Abstract
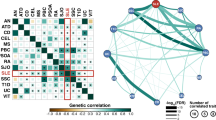
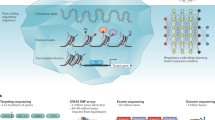
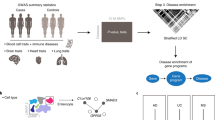
In the past decade, the availability and abundance of individual-level molecular data, such as gene expression, proteomics and sequence data, has enabled the use of integrative computational approaches to pose and answer novel questions about disease. In this article, we discuss several examples of applications of bioinformatics techniques to study autoimmune and rheumatic disorders. We focus our discussion on how integrative techniques can be applied to analyze gene expression and genetic variation data across different diseases, and discuss the implications of such analyses. We also outline current challenges and future directions of these approaches. We show that integrative computational methods are essential for translational research and provide a powerful opportunity to improve human health by refining the current knowledge about diagnostics, therapeutics and mechanisms of disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Barrett, T. et al. NCBI GEO: archive for high-throughput functional genomic data. Nucleic Acids Res. 37, D885–D890 (2009).
Butte, A. J. Translational bioinformatics applications in genome medicine. Genome Med. 1, 64 (2009).
Walsh, N. C., Crotti, T. N., Goldring, S. R. & Gravallese, E. M. Rheumatic diseases: the effects of inflammation on bone. Immunol. Rev. 208, 228–251 (2005).
Coward, J., Germain, R. N. & Altan-Bonnet, G. Perspectives for computer modeling in the study of T cell activation. Cold Spring Harb. Perspect. Biol. 2, a005538 (2010).
Germain, R. N. Computational analysis of T cell receptor signaling and ligand discrimination—past, present, and future. FEBS Lett. 584, 4814–4822 (2010).
Crow, M. K. & Wohlgemuth, J. Microarray analysis of gene expression in lupus. Arthritis Res. Ther. 5, 279–287 (2003).
Han, G. M. et al. Analysis of gene expression profiles in human systemic lupus erythematosus using oligonucleotide microarray. Genes Immun. 4, 177–186 (2003).
Booth, G. et al. Gene expression profiles at different stages of collagen-induced arthritis. Autoimmunity 41, 512–521 (2008).
Raychaudhuri, S. et al. Genetic variants at CD28, PRDM1 and CD2/CD58 are associated with rheumatoid arthritis risk. Nat. Genet. 41, 1313–1318 (2009).
Ptacek, T., Li, X., Kelley, J. M. & Edberg, J. C. Copy number variants in genetic susceptibility and severity of systemic lupus erythematosus. Cytogenet. Genome Res. 123, 142–147 (2008).
Graham, R. R., Hom, G., Ortmann, W. & Behrens, T. W. Review of recent genome-wide association scans in lupus. J. Intern. Med. 265, 680–688 (2009).
Freudenberg, J. et al. Locus category based analysis of a large genome-wide association study of rheumatoid arthritis. Hum. Mol. Genet. 19, 3863–3872 (2010).
Stahl, E. A. et al. Genome-wide association study meta-analysis identifies seven new rheumatoid arthritis risk loci. Nat. Genet. 42, 508–514 (2010).
Tsuchiya, N., Ito, I. & Kawasaki, A. Association of IRF5, STAT4 and BLK with systemic lupus erythematosus and other rheumatic diseases. Nihon Rinsho Meneki Gakkai Kaishi 33, 57–65 (2010).
Mackay, I. R. Clustering and commonalities among autoimmune diseases. J. Autoimmun. 33, 170–177 (2009).
Orozco, G. et al. Analysis of a functional BTNL2 polymorphism in type 1 diabetes, rheumatoid arthritis, and systemic lupus erythematosus. Hum. Immunol. 66, 1235–1241 (2005).
Wang, K. et al. Comparative genetic analysis of inflammatory bowel disease and type 1 diabetes implicates multiple loci with opposite effects. Hum. Mol. Genet. 19, 2059–2067 (2010).
Huang, W., Wang, P., Liu, Z. & Zhang, L. Identifying disease associations via genome-wide association studies. BMC Bioinformatics 10 (Suppl. 1), S68 (2009).
Torkamani, A., Topol, E. J. & Schork, N. J. Pathway analysis of seven common diseases assessed by genome-wide association. Genomics 92, 265–272 (2008).
