Abstract
Heavy metals in cells are typically regulated by PIB-type ATPases. The first structure of the class, a Cu+-ATPase from Legionella pneumophila (LpCopA), outlined a copper transport pathway across the membrane, which was inferred to be occluded. Here we show by molecular dynamics simulations that extracellular water solvated the transmembrane (TM) domain, results indicative of a Cu+-release pathway. Furthermore, a new LpCopA crystal structure determined at 2.8-Å resolution, trapped in the preceding E2P state, delineated the same passage, and site-directed-mutagenesis activity assays support a functional role for the conduit. The structural similarities between the TM domains of the two conformations suggest that Cu+-ATPases couple dephosphorylation and ion extrusion differently than do the well-characterized PII-type ATPases. The ion pathway explains why certain Menkes' and Wilson's disease mutations impair protein function and points to a site for inhibitors targeting pathogens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lutsenko, S. & Kaplan, J.H. Organization of P-type ATPases: significance of structural diversity. Biochemistry 34, 15607–15613 (1995).
Kühlbrandt, W. Biology, structure and mechanism of P-type ATPases. Nat. Rev. Mol. Cell Biol. 5, 282–295 (2004).
Argüello, J.M., Eren, E. & Gonzalez-Guerrero, M. The structure and function of heavy metal transport P1B-ATPases. Biometals 20, 233–248 (2007).
Vulpe, C., Levinson, B., Whitney, S., Packman, S. & Gitschier, J. Isolation of a candidate gene for Menkes disease and evidence that it encodes a copper-transporting ATPase. Nat. Genet. 3, 7–13 (1993).
Bull, P.C., Thomas, G.R., Rommens, J.M., Forbes, J.R. & Cox, D.W. The Wilson disease gene is a putative copper transporting P-type ATPase similar to the Menkes gene. Nat. Genet. 5, 327–337 (1993).
Albers, R.W. Biochemical aspects of active transport. Annu. Rev. Biochem. 36, 727–756 (1967).
Post, R.L., Hegyvary, C. & Kume, S. Activation by adenosine triphosphate in the phosphorylation kinetics of sodium and potassium ion transport adenosine triphosphatase. J. Biol. Chem. 247, 6530–6540 (1972).
Toyoshima, C., Nakasako, M., Nomura, H. & Ogawa, H. Crystal structure of the calcium pump of sarcoplasmic reticulum at 2.6 A resolution. Nature 405, 647–655 (2000).
Toyoshima, C. & Nomura, H. Structural changes in the calcium pump accompanying the dissociation of calcium. Nature 418, 605–611 (2002).
Sørensen, T.L., Moller, J.V. & Nissen, P. Phosphoryl transfer and calcium ion occlusion in the calcium pump. Science 304, 1672–1675 (2004).
Toyoshima, C. & Mizutani, T. Crystal structure of the calcium pump with a bound ATP analogue. Nature 430, 529–535 (2004).
Olesen, C., Sorensen, T.L., Nielsen, R.C., Moller, J.V. & Nissen, P. Dephosphorylation of the calcium pump coupled to counterion occlusion. Science 306, 2251–2255 (2004).
Olesen, C. et al. The structural basis of calcium transport by the calcium pump. Nature 450, 1036–1042 (2007).
Winther, A.M. et al. The sarcolipin-bound calcium pump stabilizes calcium sites exposed to the cytoplasm. Nature 495, 265–269 (2013).
Toyoshima, C. et al. Crystal structures of the calcium pump and sarcolipin in the Mg2+-bound E1 state. Nature 495, 260–264 (2013).
Møller, J.V., Olesen, C., Winther, A.M. & Nissen, P. The sarcoplasmic Ca2+-ATPase: design of a perfect chemi-osmotic pump. Q. Rev. Biophys. 43, 501–566 (2010).
Mattle, D. et al. On allosteric modulation of P-type Cu-ATPases. J. Mol. Biol. 425, 2299–2308 (2013).
Gourdon, P. et al. Crystal structure of a copper-transporting PIB-type ATPase. Nature 475, 59–64 (2011).
González-Guerrero, M., Eren, E., Rawat, S., Stemmler, T.L. & Arguello, J.M. Structure of the two transmembrane Cu+ transport sites of the Cu+-ATPases. J. Biol. Chem. 283, 29753–29759 (2008).
