Key Points
-
Approximately 20% of cases of chronic kidney disease (CKD) that manifest before the age of 25 years are caused by single gene mutations in any one of >200 different genes
-
Molecular genetic diagnostics can provide patients with a molecular diagnosis for their disease and can generate new insights into disease mechanisms
-
Molecular genetic diagnostics might also have consequences for personalized treatment and prevention of CKD
-
Indication-driven mutation analysis panels are available to guide the genetic diagnosis of common early-onset kidney diseases, such as congenital anomalies of the kidneys and urinary tract, steroid-resistant nephrotic syndrome, ciliopathies and nephrolithiasis
Abstract
The primary causes of chronic kidney disease (CKD) in children differ from those of CKD in adults. In the USA the most common diagnostic groups of renal disease that manifest before the age of 25 years are congenital anomalies of the kidneys and urinary tract, steroid-resistant nephrotic syndrome, chronic glomerulonephritis and renal cystic ciliopathies, which together encompass >70% of early-onset CKD diagnoses. Findings from the past decade suggest that early-onset CKD is caused by mutations in any one of over 200 different monogenic genes. Developments in high-throughput sequencing in the past few years has rendered identification of causative mutations in this high number of genes feasible. Use of genetic analyses in patients with early onset-CKD will provide patients and their families with a molecular genetic diagnosis, generate new insights into disease mechanisms, facilitate aetiology-based classifications of patient cohorts for clinical studies, and might have consequences for personalized approaches to the prevention and treatment of CKD. In this Review, we discuss the implications of next-generation sequencing in clinical genetic diagnostics and the discovery of novel genes in early-onset CKD. We also delineate the resulting opportunities for deciphering disease mechanisms and the therapeutic implications of these findings.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Inker, L. A. et al. KDOQI US commentary on the 2012 KDIGO clinical practice guideline for the evaluation and management of CKD. Am. J. Kidney Dis. 63, 713–735 (2014).
Coresh, J. et al. Prevalence of chronic kidney disease in the United States. JAMA 298, 2038–2047 (2007).
Gee, H. Y. et al. Whole-exome resequencing distinguishes cystic kidney diseases from phenocopies in renal ciliopathies. Kidney Int. 85, 880–887 (2014).
Devuyst, O. et al. Rare inherited kidney diseases: challenges, opportunities, and perspectives. Lancet 383, 1844–1859 (2014).
Vivante, A., Kohl, S., Hwang, D. Y., Dworschak, G. C. & Hildebrandt, F. Single-gene causes of congenital anomalies of the kidney and urinary tract (CAKUT) in humans. Pediatr. Nephrol. 29, 695–704 (2014).
Saisawat, P. et al. Whole-exome resequencing reveals recessive mutations in TRAP1 in individuals with CAKUT and VACTERL association. Kidney Int. 85, 1310–1317 (2014).
Ebarasi, L. et al. Defects of CRB2 cause steroid-resistant nephrotic syndrome. Am. J. Hum. Genet. 96, 153–161 (2014).
Lovric, S. et al. Rapid detection of monogenic causes of childhood-onset steroid-resistant nephrotic syndrome. Clin. J. Am. Soc. Nephrol. 9, 1109–1116 (2014).
Kohl, S. et al. Mild recessive mutations in six Fraser syndrome-related genes cause isolated congenital anomalies of the kidney and urinary tract. J. Am. Soc. Nephrol. 25, 1917–1922 (2014).
Hwang, D. Y. et al. Mutations in 12 known dominant disease-causing genes clarify many congenital anomalies of the kidney and urinary tract. Kidney Int. 85, 1429–1433 (2014).
Hildebrandt, F. Genetic kidney diseases. Lancet 375, 1287–1295 (2010).
Smith, J. M., Stablein, D. M., Munoz, R., Hebert, D. & McDonald, R. A. Contributions of the Transplant Registry: the 2006 Annual Report of the North American Pediatric Renal Trials and Collaborative Studies (NAPRTCS). Pediatr. Transplant. 11, 366–373 (2007).
