Abstract
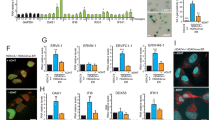
Trim24 (Tif1α) and Trim33 (Tif1γ) interact to form a co-repressor complex that suppresses murine hepatocellular carcinoma. Here we show that Trim24 and Trim33 cooperatively repress retinoic acid receptor–dependent activity of VL30-class endogenous retroviruses (ERVs) in liver. In Trim24-knockout hepatocytes, VL30 derepression leads to accumulation of reverse-transcribed VL30 cDNA in the cytoplasm that correlates with activation of the viral-defense interferon responses mimicking the preneoplastic inflammatory state seen in human liver following exogenous viral infection. Furthermore, upon derepression, VL30 long terminal repeats (LTRs) act as promoter and enhancer elements deregulating expression of neighboring genes and generating enhancer RNAs that are required for LTR enhancer activity in hepatocytes in vivo. These data reinforce the role of the TRIM family of proteins in retroviral restriction and antiviral defense and provide an example of an ERV-derived oncogenic regulatory network.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Feschotte, C. & Gilbert, C. Endogenous viruses: insights into viral evolution and impact on host biology. Nat. Rev. Genet. 13, 283–296 (2012).
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat. Genet. 42, 631–634 (2010).
Lynch, V.J., Leclerc, R.D., May, G. & Wagner, G.P. Transposon-mediated rewiring of gene regulatory networks contributed to the evolution of pregnancy in mammals. Nat. Genet. 43, 1154–1159 (2011).
Wolf, D. & Goff, S.P. Embryonic stem cells use ZFP809 to silence retroviral DNAs. Nature 458, 1201–1204 (2009).
Rowe, H.M. et al. KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463, 237–240 (2010).
McNab, F.W., Rajsbaum, R., Stoye, J.P. & O′Garra, A. Tripartite-motif proteins and innate immune regulation. Curr. Opin. Immunol. 23, 46–56 (2011).
Nisole, S., Stoye, J.P. & Saib, A. TRIM family proteins: retroviral restriction and antiviral defence. Nat. Rev. Microbiol. 3, 799–808 (2005).
Kawai, T. & Akira, S. Regulation of innate immune signalling pathways by the tripartite motif (TRIM) family proteins. EMBO Mol. Med. 3, 513–527 (2011).
Herquel, B., Ouararhni, K. & Davidson, I. The TIF1α-related TRIM cofactors couple chromatin modifications to transcriptional regulation, signaling and tumor suppression. Transcription 2, 231–236 (2011).
Tsai, W.W. et al. TRIM24 links a non-canonical histone signature to breast cancer. Nature 468, 927–932 (2010).
Agricola, E., Randall, R.A., Gaarenstroom, T., Dupont, S. & Hill, C.S. Recruitment of TIF1γ to chromatin via Its PHD finger-bromodomain activates its ubiquitin ligase and transcriptional repressor activities. Mol. Cell 43, 85–96 (2011).
Khetchoumian, K. et al. Loss of Trim24 (Tif1α) gene function confers oncogenic activity to retinoic acid receptor alpha. Nat. Genet. 39, 1500–1506 (2007).
Khetchoumian, K. et al. Trim24 (Tif1α): an essential ′brake′ for retinoic acid-induced transcription to prevent liver cancer. Cell Cycle 7, 3647–3652 (2008).
Herquel, B. et al. Transcription cofactors TRIM24, TRIM28, and TRIM33 associate to form regulatory complexes that suppress murine hepatocellular carcinoma. Proc. Natl. Acad. Sci. USA 108, 8212–8217 (2011).
French, N.S. & Norton, J.D. Structure and functional properties of mouse VL30 retrotransposons. Biochim. Biophys. Acta 1352, 33–47 (1997).
Nilsson, M. & Bohm, S. Inducible and cell type-specific expression of VL30 U3 subgroups correlate with their enhancer design. J. Virol. 68, 276–288 (1994).
Tisserand, J. et al. Tripartite motif 24 (Trim24/Tif1α) tumor suppressor protein is a novel negative regulator of interferon (IFN)/signal transducers and activators of transcription (STAT) signaling pathway acting through retinoic acid receptor α (Rarα) inhibition. J. Biol. Chem. 286, 33369–33379 (2011).
Bhoj, V.G. & Chen, Z.J. Linking retroelements to autoimmunity. Cell 134, 569–571 (2008).
Stetson, D.B., Ko, J.S., Heidmann, T. & Medzhitov, R. Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell 134, 587–598 (2008).
Islam, T.C. & Toftgard, R. Nuclear orphan receptor-binding retinoic acid response elements in keratinocytes. Biochem. Biophys. Res. Commun. 203, 545–552 (1994).
Benhaddou, A. et al. Transcription factor TEAD4 regulates expression of Myogenin and the unfolded protein response genes during C2C12 cell differentiation. Cell Death Differ. 19, 220–231 (2012).
Kim, T.K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Kowalczyk, M.S. et al. Intragenic enhancers act as alternative promoters. Mol. Cell 45, 447–458 (2012).
Koch, F. et al. Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters. Nat. Struct. Mol. Biol. 18, 956–963 (2011).
