Abstract
Cancer therapies that target epigenetic repressors can mediate their effects by activating retroelements within the human genome. Retroelement transcripts can form double-stranded RNA (dsRNA) that activates the MDA5 pattern recognition receptor1,2,3,4,5,6. This state of viral mimicry leads to loss of cancer cell fitness and stimulates innate and adaptive immune responses7,8. However, the clinical efficacy of epigenetic therapies has been limited. To find targets that would synergize with the viral mimicry response, we sought to identify the immunogenic retroelements that are activated by epigenetic therapies. Here we show that intronic and intergenic SINE elements, specifically inverted-repeat Alus, are the major source of drug-induced immunogenic dsRNA. These inverted-repeat Alus are frequently located downstream of ‘orphan’ CpG islands9. In mammals, the ADAR1 enzyme targets and destabilizes inverted-repeat Alu dsRNA10, which prevents activation of the MDA5 receptor11. We found that ADAR1 establishes a negative-feedback loop, restricting the viral mimicry response to epigenetic therapy. Depletion of ADAR1 in patient-derived cancer cells potentiates the efficacy of epigenetic therapy, restraining tumour growth and reducing cancer initiation. Therefore, epigenetic therapies trigger viral mimicry by inducing a subset of inverted-repeats Alus, leading to an ADAR1 dependency. Our findings suggest that combining epigenetic therapies with ADAR1 inhibitors represents a promising strategy for cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-sequencing and CUT&RUN data have been deposited at the Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo) under the accession number GSE145639. Source data are provided with this paper.
Code availability
Code is available at https://github.com/smarhon/IR.
Change history
02 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03329-1
References
Roulois, D. et al. DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162, 961–973 (2015).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Sheng, W. et al. LSD1 ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell 174, 549–563 (2018).
Cuellar, T. L. et al. Silencing of retrotransposons by SETDB1 inhibits the interferon response in acute myeloid leukemia. J. Cell Biol. 216, 3535–3549 (2017).
Guler, G. D. et al. Repression of stress-induced LINE-1 expression protects cancer cell subpopulations from lethal drug exposure. Cancer Cell 32, 221–237 (2017).
Loo Yau, H., Ettayebi, I. & De Carvalho, D. D. The cancer epigenome: exploiting its vulnerabilities for immunotherapy. Trends Cell Biol. 29, 31–43 (2019).
Jones, P. A., Ohtani, H., Chakravarthy, A. & De Carvalho, D. D. Epigenetic therapy in immune-oncology. Nat. Rev. Cancer 19, 151–161 (2019).
Deaton, A. M. & Bird, A. CpG islands and the regulation of transcription. Genes Dev. 25, 1010–1022 (2011).
Levanon, E. Y. et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nat. Biotechnol. 22, 1001–1005 (2004).
Liddicoat, B. J. et al. RNA editing by ADAR1 prevents MDA5 sensing of endogenous dsRNA as nonself. Science 349, 1115–1120 (2015).
Leruste, A. et al. Clonally expanded T cells reveal immunogenicity of rhabdoid tumors. Cancer Cell 36, 597–612 (2019).
Krug, B. et al. Pervasive H3K27 acetylation leads to ERV expression and a therapeutic vulnerability in H3K27M gliomas. Cancer Cell 35, 782–797 (2019).
Ahmad, S. et al. Breaching self-tolerance to Alu duplex RNA underlies MDA5-mediated inflammation. Cell 172, 797–810 (2018).
Chung, H. et al. Human ADAR1 prevents endogenous RNA from triggering translational shutdown. Cell 172, 811–824 (2018).
Deininger, P. Alu elements: know the SINEs. Genome Biol. 12, 236 (2011).
Iorio, F. et al. A landscape of pharmacogenomic interactions in cancer. Cell 166, 740–754 (2016).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Gruber, A. J. et al. A comprehensive analysis of 3′ end sequencing data sets reveals novel polyadenylation signals and the repressive role of heterogeneous ribonucleoprotein C on cleavage and polyadenylation. Genome Res. 26, 1145–1159 (2016).
Samuel, C. E. Adenosine deaminases acting on RNA (ADARs) are both antiviral and proviral. Virology 411, 180–193 (2011).
