska, I. et al. Filament formation by metabolic enzymes is a specific adaptation to an advanced state of cellular starvation. eLife 3, e02409 (2014).
Noree, C., Monfort, E., Shiau, A.K. & Wilhelm, J.E. Common regulatory control of CTP synthase enzyme activity and filament formation. Mol. Biol. Cell 25, 2282–2290 (2014).
Chang, C.C. et al. Cytoophidium assembly reflects upregulation of IMPDH activity. J. Cell Sci. 128, 3550–3555 (2015).
Chen, K. et al. Glutamine analogs promote cytoophidium assembly in human and Drosophila cells. J. Genet. Genomics 38, 391–402 (2011).
Endrizzi, J.A., Kim, H., Anderson, P.M. & Baldwin, E.P. Crystal structure of Escherichia coli cytidine triphosphate synthetase, a nucleotide-regulated glutamine amidotransferase/ATP-dependent amidoligase fusion protein and homologue of anticancer and antiparasitic drug targets. Biochemistry 43, 6447–6463 (2004).
Goto, M., Omi, R., Nakagawa, N., Miyahara, I. & Hirotsu, K. Crystal structures of CTP synthetase reveal ATP, UTP, and glutamine binding sites. Structure 12, 1413–1423 (2004).
Endrizzi, J.A., Kim, H., Anderson, P.M. & Baldwin, E.P. Mechanisms of product feedback regulation and drug resistance in cytidine triphosphate synthetases from the structure of a CTP-inhibited complex. Biochemistry 44, 13491–13499 (2005).
Kursula, P. et al. Structure of the synthetase domain of human CTP synthetase, a target for anticancer therapy. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 62, 613–617 (2006).
Lauritsen, I., Willemoës, M., Jensen, K.F., Johansson, E. & Harris, P. Structure of the dimeric form of CTP synthase from Sulfolobus solfataricus . Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 67, 201–208 (2011).
Aughey, G.N. & Liu, J.L. Metabolic regulation via enzyme filamentation. Crit. Rev. Biochem. Mol. Biol. 51, 282–293 (2016).
Aughey, G.N. et al. Nucleotide synthesis is regulated by cytoophidium formation during neurodevelopment and adaptive metabolism. Biol. Open 3, 1045–1056 (2014).
Calise, S.J. et al. Glutamine deprivation initiates reversible assembly of mammalian rods and rings. Cell. Mol. Life Sci. 71, 2963–2973 (2014).
Martin, E. et al. CTP synthase 1 deficiency in humans reveals its central role in lymphocyte proliferation. Nature 510, 288–292 (2014).
Hofer, A., Steverding, D., Chabes, A., Brun, R. & Thelander, L. Trypanosoma brucei CTP synthetase: a target for the treatment of African sleeping sickness. Proc. Natl. Acad. Sci. USA 98, 6412–6416 (2001).
Hindenburg, A.A., Taub, R.N., Grant, S., Chang, G. & Baker, M.A. Effects of pyrimidine antagonists on sialic acid regeneration in HL-60 cells. Cancer Res. 45, 3048–3052 (1985).
Kang, G.J. et al. Cyclopentenylcytosine triphosphate. Formation and inhibition of CTP synthetase. J. Biol. Chem. 264, 713–718 (1989).
Politi, P.M. et al. Phase I clinical trial of continuous infusion cyclopentenyl cytosine. Cancer Chemother. Pharmacol. 36, 513–523 (1995).
Trudel, M., Van Genechten, T. & Meuth, M. Biochemical characterization of the hamster thy mutator gene and its revertants. J. Biol. Chem. 259, 2355–2359 (1984).
Raushel, F.M., Thoden, J.B. & Holden, H.M. Enzymes with molecular tunnels. Acc. Chem. Res. 36, 539–548 (2003).
Chang, Y.F., Martin, S.S., Baldwin, E.P. & Carman, G.M. Phosphorylation of human CTP synthetase 1 by protein kinase C: identification of Ser(462) and Thr(455) as major sites of phosphorylation. J. Biol. Chem. 282, 17613–17622 (2007).
Choi, M.G. & Carman, G.M. Phosphorylation of human CTP synthetase 1 by protein kinase A: identification of Thr455 as a major site of phosphorylation. J. Biol. Chem. 282, 5367–5377 (2007).
Choi, M.G., Park, T.S. & Carman, G.M. Phosphorylation of Saccharomyces cerevisiae CTP synthetase at Ser424 by protein kinases A and C regulates phosphatidylcholine synthesis by the CDP-choline pathway. J. Biol. Chem. 278, 23610–23616 (2003).
Han, G.S. et al. Expression of Human CTP synthetase in Saccharomyces cerevisiae reveals phosphorylation by protein kinase A. J. Biol. Chem. 280, 38328–38336 (2005).