Sirota, M., Schaub, M. A., Batzoglou, S., Robinson, W. H. & Butte, A. J. Autoimmune disease classification by inverse association with SNP alleles. PLoS Genet. 5, e1000792 (2009).
Dudley, J. T., Tibshirani, R., Deshpande, T. & Butte, A. J. Disease signatures are robust across tissues and experiments. Mol. Syst. Biol. 5, 307 (2009).
Ungethuem, U. et al. Molecular signatures and new candidates to target the pathogenesis of rheumatoid arthritis. Physiol. Genomics 42A, 267–282 (2010).
Chaussabel, D. et al. A modular analysis framework for blood genomics studies: application to systemic lupus erythematosus. Immunity 29, 150–164 (2008).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Iskar, M. et al. Drug-induced regulation of target expression. PLoS Comput. Biol. 6, e1000925 (2010).
Iorio, F. et al. Discovery of drug mode of action and drug repositioning from transcriptional responses. Proc. Natl Acad. Sci. USA 107, 14621–14626 (2010).
Garman, K. S. et al. A genomic approach to colon cancer risk stratification yields biologic insights into therapeutic opportunities. Proc. Natl Acad. Sci. USA 105, 19432–19437 (2008).
Setlur, S. R. et al. Estrogen-dependent signaling in a molecularly distinct subclass of aggressive prostate cancer. J. Natl Cancer Inst. 100, 815–825 (2008).
Pujol, A., Mosca, R., Farrés, J. & Aloy, P. Unveiling the role of network and systems biology in drug discovery. Trends Pharmacol. Sci. 31, 115–123 (2010).
Zhong, H., Yang, X., Kaplan, L. M., Molony, C. & Schadt, E. E. Integrating pathway analysis and genetics of gene expression for genome-wide association studies. Am. J. Hum. Genet. 86, 581–591 (2010).
Chen, R. et al. FitSNPs: highly differentially expressed genes are more likely to have variants associated with disease. Genome Biol. 9, R170 (2008).
Hsu, Y. H. et al. An integration of genome-wide association study and gene expression profiling to prioritize the discovery of novel susceptibility Loci for osteoporosis-related traits. PLoS Genet. 6, e1000977 (2010).
Chen, D. P. et al. Clinical arrays of laboratory measures, or “clinarrays”, built from an electronic health record enable disease subtyping by severity. AMIA Annu. Symp. Proc. 2007, 115–119 (2007).
Patel, C. J., Bhattacharya, J. & Butte, A. J. An Environment-Wide Association Study (EWAS) on type 2 diabetes mellitus. PLoS ONE 5, e10746 (2010).
Utz, P. J. Multiplexed assays for identification of biomarkers and surrogate markers in systemic lupus erythematosus. Lupus 13, 304–311 (2004).
Hueber, W. et al. Antigen microarray profiling of autoantibodies in rheumatoid arthritis. Arthritis Rheum. 52, 2645–2655 (2005).
Raychaudhuri, S. et al. Common variants at CD40 and other loci confer risk of rheumatoid arthritis. Nat. Genet. 40, 1216–1223 (2008).
Pointon, J. J. et al. Elucidating the chromosome 9 association with AS; CARD9 is a candidate gene. Genes Immun. 11, 490–496 (2010).
Zhang, H. G. et al. Novel tumor necrosis factor α-regulated genes in rheumatoid arthritis. Arthritis Rheum. 50, 420–431 (2004).
Acknowledgements
This work was supported by the Lucile Packard Foundation for Children's Health, the Hewlett Packard Foundation, US National Library of Medicine (R01 LM009719 and T15 LM007033), Howard Hughes Medical Institute, and Pharmaceutical Research and Manufacturers of America Foundation. We thank the anonymous reviewers for helping us improve the manuscript and members of the Butte Lab from Stanford University for constructive discussion.
Author information
Authors and Affiliations
Contributions
M. Sirota and A. J. Butte contributed equally to all aspects of preparation of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Sirota, M., Butte, A. The role of bioinformatics in studying rheumatic and autoimmune disorders. Nat Rev Rheumatol 7, 489–494 (2011). https://doi.org/10.1038/nrrheum.2011.87
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrrheum.2011.87