Hou, F. et al. MAVS forms functional prion-like aggregates to activate and propagate antiviral innate immune response. Cell 146, 448–461 (2011).
Issa, J. P. & Kantarjian, H. M. Targeting DNA methylation. Clin. Cancer Res. 15, 3938–3946 (2009).
Ishizuka, J. J. et al. Loss of ADAR1 in tumours overcomes resistance to immune checkpoint blockade. Nature 565, 43–48 (2019).
Gannon, H. S. et al. Identification of ADAR1 adenosine deaminase dependency in a subset of cancer cells. Nat. Commun. 9, 5450 (2018).
Kreso, A. et al. Self-renewal as a therapeutic target in human colorectal cancer. Nat. Med. 20, 29–36 (2014).
Saito, Y. et al. Inhibition of DNA methylation suppresses intestinal tumor organoids by inducing an anti-viral response. Sci. Rep. 6, 25311 (2016).
Lima-Fernandes, E. et al. Targeting bivalency de-represses Indian Hedgehog and inhibits self-renewal of colorectal cancer-initiating cells. Nat. Commun 10, 1436 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Jun, G., Wing, M. K., Abecasis, G. R. & Kang, H. M. An efficient and scalable analysis framework for variant extraction and refinement from population-scale DNA sequence data. Genome Res. 25, 918–925 (2015).
Narasimhan, V. et al. BCFtools/RoH: a hidden Markov model approach for detecting autozygosity from next-generation sequencing data. Bioinformatics 32, 1749–1751 (2016).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165, (2016).
Kwon, M. & Firestein, B. L. DNA transfection: calcium phosphate method. Methods Mol. Biol. 1018, 107–110 (2013).
Halfmann, R. & Lindquist, S. Screening for amyloid aggregation by semi-denaturing detergent-agarose gel electrophoresis. J. Vis. Exp. 17, e838 (2008).
Ettayebi, I., Yau, H. L. & De Carvalho, D. D. Methods to detect endogenous dsRNA induction and recognition. Methods Enzymol. 629, 35–51 (2019).
Teissandier, A., Servant, N., Barillot, E. & Bourc’his, D. Tools and best practices for retrotransposon analysis using high-throughput sequencing data. Mob. DNA 10, 52 (2019).
Sergushichev, A. A. An algorithm for fast preranked gene set enrichment analysis using cumulative statistic calculation. Preprint at https://doi.org/10.1101/060012 (2016).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Acknowledgements
We thank S. Hur and X. Mu for providing the recombinant MDA5 proteins and the protocol for the MDA5-protection assay. We thank the Ontario Institute for Cancer Research (OICR) Genomics for conducting library preparation and RNA-sequencing for the MDA5-protection assay. We also thank the Princess Margaret Cancer Centre Genomics core (PM Genomics) for library preparation and RNA sequencing. This work was funded by Canadian Institute of Health Research (CIHR), New Investigator salary award (201512MSH360794-228629), Helen M Cooke professorship from Princess Margaret Cancer Foundation, Canada Research Chair, CIHR Foundation Grant (FDN 148430), CIHR Project Grant (PJT 165986), NSERC (489073) and Ontario Institute for Cancer Research (OICR) with funds from the province of Ontario to D.D.D.C. P.M. was supported by a Princess Margaret Post-doctoral fellowship- Hold’em for life. I.E. was supported by the Canadian Institutes of Health Research Canada Graduate Scholarships—Masters Award (CGS-M). F.A.d.C. was supported by FAPESP fellowship (2015/21237-4).
Author information
Authors and Affiliations
Contributions
P.M., S.A.M., I.E. and D.D.D.C. contributed to the study design. P.M. and I.E. performed the in vitro experiments with technical support from A.H., F.A.d.C., H.L.Y., C.I. and S.A. P.M. and Y.W. performed the in vivo experiments, C.A.O. contributed to designing the in vivo experiments. S.A.M. performed the bioinformatics data analysis and developed the RNA-editing and the inverted-repeat pipelines. A.C. performed public DNA methylation data analysis. P.M., S.A.M., I.E. and D.D.D.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.D.D.C. received research funds from Pfizer and Nektar therapeutics. D.D.D.C. is co-founder and shareholder of DNAMx, Inc. All the other authors declare no conflict of interest.