Bulger, M. & Groudine, M. Functional and mechanistic diversity of distal transcription enhancers. Cell 144, 327–339 (2011).
Islam, T.C., Bugge, T.H. & Bohm, S. The long terminal repeat of VL30 retrotransposons contains sequences that determine retinoic acid-induced transcription in cultured keratinocytes. J. Biol. Chem. 268, 3251–3259 (1993).
Brunmeir, R. et al. Epigenetic regulation of a murine retrotransposon by a dual histone modification mark. PLoS Genet. 6, e1000927 (2010).
Nielsen, A.L. et al. Interaction with members of the heterochromatin protein 1 (HP1) family and histone deacetylation are differentially involved in transcriptional silencing by members of the TIF1 family. EMBO J. 18, 6385–6395 (1999).
Crow, Y.J. & Rehwinkel, J. Aicardi-Goutieres syndrome and related phenotypes: linking nucleic acid metabolism with autoimmunity. Hum. Mol. Genet. 18, R130–R136 (2009).
Hatakeyama, S. TRIM proteins and cancer. Nat. Rev. Cancer 11, 792–804 (2011).
Sun, B. & Karin, M. Obesity, inflammation, and liver cancer. J. Hepatol. 56, 704–713 (2012).
Alison, M.R., Nicholson, L.J. & Lin, W.R. Chronic inflammation and hepatocellular carcinoma. Recent Results Cancer Res. 185, 135–148 (2011).
Crow, Y.J. et al. Mutations in genes encoding ribonuclease H2 subunits cause Aicardi-Goutieres syndrome and mimic congenital viral brain infection. Nat. Genet. 38, 910–916 (2006).
Crow, Y.J. et al. Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 cause Aicardi-Goutieres syndrome at the AGS1 locus. Nat. Genet. 38, 917–920 (2006).
Ørom, U.A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Ørom, U.A. & Shiekhattar, R. Long non-coding RNAs and enhancers. Curr. Opin. Genet. Dev. 21, 194–198 (2011).
Ho, Y., Elefant, F., Liebhaber, S.A. & Cooke, N.E. Locus control region transcription plays an active role in long-range gene activation. Mol. Cell 23, 365–375 (2006).
De Santa, F. et al. A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol. 8, e1000384 (2010).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Choukrallah, M.A. et al. Interconversion between active and inactive TATA-binding protein transcription complexes in the mouse genome. Nucleic Acids Res. 40, 1446–1459 (2012).
Ye, T. et al. seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res. 39, e35 (2011).
Acknowledgements
We thank the following members of the Institute de Génétique et de Biologie Moléculaire et Cellulaire, Illkirch, France: P. Koebel for AAV production, M. Cervino for mouse genotyping, Y. Lutz for microscopy, D. Umlauf for gift of digitonin, M. Philipps, and S. Vicaire for high-throughput sequencing. We thank the Penn Vector Core, School of Medicine Gene Therapy Program, University of Pennsylvania for the pAAV2/9 plasmid. This work was supported by institutional grants from the Centre National de la Recherche Scientifique, the Institut National de Sante et de la Recherche Médicale, the Université de Strasbourg, the Association pour la Recherche contre le Cancer, the Ligue Nationale contre le Cancer, and the Institut National du Cancer grant No 2008-037, the EuTRACC program of the European Union and the French Agence National pour la Recherche. I.D. is supported as an 'équipe labellisée' of the Ligue Nationale contre le Cancer. B.H. was supported by a fellowship from the Association pour la Recherche contre le Cancer.
Author information
Authors and Affiliations
Contributions
B.H. performed all the RNA-seq, qPCR and ChIP-seq and generated and maintained the different mouse lines together with K.O.; F.C. and K.O. generated and analyzed the MEFs; I.M. performed the Trim24 ChIPs; S.L.G., T.Y. and C.K. performed the bioinformatics analysis of the ChIP-seq and RNA-seq data; T.L. performed the 5′ RACE; B.J. generated the libraries for the ChIP-seq and RNA-seq; B.H., R.L. and I.D. designed experiments analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 12139 kb)
Supplementary Data set
Supplementary Table 1 (XLS 221 kb)
Supplementary Data set
Supplementary Table 2 (XLS 53 kb)
Supplementary Data set
Supplementary Table 3 (XLS 78 kb)
Supplementary Data set
Supplementary Table 4 (XLS 48 kb)
Rights and permissions
About this article
Cite this article
Herquel, B., Ouararhni, K., Martianov, I. et al. Trim24-repressed VL30 retrotransposons regulate gene expression by producing noncoding RNA. Nat Struct Mol Biol 20, 339–346 (2013). https://doi.org/10.1038/nsmb.2496
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2496
This article is cited by
-
TRIM proteins in hepatocellular carcinoma
Journal of Biomedical Science (2022)
-
KDM5B promotes immune evasion by recruiting SETDB1 to silence retroelements
Nature (2021)
-
The mouse HP1 proteins are essential for preventing liver tumorigenesis
Oncogene (2020)
-
Functional TRIM24 degrader via conjugation of ineffectual bromodomain and VHL ligands
Nature Chemical Biology (2018)
-
A somatic role for the histone methyltransferase Setdb1 in endogenous retrovirus silencing
Nature Communications (2018)