Jardetzky, O. Simple allosteric model for membrane pumps. Nature 211, 969–970 (1966).
Petrek, M. et al. CAVER: a new tool to explore routes from protein clefts, pockets and cavities. BMC Bioinformatics 7, 316 (2006).
Shannon, R.D. Revised effective ionic-radii and systematic studies of interatomic distances in halides and chalcogenides. Acta Crystallogr. A 32, 751–767 (1976).
Huang, Z. & Tajkhorshid, E. Identification of the third Na+ site and the sequence of extracellular binding events in the glutamate transporter. Biophys. J. 99, 1416–1425 (2010).
Renthal, R. Buried water molecules in helical transmembrane proteins. Protein Sci. 17, 293–298 (2008).
Baginski, E.S., Foa, P.P. & Zak, B. Microdetermination of inorganic phosphate, phospholipids, and total phosphate in biologic materials. Clin. Chem. 13, 326–332 (1967).
Gourdon, P., Sitsel, O., Karlsen, J.L., Moller, L.B. & Nissen, P. Structural models of the human copper P-type ATPases ATP7A and ATP7B. Biol. Chem. 393, 205–216 (2012).
Barry, A.N. et al. The lumenal loop Met672-Pro707 of copper-transporting ATPase ATP7A binds metals and facilitates copper release from the intramembrane sites. J. Biol. Chem. 286, 26585–26594 (2011).
Tadini-Buoninsegni, F. et al. ATP dependent charge movement in ATP7B Cu+-ATPase is demonstrated by pre-steady state electrical measurements. FEBS Lett. 584, 4619–4622 (2010).
Lewis, D. et al. Distinctive features of catalytic and transport mechanisms in mammalian Ca2+ (SERCA) and Cu+ (ATP7A/B) ATPases. J. Biol. Chem. 278, 32717–32727 (2012).
Hatori, Y. et al. Intermediate phosphorylation reactions in the mechanism of ATP utilization by the copper ATPase (CopA) of Thermotoga maritima. J. Biol. Chem. 283, 22541–22549 (2008).
Poulsen, H. et al. Neurological disease mutations compromise a C-terminal ion pathway in the Na+/K+-ATPase. Nature 467, 99–102 (2010).
Hollenstein, K., Dawson, R.J. & Locher, K.P. Structure and mechanism of ABC transporter proteins. Curr. Opin. Struct. Biol. 17, 412–418 (2007).
Raimunda, D., Gonzalez-Guerrero, M., Leeber, B.W. III & Arguello, J.M. The transport mechanism of bacterial Cu+-ATPases: distinct efflux rates adapted to different function. Biometals 24, 467–475 (2011).
Waldron, K.J. et al. Structure and metal loading of a soluble periplasm cuproprotein. J. Biol. Chem. 285, 32504–32511 (2010).
Hassani, B.K., Astier, C., Nitschke, W. & Ouchane, S. CtpA, a copper-translocating P-type ATPase involved in the biogenesis of multiple copper-requiring enzymes. J. Biol. Chem. 285, 19330–19337 (2010).
Ward, S.K., Abomoelak, B., Hoye, E.A., Steinberg, H. & Talaat, A.M. CtpV: a putative copper exporter required for full virulence of Mycobacterium tuberculosis. Mol. Microbiol. 77, 1096–1110 (2010).
Festa, R.A. & Thiele, D.J. Copper at the front line of the host-pathogen battle. PLoS Pathog. 8, e1002887 (2012).
Hub, J.S., de Groot, B.L. & van der Spoel, D. g_wham: a free weighted histogram analysis implementation including robust error and autocorrelation estimates. J. Chem. Theory Comput. 6, 3713–3720 (2010).
Schneider, C.A., Rasband, W.S. & Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Petersen, C. & Moller, L.B. Control of copper homeostasis in Escherichia coli by a P-type ATPase, CopA, and a MerR-like transcriptional activator, CopR. Gene 261, 289–298 (2000).
Gourdon, P. et al. HiLiDe: systematic approach to membrane protein crystallization in lipid and detergent. Cryst. Growth Des. 11, 2098–2106 (2011).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Jo, S., Lim, J.B., Klauda, J.B. & Im, W. CHARMM-GUI membrane builder for mixed bilayers and its application to yeast membranes. Biophys. J. 97, 50–58 (2009).