Additional information
Peer review information Nature thanks Gordon Carmichael, William Haining, Kenneth Nephew and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Detecting dsRNA forming repeat element transcription after 5-AZA-CdR treatment.
a, Effect of 5-AZA-CdR on expression of repeat elements in patient-derived xenograft CRC cells after 5 days of treatment. Plots show log-transformed fold change for 5-AZA-CdR-treated cells compared with mock-treated cells for antisense (y axis) and sense (x axis) transcripts of each SINE, LINE and ERV repeat element (CPM). Green dots in the top-right quadrant represent repeats that are significantly upregulated in 5-AZA-CdR-treated condition at both the antisense and sense strands as compared to the mock condition. Blue dots in the top-right quadrant represent repeats that have baseline expression (CPM ≥ 1) at the sense strand and are significantly upregulated at the antisense strand after 5-AZA-CdR treatment. Red dots in the top-right quadrant represent the repeats that have baseline expression (CPM ≥ 1) at the antisense strand and are significantly upregulated at the sense strand after 5-AZA-CdR treatment. Significance was determined as P < 0.05 and |logFC| ≥ 1 at each strand, BH (Benjamini–Hochberg)-corrected for multiple testing. b, c, Counts of expressed Alus in terms of the log10-transformed fold change of their MDA5-protected expression compared with total CytoRNA expression for baseline immunogenic RNA (n = 187 non-IR-Alus and 1,602 IR-Alus) (b), and the treatment-induced immunogenic RNA (n = 992 non-IR-Alus and 5,199 IR-Alus) (c). Histograms on the left and right show the count of Alus from mock- and 5-AZA-CdR-treated cells respectively. The colour code represents Alus making IR-Alus (red) and non-IR Alus (blue). IR-Alus that are MDA5 dsRNA agonists have positive values on the x axis. d, e, Scatterplots showing the log10-transformed fold change of MDA5-protected compared with total CytoRNA for each repeat in the pair of Alus (y axis represents the fold change in the first repeat, x axis represents the fold change in the second repeat in the pair) identified for baseline immunogenic RNA (n = 1,040 IR-Alu pairs) (d) and treatment-induced immunogenic RNA (n = 3,687 IR-Alu pairs) (e).
Extended Data Fig. 2 Detecting immunogenic RNAs that are induced after treatment with 5-AZA-CdR using MDA5/RNase-protection assay.
a, MDA5/RNase-protection assay experimental scheme created with BioRender.com. To identify the primary ligand for MDA5, total cytosolic RNA (5 ng μl−1) was purified from mock or 5-AZA-CdR treated (300 nM for 5 days) patient-derived CRC cells and pre-incubated with MDA5-∆2CARD protein, followed by either RNase A (+RNase A) digestion or without RNase A digestion (−RNase A). The remaining RNA was then purified and sequenced. b, Venn diagram showing the number of IR-Alu pairs at the baseline (n = 1,040 IR-Alu pairs), immunogenic (5-AZA-CdR) (n = 4,482 IR-Alu pairs) and treatment-induced (n = 3,687 IR-Alu pairs) conditions. c, Bar plot showing the number of baseline immunogenic RNA (IR-Alu pairs = 1,040, non-IR Alu pairs = 97). d, Bar plot showing the number of treatment-induced immunogenic RNA (IR-Alu pairs = 3,687, non-IR Alu pairs = 547). e, f, Average reads per million (RPM) profile of baseline MDA5-protected inverted-repeat transcripts (n = 1,176 transcript regions) (e) and treatment-induced MDA5-protected inverted-repeat transcripts (n = 3,895 transcript regions) (f). The box plots include the RPM scores of these transcripts. P value was calculated using a two-sided Wilcoxon signed-rank test. Box plot statistics containing the minimum, first quantile, median, third quantile, and maximum are 4.76, 10.21, 15.15, 27.44, and 52.53, respectively, for the mock-treated sample, and 0.0, 7.69, 15.24, 29.96, and 63.35 for the 5-AZA-CdR-treated sample in e, and 0.00, 0.79, 2.976, 6.185, and 14.26 for the mock-treated sample, and 2.41, 5.47, 7.47, 10.98, and 19.22 for the 5-AZA-CdR-treated sample in f. g, Schematic representation of inverted-repeats intramolecular pairing versus sense and antisense transcription created with BioRender.com. Inverted repeats can form intramolecular stem–loops when one strand is transcribed, as well as intermolecular duplexes when both strands are transcribed.