Long, C.W. & Pardee, A.B. Cytidine triphosphate synthetase of Escherichia coli B. I. Purification and kinetics. J. Biol. Chem. 242, 4715–4721 (1967).
Habrian, C. et al. Inhibition of Escherichia coli CTP synthetase by NADH and other nicotinamides and their mutual interactions with CTP and GTP. Biochemistry 55, 5554–5565 (2016).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Lander, G.C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Egelman, E.H. The iterative helical real space reconstruction method: surmounting the problems posed by real polymers. J. Struct. Biol. 157, 83–94 (2007).
Sachse, C. et al. High-resolution electron microscopy of helical specimens: a fresh look at tobacco mosaic virus. J. Mol. Biol. 371, 812–835 (2007).
Egelman, E.H. A robust algorithm for the reconstruction of helical filaments using single-particle methods. Ultramicroscopy 85, 225–234 (2000).
Voss, N.R., Yoshioka, C.K., Radermacher, M., Potter, C.S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009).
Scheres, S.H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 54, 5.6.1–5.6.37 (2016).
DiMaio, F. et al. Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, W5–W9 (2008).
Katoh, K. & Standley, D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Acknowledgements
The authors are grateful to G. Carman (Rutgers University) for the hCTPS1-expressing S. cerevisiae strain. This work was supported by the US National Institutes of Health (R01 GM118396 to J.M.K.) and the Human Frontier Science Program (RGY0076/2013 to Z.G. and J.M.K.). Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research and by the National Institutes of Health, National Institute of General Medical Sciences (including P41GM103393). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIGMS or NIH.
Author information
Authors and Affiliations
Contributions
E.M.L. and J.M.K. wrote the manuscript. E.M.L., J.M.K., and D.R.H. solved the EM structures. J.A.E. and E.P.B. solved the crystal structure of ecCTPS. M.S. and J.M.H. generated disulfide cross-linked ecCTPS constructs and optimized purification and assembly conditions. E.M.L. and A.M. carried out CTPS activity assays. R.M.B. and Z.G. performed initial characterization of ecCTPS filaments. E.M.L., Z.G., E.P.B. and J.M.K. contributed to experimental design and data analysis and interpretation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cryo-EM reconstruction of ecCTPS.
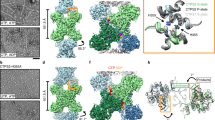
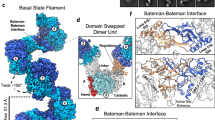
a, Cryo-electron micrograph of ecCTPS filaments. b, The resolution of the structure is 4.6 Å by gold-standard FSC. c, Local resolution map of one ecCTPS tetramer in the filament. d, A central slice through the unfiltered ecCTPS filament reconstruction shows less well-defined features in the GAT domains (yellow arrow) than in the AL domains (white arrowhead).
Supplementary Figure 2 Details of structural rearrangements at the CTP binding site that create new intersubunit contacts and drive the conformational switch to the filament-forming conformer.
a, The alternative ecCTPS-CTP complex structure (5TKV) is shown in yellow (subunit A), red (subunit B) and green (subunit A’, two-fold symmetry related to A, symmop 4). The previously published CTP complex (2AD5)16 is shown in gray, after superposition of the A subunits (residues 1-266, 0.42 Å Cα rmsd after superposition). Side chains that have altered interactions or conformations relative to 2AD5 are displayed. Hydrogen bonds unique to 5TKV are shown as cyan dotted lines, and heavy atom distances are indicated. The 2AD5 CTP is depicted using narrow sticks. In 5TKV, an octahedral Mg+2 ion coordinates the β- and γ-phosphate oxygens and 3 water molecules. Relative to the A subunit, helices α5(A’) and α7(A) Cα atoms shifted 1.9 Å and 1.1 Å respectively. F227 and N229 side chains changed rotamers, with the F227 phenyl ring packed against the CTP C5 atom, while N229 formed new inter-subunit hydrogen bonds with R158. These new hydrogen bonds became possible due to a quaternary rearrangement of subunits that was similar to the rearrangement of N-terminal domains in the filament-binding conformer observed by cryoEM. This structure validates the sequence conservation for these residues. Mutations at E155, R158 and position 229 confer pyrimidine drug resistance in CTPSs, presumably by hindering the conformational change required for formation of the filament (bacterial49) or inhibited tetramer (yeast50, mammalian51). Similar intersubunit contacts are observed in tetramer crystal structures of the M. tuberculosis CTPS-diUTP complex (PDB 4ZDJ), T. thermophilus CTPS-glutamine (PDB 1VCO) and apo (PDB 1VCM) complexes, and the hCTPS1 AL domain (PDB 2VO1). b, E155 and N229 participate in extended hydrogen bond networks in the A’ and B subunits. Hydrogen bonds specific to 5TKV are depicted in magenta, while those common to both 2AD5 and 5TKV are colored light blue. R158 and N229 form the center of this network. 49. Wylie, J.L., Wang, L.L., Tipples, G. & McClarty, G. A single point mutation in CTP synthetase of Chlamydia trachomatis confers resistance to cyclopentenyl cytosine. J Biol Chem 271, 15393-400 (1996). 50. Ostrander, D.B., O'Brien, D.J., Gorman, J.A. & Carman, G.M. Effect of CTP synthetase regulation by CTP on phospholipid synthesis in Saccharomyces cerevisiae. J Biol Chem 273, 18992-9001 (1998). 51. Whelan, J., Phear, G., Yamauchi, M. & Meuth, M. Clustered base substitutions in CTP synthetase conferring drug resistance in Chinese hamster ovary cells. Nat Genet 3, 317-22 (1993).