Sansom, M.S., Scott, K.A. & Bond, P.J. Coarse-grained simulation: a high-throughput computational approach to membrane proteins. Biochem. Soc. Trans. 36, 27–32 (2008).
Lomize, M.A., Lomize, A.L., Pogozheva, I.D. & Mosberg, H.I. OPM: orientations of proteins in membranes database. Bioinformatics 22, 623–625 (2006).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Phillips, J.C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Klauda, J.B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Mackerell, A.D., Feig, M. & Brooks, C.L. Extending the treatment of backbone energetics in protein force fields: Limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Jorgensen, W.L., Chandrasekhar, J., Madura, J.D., Impey, R.W. & Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Grubmüller, H., Heller, H., Windemuth, A. & Schulten, K. Generalized verlet algorithm for efficient molecular dynamics simulations with long-range interactions. Mol. Simul. 6, 121–142 (1991).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an n.log(n) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Ryckaert, J.P., Ciccotti, G. & Berendsen, H.J.C. Numerical-integration of cartesian equations of motion of a system with constraints: molecular-dynamics of N-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Miyamoto, S. & Kollman, P.A. Settle: an analytical version of the shake and rattle algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992).
Feller, S.E., Zhang, Y.H., Pastor, R.W. & Brooks, B.R. Constant-pressure molecular-dynamics simulation: the Langevin piston method. J. Chem. Phys. 103, 4613–4621 (1995).
Martyna, G.J., Tobias, D.J. & Klein, M.L. Constant-pressure molecular-dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Oostenbrink, C., Villa, A., Mark, A.E. & van Gunsteren, W.F. A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J. Comput. Chem. 25, 1656–1676 (2004).
Poger, D., Van Gunsteren, W.F. & Mark, A.E. A new force field for simulating phosphatidylcholine bilayers. J. Comput. Chem. 31, 1117–1125 (2010).
Berendsen, H.J.C., Postma, J.P.M., van Gunsteren, W.F. & Hermans, J. in Intermolecular Forces (ed. Pullman, B.) (Reidel, Dordrecht, 1981).
Hoover, W.G. Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A 31, 1695–1697 (1985).
Nose, S. A unified formulation of the constant temperature molecular-dynamics methods. J. Chem. Phys. 81, 511–519 (1984).
Nose, S. & Klein, M.L. Constant pressure molecular-dynamics for molecular-systems. Mol. Phys. 50, 1055–1076 (1983).
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals: a new molecular-dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Hess, B., Bekker, H., Berendsen, H.J.C. & Fraaije, J.G.E.M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Kumar, S., Bouzida, D., Swendsen, R.H., Kollman, P.A. & Rosenberg, J.M. The weighted histogram analysis method for free-energy calculations on biomolecules: 1. The method. J. Comput. Chem. 13, 1011–1021 (1992).
Acknowledgements
We thank J.L. Karlsen for support on crystallographic computing. M.A. was supported in part by a Senior Postdoctoral Fellowship from the US National Science Foundation I2CAM International Materials Institute and a Marie Curie Career Integration Grant (FP7-MC-CIG-618558). The research was also supported by US National Institutes of Health grants GM074637 and GM086685 (S.H.W.). D.M. and O.S. were supported by the Graduate School of Science and Technology at Aarhus University. P.N. was supported by an advanced research grant (Biomemos) of the European Research Council. MD simulations were supported in part by the US National Science Foundation through TeraGrid (now Xsede) resources provided by the Texas Advanced Computing Center at the University of Texas at Austin.
Author information
Authors and Affiliations
Contributions
M.A. performed MD simulations; L.B.M. provided the in vivo strain and plasmid; D.M., O.S., T.K. and A.M.N. established DNA constructs; D.M. and O.S. performed the protein purifications; O.S. executed activity measurements; D.M. crystallized proteins, processed data and solved the crystal structures, guided by P.N. and P.G.; and all authors analyzed results. M.A. and P.G. designed the project, supervised by S.H.W. and P.N.; M.A., D.M., O.S. and P.G. generated figures; and all authors contributed to the writing of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
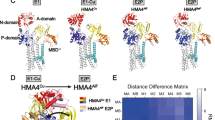
Supplementary Figure 1 Comparison of the E2.Pi and E2P states of LpCopA and SERCA.