Extended Data Fig. 3 Genomic distribution and regulatory features of baseline and treatment-induced IR-Alus.
a, b, Distribution of the IR-Alu pairs based on their genomic orientation for the baseline immunogenic RNA (a) and treatment-induced immunogenic RNA (b). Arrows represent the genomic orientation (5′ to 3′) and arrow colour represent the transcriptional orientation (blue for sense and red for antisense). The number of IR-Alu pairs is represented by n. c, Distribution of Alu subfamilies from baseline immunogenic IR-Alus (n = 1,602 IR Alus), treatment-induced immunogenic IR-Alus (n = 5,199 IR Alus), and IR-Alus present in the human genome (n = 641,262 IR Alus). Odds ratio shows depletion or enrichment for each Alu subfamily (AluS, AluJ and AluY) at baseline immunogenic IR-Alus or treatment-induced immunogenic IR-Alus compared to Alu subfamily distribution of all IR-Alus present in the human genome. ****P < 0.0001, two-sided Fisher exact test. d, Profile of average CpG density 50 kb upstream and 50 kb downstream of the baseline IR-Alu pairs and all existing IR-Alu pairs in the human genome. e, Profile of average bona-fide CpG island (CGI) intersection density flanking (−50 kb/+50 kb; upstream/downstream) the treatment-induced IR-Alu pairs (n = 3,687 IR-Alu pairs, red line) and all existing IR-Alu pairs in the human genome (n = 746,470 IR-Alu pairs, blue line). For treatment-induced IR-Alu pairs, orientation was based on the transcriptional orientation in the MDA5-protected RNA-seq data. f, Profile of average CpG island intersection density flanking (−50 kb/+50 kb; upstream/downstream) the baseline IR-Alu pairs (n = 1,040 IR-Alu pairs, red line) and all existing IR-Alu pairs in the human genome (n = 746,470 IR-Alu pairs, blue line). g–i, Representative genomic tracks of treatment-induced IR-Alu pairs located at intergenic regions (g, h) and baseline IR-Alu pair located at a 3′ UTR region (i). The top four tracks represent antisense transcription (in red) and sense transcription (in blue). The bottom track represents CpG density (green). j, Schematic representation of baseline and treatment-induced immunogenic IR-Alu pairs.
Extended Data Fig. 4 DNA methylation status of CGI adjacent to immunogenic IR-Alus and correlation of methylation score of immunogenic IR-Alus regulatory regions with viral mimicry signature.
a, b, Percentage of CGIs (n = 1338) directly upstream of 5-AZA-CdR induced immunogenic IR-Alus with at least one fully methylated (green), partially methylated (blue), or fully unmethylated (red) CpG site in CRC cells (n = 51) (a) or pan-cancer cell lines (n = 988) (b) from the GDSC project. c, d, Regions that after epigenetic therapy become H3K4me3 marked directly upstream of induced immunogenic IR-Alus (n = 991 peaks) with at least one fully methylated (green), partially methylated (blue), or fully unmethylated (red) CpG site in CRC cells (c) or pan-cancer cell lines (d) from the GDSC project. e, Scatter plot showing inverse correlation (r = −0.23, P = 7.58 × 10−13) between the DNA methylation score of regulatory regions that after epigenetic therapy become H3K4me3 marked directly upstream to the induced immunogenic IR-Alus (n = 991 H3K4me3 peaks) and viral mimicry ISG signature ssGSEA score (n = 22 ISGs). Each dot is one pan-cancer cell line (n = 988) from the GDSC project. The grey area represents the 95% confidence interval around the linear model fit represented by the black line. The test used was a two-sided Pearson’s correlation test, uncorrected for multiple testing.