Supplementary Figure 3 CTP promotes disassembly of hCTPS1 filaments.
a, Adding glutamine to hCTPS1 filaments assembled with UTP, ATP, and GTP causes them to disassemble over time. b, Adding glutamine to hCTPS1 filaments assembled with UTP, AMPPNP, and GTP does not lead to filament disassembly, suggesting CTP synthesis is required to promoted hCTPS1 depolymerization. c, Directly adding CTP to hCTPS1 filaments assembled with UTP, ATP, and GTP causes filaments to disassemble. Scale bars are 50 nm.
Supplementary Figure 4 Cryo-EM reconstruction of hCTPS1.
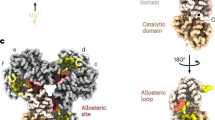
a, Section from a micrograph of frozen hCTPS1. b, FSC curve for the hCTPS1 filament, showing a resolution of 6.1Å. c, ResMap local resolution map of the hCTPS1 filament. d, Atomic model of the hCTPS1 monomer fit into the hCTPS1 filament cryoEM map, colored by domain. e, Individual helices are well-defined in the hCTPS1 filament cryoEM map. f, Comparison of the ecCTPS and hCTPS1 filament contact sites. hCTPS1 filaments lack the linker-linker contact site, and have a completely rearranged GAT contact site. g, Poorly-resolved C-terminal contact site (orange) in the hCTPS1 filament.
Supplementary Figure 5 Mutation of a conserved histidine in the eukaryotic GAT-domain insert abolishes hCTPS1 polymerization without disrupting tetramer formation.
a, CTPS sequence alignments, showing the appearance of the GAT domain helical insert early in eukaryotic evolution. The conserved histidine (H355 in humans) is shown in red. b, hCTPS1-H355A does not polymerize in the presence of UTP, ATP, and GTP. c, Negative stain reconstruction of the hCTPS1-H355A tetramer in the presence of substrates (grey), fit with an atomic model of hCTPS1 in the filament conformation (colored by protomer). Scale bar is 50 nm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5. (PDF 10328 kb)
Supplementary Video 1
Animation of ecCTPS tetramer morphing between the apo conformation and product-bound filament conformation. Monomers are shown in different colors, and filament contact sites are shown in red. (MOV 6109 kb)
Supplementary Video 2
Animation of ecCTPS tetramer morphing between the apo conformation and product-bound filament conformation (alternate view). Same animation as in Video 1, viewed down the filament axis. (MOV 3292 kb)
Supplementary Video 3
Animation of hCTPS1 tetramer morphing between the apo conformation and substrate-bound filament conformation. Monomers are shown in different colors, and filament contact sites are shown in red. (MOV 8709 kb)
Supplementary Video 4
Animation of hCTPS1 tetramer morphing between the apo conformation and substrate-bound filament conformation (alternate view). Same animation as in Video 3, viewed down the filament axis. (MOV 7319 kb)
Rights and permissions
About this article
Cite this article
Lynch, E., Hicks, D., Shepherd, M. et al. Human CTP synthase filament structure reveals the active enzyme conformation. Nat Struct Mol Biol 24, 507–514 (2017). https://doi.org/10.1038/nsmb.3407
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3407
This article is cited by
-
Structural basis of human PRPS2 filaments
Cell & Bioscience (2023)
-
Fine-mapping and candidate gene analysis of the Mcgy1 locus responsible for gynoecy in bitter gourd (Momordica spp.)
Theoretical and Applied Genetics (2023)
-
Agglomeration: when folded proteins clump together
Biophysical Reviews (2023)
-
Principles and functions of metabolic compartmentalization
Nature Metabolism (2022)
-
Rubisco forms a lattice inside alpha-carboxysomes
Nature Communications (2022)