(a), Domain arrangements in the E2P states of LpCopA and SERCA. LpCopA is shown as in Fig. 1b and SERCA (PDB 3B9B13) in green. Superimpositions were made of the intracellular domains using super in PYMOL. TM helices MA and MB, M7-M10 as well as SERCA insertions have been removed for clarity. (b-e), Close-up view of the phosphorylation site of LpCopA and SERCA. The A and P domains (LpCopA and SERCA) are indicated by colors as in Fig. 1a and the catalytic aspartate (Asp426 in LpCopA) and selected residues important for (de)phosphorylation are shown as sticks. (b), The E2P state in complex with BeF3- (Be in black, F in pink) and the Mg2+ ion (green) associated with Asp426. (c), The subsequent (forward reaction) E2.Pi conformation with AlF4- (Al in brown, F in pink) and the Mg2+ ion (green) associated with Asp426 (PDB 3RFU18). Note the shift in the position of the A domain relative to the P domain between the conformations. (d), Equivalent view as in (b) for the E2P state of SERCA (PDB 3B9B13). (e), Equivalent view as in (c) for the E2.Pi state of SERCA (PDB 3B9R13).
Supplementary Figure 2 Two E2P crystal forms of LpCopA.
(a-d), Crystal packing with the domains are colored as in Fig. 1b. The proteins are arranged as stacked bilayers, held together by hydrophobic interactions between their membrane-spanning regions, typical of the HiLiDe crystallization method41. (a), View along the membrane bilayer for the C2 (high-resolution) crystal form. (b), Equivalent view for the P212121 crystal form. Approximate perpendicular views of (a) and (b) are shown in (c) and (d), respectively. Note the more loose packing of the C2 lattice. The unit cell parameters can be found in Table 1. (e), Electron density maps of the low resolution P212121 form of the E2-BeF3- complex. The search model is based on the high-resolution structure (C2 space group) and is colored as in Fig. 1b. The 2mFo-DFc (blue, 1σ contour level) and mFo-DFc (±14σ contour levels, green and red, respectively) electron density maps were derived upon rigid body refinement and indicate no major deviations from the high-resolution structure
Supplementary Figure 3 MD simulation analyses.
(a), Evolution of the simulation box cell height. (b), The corresponding relative cell area (X,Y) evolution. (c-d), Backbone RMSD measured over the MD trajectory for states E2.Pi and E2P, and E2.Pi mutants P94A and P710A in the full protein (c) and the TM domain (d). (e), Centers of mass in x, y and z dimensions of the intracellular domains during the E2.Pi simulation. (f-g), Residence times for water molecules within 7 Å of Glu189 associated with the release pathway in the E2.Pi (f) and E2P (g) simulations. Two crystal waters remained associated with the internal water pockets within the release pathway for the entire E2P simulation and hence have residency times of 85 ns (not included in (f)). (h-i), Radii analyses. To determine structural variations of the release pathway within the 10 ns average from the MD simulations presented in Fig. 3d, the last 10 ns of the E2P (h) and E2.Pi (i) simulations were divided into 10 equally spaced 100 ps averages and subjected to Caver analyses.
Supplementary Figure 4 Functional analysis of mutant LpCopA forms.
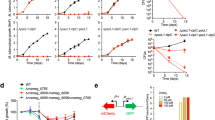
(a), Purity of the assayed constructs used for the in vitro assay. SDS-PAGE gels of the purified LpCopA constructs following a first round of scaling using ImageJ. Identical relative amounts of protein were used for the Baginski assay25 for generating the raw data shown in Supplementary Table 1. (b-c), The in vivo copper susceptibility assay. E. coli growth complementation curves from three independent experiments for wild-type LpCopA (WT, red), the inactive Asp426Asn mutant (gray), the high-affinity coordinator mutant Met717Val (yellow) as well as various release pathway mutants are displayed. Experiments 1 and 3 used three replicates each, experiment 2 used 15 replicates. 0 and 3 mM copper ((b) and (c), respectively) were used in the growth medium. (d), Assessment of the reproducibility of the in vitro assay. The in vitro data shown in Fig. 4c are based on a single experiment (experiment 1, with nine replicates), so wild type, Asp426Asn, Ala714Thr and Met100Glu LpCopA were reproduced in additional experiments (experiment 2, with six replicates). The two independent sets of data are highly consistent (merged data).