Extended Data Fig. 5 Characterization of the cytoplasmic immunogenic IR-Alus.
a, b, Distribution of each known human polyA signal (PAS) motif with respect to the distance from the end of the downstream Alu of each IR-Alu pair in the set of baseline immunogenic IR-Alus (n = 1,040 IR-Alu pairs) (a) and the set of treatment-induced immunogenic IR-Alus (n = 3,687 IR-Alu pairs) (b). The y axis is the counts of the IR-Alu pairs that include the motif in the MDA5-protected RNA-seq, and the x axis is the distance in bp from the end of the downstream Alu in an IR-Alu pair. c, d, Heatmap and average profile of the MDA5-protected RNA-seq signal centred at the PAS locations detected in a and b. Each row represents the downstream Alu for each IR-Alu pair. The orientation and the strand are based on the MDA5-protected transcriptional orientation. e, Percentage of non-repeats and several families of repetitive elements in total RNA-seq (nuclear and cytoplasmic RNA) and in RNA-seq from the total cytoplasmic fraction after 5-AZA-CdR treatment at 300 nM for 5 days. The total Cyto RNA-Seq donut plot (on right) is as in Fig. 1b and plotted here for reference. The odds ratio for SINEs is 1.68 (P < 2.2 × 10−16) and 2.21 (P < 2.2 × 10−16) for Alus. Odds ratio was calculated between total cytoplasmic RNA-seq and total RNA-seq. ****P < 0.0001, two-sided Fisher exact test. f, Representative confocal microscopy images from two independent experiments of mock-treated and 5-AZA-CdR-treated ADAR1WT patient-derived xenograft CRC cells. DNA was stained with DAPI (blue) and dsRNA was stained using the J2 antibody (red). Scale bar, 50 μm.
Extended Data Fig. 6 ADAR1 induces immunogenic inverted Alus.
a, dsRNA quantification based on J2/DAPI staining measured by ImageJ. Data are mean ± s.d. from n = 20 randomly sampled regions of two independent experiments. ***P < 0.001, ****P < 0.0001, Dunnett-corrected ordinary one-way ANOVA. b, qPCR analysis of total ADAR1 mRNA level after treatment with 5-AZA-CdR (300 nM, for 5 days) or transfection with 100 ng ml−1 poly(I:C) over a course of 24 days. Cells were washed out at day 5 and seeded in drug-free medium. Data are mean ± s.e.m. (n = 3 from three independent experiments). ****P < 0.0001, Sidak-corrected two-way ANOVA. c, Venn-diagram showing the number of IR-Alu pairs at baseline (n = 1,040 IR-Alu pairs), ADAR1KD-baseline immunogenic (n = 9,030 IR-Alu pairs) and ADAR1KD-induced immunogenic conditions. d, e, log10-transformed fold change of MDA5-protected total CytoRNA enriched IR-Alus and non-IR Alus for baseline immunogenic RNA (d) and ADAR1KD-induced immunogenic RNA (e). The histogram on the left shows the count of IR-Alus from the ADAR1WT cells, and the histogram on the right shows the counts from the ADAR1KD-treated cells. The colour code represents Alus making inverted-repeats (red) and non-IR-Alus (blue) for baseline immunogenic RNAs (n = 187 non-IR-Alus and 1,602 IR-Alus) (d; ADAR1WT data are the same as mock-treated data in Extended Data Fig. 1b and plotted here for reference) and for the ADAR1KD-induced immunogenic RNA (n = 1,684 non-IR-Alus and 11,085 IR-Alus) (e). f, g, Scatterplots showing the log10-transformed fold change of MDA5-protected over total CytoRNA for each repeat in the pair of Alus (y axis represents the fold change in the first repeat, x axis represents the fold change in the second repeat in the pair) identified for baseline immunogenic RNA (n = 1,040 IR-Alu pairs) (f; ADAR1WT data are the same as mock-treated data in Extended Data Fig. 1d) ADAR1KD patient-derived CRCs (n = 8,148 IR-Alu pairs) (g). h, i, Transcriptional orientation of each IR-Alu pair identified as immunogenic RNA at baseline (n = 1,040 IR-Alu pairs) (h) and ADAR1KD-induced immunogenic RNA (n = 8,148 IR-Alu pairs) (i). Each bar shows the count of IR-pairs where both repeats are in the sense strand (+/+, blue), antisense strand (−/−, red), or discordant strands (+/− or −/+). The plot on the left shows the counts in the RNA-seq data from the ADAR1WT cells and the plot on the right shows the counts in the RNA-seq data from the ADAR1KD cells. j, k, Average RPM profile of baseline MDA5-protected RNA transcripts (n = 1,176 transcript regions) that include baseline IR-Alus (j; ADAR1WT data are the same as mock-treated data in Extended Data Fig. 2e), and ADAR1KD-induced MDA5-protected transcripts (n = 8,432 transcript regions) that include ADAR1KD-induced IR-Alus (k). The box plots indicate the RPM scores of these transcripts. P values determined by two-sided Wilcoxon signed-rank test. Box plot statistics that contain the minimum, first quantile, median, third quantile, and maximum are 4.76, 10.21, 15.15, 27.44, and 52.53, respectively, for ADAR1WT, and 0, 7.52, 14.71, 28.38, and 59.37 for the ADAR1KD box plots in j, and 0, 0, 2.55, 5.06, and 12.65 for the ADAR1WT and 1.21, 3.17, 4.62, 7.23, and 13.32 for the ADAR1KD box plots in k.
Extended Data Fig. 7 ADAR1 depletion synergizes with anti-tumour effects of DNMTi through loss of its catalytic activity.
a, Representative immunoblot of two independent experiments showing MAVS aggregation analysed by SDS-AGE and MAVS protein level analysed by SDS–PAGE. VDAC served as a loading control in SDS–PAGE. b, ADAR1 isoforms (p110 and p150) relative expression analysed by qPCR. LACZ denotes ADAR1WT; PIC denotes 100 ng ml−1 poly (I:C) for 5 days; AZA denotes 300 nM 5-AZA-CdR for 5 days). Data are mean ± s.e.m. (n = 3 from three independent experiments). ***P < 0.001, ****P < 0.0001, Tukey-corrected two-way ANOVA. c, Kinetics of ISGs (ISG15, DDX58 and IRF7) relative expression in ADAR1WT (grey) and ADAR1KD (salmon) patient-derived CRCs treated with 5-AZA-CdR analysed by qPCR. Data are mean ± s.e.m. (n = 3 from three independent experiments). *P < 0.05, **P < 0.01, ****P < 0.0001, Sidak-corrected two-way ANOVA. d, Interferon-responsive ssGSEA score in ADAR1WT and ADAR1KD mock-treated and poly(I:C)-transfected samples collected 5 days, 14 days and 24 days after treatment. e, Kinetics of ISGs (ISG15, DDX58 and IRF7) relative expression in ADAR1WT (black) and ADAR1KD (pink) patient-derived CRCs transfected with poly(I:C), analysed by qPCR. Data are mean ± s.e.m. (n = 3 from three independent experiments). **P < 0.01, ****P < 0.0001, Sidak-corrected two-way ANOVA. f, qPCR (left) and immunoblot (right) analysis of ADAR1-p150 overexpression efficiency (ADAR1p150-OE) in patient-derived xenograft CRCs. Data are mean ± s.e.m. (n = 3 from three independent experiments). P = 0.0144, Wilcoxon test. Representative immunoblot of two independent experiments depicts two ADAR1 isoforms (p110 and p150). α-Tubulin served as a loading control. g, ISGs (ISG15, DDX58 and IRF7) relative expression in control (dark blue) and ADAR1p150-OE (light blue) patient-derived CRCs treated with 5-AZA-CdR, analysed by qPCR. Data are mean ± s.e.m. (n = 3 from three independent experiments). **P < 0.01, ***P < 0.001, Sidak-corrected two-way ANOVA. h, IFN-responsive ssGSEA score in mock-treated or 5-AZA-CdR-treated control and ADAR1p150-OE samples collected 5 days after treatment and measured by RNA-seq. Box plot statistics that contain the minimum, first quantile, median, third quantile, and maximum are 0.67, 0.73, 0.78, 0.84, and 0.90 respectively for 5-AZA-CdR control and 0.12, 0.23,0.35,0.46,0.58 for the ADAR1-p150-overexpressing 5-AZA-CdR samples. i, dsRNA quantification by ImageJ in control and ADAR1p150-OE cells mock-treated or treated with 5-AZA-CdR. Data are mean ± s.d. from n = 20 randomly sampled regions of two independent experiments. ****P < 0.0001, Tukey-corrected ordinary one-way ANOVA. ns, not significant. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 ADAR1 depletion synergizes with DNMTi in induction of ISGs in CRCs.