Supplementary Figure 5 Support of a proline-dependent opening mechanism.
(a-b), Helix dynamics in the E2.Pi simulation with LpCopA colored as in Fig. 1b. (a), Calculated inter-helical distances with spheres indicating Cα atoms. Three equally interspersed distances (along the normal to the membrane) in the TM domain were measured for each TM helix pair; gray (intracellular end), red and cyan (extracellular end) dotted lines between helices MA:M6, M1:M4 and M4:M5. (b), The helix-pair distances in (a) plotted as gray (intracellular end), red and cyan (extracellular end) lines. (c-g), Altered hydration patterns for simulations of the Pro94Ala (purple) and Pro710Ala (blue) mutants compared to wild-type (black). (c), Pore radius analyses of average structures from the E2.Pi simulations. (d), Structural representation of the pores predicted by CAVER. (e), Number of water molecules associated with the release pathway. (f), Average representations from the simulations of wild type (solid) and the Pro94Ala mutant (transparent) with the side chains of Pro94, Met717 and Glu189 pinpointed. Water is shown as red (wild type) and green (mutant) iso-density surfaces at 22 % occupancy. (g), Equivalent view as in (f) for wild type (solid, red water) and the Pro710Ala mutant (transparent, blue water).
Supplementary Figure 6 Sequence and secondary-structure conservation of the PIB-1, PIB-2 and PIB-4 ATPases.
(a), Alignment and sequence conservation of three representative members of the , and ATPases. The sequences used have locus tags lpg1024 (CopA), JW3434 (ZntA) and msmeg_5403 (CtpD). For analysis of conservation level, the ConSurf server was used72. Determination of conservation was based on 617 sequences having a sequence identity below 95% for ATPases, 520 sequences below identity=99% for ATPases and 607 sequences below identity=99% for ATPases. Information from (b) was used for adjusting the alignments of helices MA and MB due to poor sequence conservation in this region. The black box indicates the position of Pro94 in LpCopA. (b), Secondary structure based alignment of the N-terminal segment of the ATPase core of , and ATPases. Ten weakly redundant members of each subgroup were selected and aligned based on secondary structure prediction using the PSIPRED server73. Only the region from the start of helix MA to MB' is shown. The UniProtKB entries of the sequences are indicated to the left. Highlighted in bold are first sequences in each subgroup which are also used in (a).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 1638 kb)
Movie sequence showing every 200 ps of the non-equilibrated and equilibrated E2.Pi trajectory.
The protein is colored as in Fig. 1b and important residues are displayed in sticks. Water molecules within 3 Å of the protein are represented as red and white vdW spheres. (MOV 28136 kb)
Movie sequence showing every 200 ps of the non-equilibrated and equilibrated E2P trajectory.
The protein is colored as in Fig. 1b and important residues are displayed in sticks. Water molecules within 3 Å of the protein are represented as red and white vdW spheres. (MOV 29641 kb)
Rights and permissions
About this article
Cite this article
Andersson, M., Mattle, D., Sitsel, O. et al. Copper-transporting P-type ATPases use a unique ion-release pathway. Nat Struct Mol Biol 21, 43–48 (2014). https://doi.org/10.1038/nsmb.2721
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2721
This article is cited by
-
Structural basis of ion uptake in copper-transporting P1B-type ATPases
Nature Communications (2022)
-
Stabilization of supramolecular membrane protein–lipid bilayer assemblies through immobilization in a crystalline exoskeleton
Nature Communications (2021)
-
Tracking Membrane Protein Dynamics in Real Time
The Journal of Membrane Biology (2021)
-
CtpB is a plasma membrane copper (I) transporting P-type ATPase of Mycobacterium tuberculosis
Biological Research (2020)
-
CRISPR-mediated gene correction links the ATP7A M1311V mutations with amyotrophic lateral sclerosis pathogenesis in one individual
Communications Biology (2020)