a, qPCR analysis of ADAR1 p150 and p110 isoforms relative expression in LIM1215 CRC cells. b, qPCR analysis of ISGs (MDA5, IRF7 and DDX58) relative expression in ADAR1WT and ADAR1KD LIM1215 CRCs mock-treated or treated with 300 nM 5-AZA-CdR for 5 days. c, qPCR analysis of ADAR1 p150 and p110 isoforms relative expression in HT29 CRCs. d, qPCR analysis of ISGs (MDA5, ISG15 and DDX58) relative expression in ADAR1WT and ADAR1KD HT29 CRCs mock-treated or treated with 300 nM 5-AZA-CdR for 5 days. Data are mean ± s.e.m. (n = 3 from three independent experiments). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, Tukey-corrected two-way ANOVA.
Extended Data Fig. 9 Epigenetic therapies which induce dsRNA can synergize with ADAR1 inhibition.
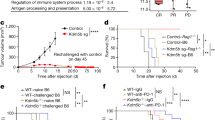
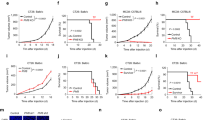
a, Colorectal cancer tumours dissected from vehicle and 5-AZA-CdR cohorts at day 14 after treatment are depicted for each condition (n = 5). b, Change in mean body weight of NSG mice transplanted with non-infected, ADAR1WT and ADAR1KDA/B patient-derived CRCs treated with vehicle or 5-AZA-CdR (0.5 mg kg−1 through intraperitoneal injection, for two cycles of 4 days with a 3 day break). c, Schematic representation of in vitro LDA in patient-derived xenograft CRCs mock-treated or treated with 5-AZA-CdR, created with BioRender.com. For each condition, cells were seeded as 1,000 cells per well, 100 cells per well, 10 cells per well and 1 cell per well. Spheroids were scored (absence or presence) 4 weeks after plating the cells. d, Normalized percentage of reduction in sphere forming ability calculated in each condition compared to mock- treated. e, Number of engrafted tumours compared to the number of injected tumours in each cohort for in vivo LDA. f, Representative confocal microscopy images from two independent experiments of ADAR1WT and ADAR1KDA cells treated with CDK4/6 inhibitor (250 nM palbociclib) for 7 days. DNA was stained with DAPI (blue) and dsRNA was stained using the J2 antibody (red). Scale bar, 50 μm. g, dsRNA quantification using ImageJ. Data are mean ± s.d. from n = 15 randomly sampled regions of two independent experiments. **P < 0.01, ***P < 0.001, Tukey-corrected ordinary one-way ANOVA. h, qPCR analysis of relative expression of ISGs (ISG15, DDX58 and IRF7) in cells treated with 250 nM palbociclib. Data are mean ± s.e.m., n = 3 from three independent experiments. ***P < 0.001, ****P < 0.0001, Tukey-corrected two-way ANOVA.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1: Uncropped gel source data, Supplementary Table 1: shRNA sequences used for ADAR1 knockdown, Supplementary Table 2: qPCR primers sequences used in this study and references for Methods and supplementary tables.
Rights and permissions
About this article
Cite this article
Mehdipour, P., Marhon, S.A., Ettayebi, I. et al. Epigenetic therapy induces transcription of inverted SINEs and ADAR1 dependency. Nature 588, 169–173 (2020). https://doi.org/10.1038/s41586-020-2844-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2844-1
This article is cited by
-
The role of ADAR1 through and beyond its editing activity in cancer
Cell Communication and Signaling (2024)
-
Towards targeting transposable elements for cancer therapy
Nature Reviews Cancer (2024)
-
Novel dual inhibitors of PARP and HDAC induce intratumoral STING-mediated antitumor immunity in triple-negative breast cancer
Cell Death & Disease (2024)
-
Regulation and function of transposable elements in cancer genomes
Cellular and Molecular Life Sciences (2024)
-
The potential of epigenetic therapy to target the 3D epigenome in endocrine-resistant breast cancer
Nature Structural & Molecular Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.