Harambat, J., van Stralen, K. J., Kim, J. J. & Tizard, E. J. Epidemiology of chronic kidney disease in children. Pediatr. Nephrol. 27, 363–373 (2012).
Kestila, M. et al. Positionally cloned gene for a novel glomerular protein — nephrin — is mutated in congenital nephrotic syndrome. Mol. Cell 1, 575–582 (1998).
Wiggins, R. C. The spectrum of podocytopathies: a unifying view of glomerular diseases. Kidney Int. 71, 1205–1214 (2007).
Somlo, S. & Mundel, P. Getting a foothold in nephrotic syndrome. Nat. Genet. 24, 333–335 (2000).
Vivante, A. et al. Mutations in TBX18 cause dominant urinary tract malformations via transcriptional dysregulation of ureter development. Am. J. Hum. Genet. 97, 291–301 (2015).
Lovric, S., Ashraf, S., Tan, W. & Hildebrandt, F. Genetic testing in steroid-resistant nephrotic syndrome: when and how? Nephrol. Dial. Transplant. http://dx.doi.org/10.1093/ndt/gfv355 (2015).
Trautmann, A. et al. Spectrum of steroid-resistant and congenital nephrotic syndrome in children: the PodoNet registry cohort. Clin. J. Am. Soc. Nephrol. 10, 592–600 (2015).
Hildebrandt, F., Benzing, T. & Katsanis, N. Ciliopathies. N. Engl. J. Med. 364, 1533–1543 (2011).
Altshuler, D., Daly, M. J. & Lander, E. S. Genetic mapping in human disease. Science 322, 881–888 (2008).
Chaki, M. et al. Genotype-phenotype correlation in 440 patients with NPHP-related ciliopathies. Kidney Int. 80, 1239–1245 (2011).
Palumbo, P. et al. Variable phenotype in 17q12 microdeletions: clinical and molecular characterization of a new case. Gene 538, 373–378 (2014).
Chen, Y. Z. et al. Systematic review of TCF2 anomalies in renal cysts and diabetes syndrome/maturity onset diabetes of the young type 5. Chinese Med. J. 123, 3326–3333 (2010).
Hasselbacher, K. et al. Recessive missense mutations in LAMB2 expand the clinical spectrum of LAMB2-associated disorders. Kidney Int. 70, 1008–1012 (2006).
Tory, K. et al. Mutation-dependent recessive inheritance of NPHS2-associated steroid-resistant nephrotic syndrome. Nat. Genet. 46, 299–304 (2014).
Genovese, G. et al. Association of trypanolytic ApoL1 variants with kidney disease in African Americans. Science 329, 841–845 (2010).
Tzur, S. et al. Missense mutations in the APOL1 gene are highly associated with end stage kidney disease risk previously attributed to the MYH9 gene. Hum. Genet. 128, 345–350 (2010).
Freedman, B. I. et al. The apolipoprotein L1 (APOL1) gene and nondiabetic nephropathy in African Americans. J. Am. Soc. Nephrol. 21, 1422–1426 (2010).
Friedman, D. J., Kozlitina, J., Genovese, G., Jog, P. & Pollak, M. R. Population-based risk assessment of APOL1 on renal disease. J. Am. Soc. Nephrol. 22, 2098–2105 (2011).
Genovese, G. et al. A risk allele for focal segmental glomerulosclerosis in African Americans is located within a region containing APOL1 and MYH9. Kidney Int. 78, 698–704 (2010).
Dummer, P. D. et al. APOL1 kidney disease risk variants: an evolving landscape. Seminars Nephrol. 35, 222–236 (2015).
Kopp, J. B. et al. APOL1 genetic variants in focal segmental glomerulosclerosis and HIV-associated nephropathy. J. Am. Soc. Nephrol. 22, 2129–2137 (2011).
Khanna, H. et al. A common allele in RPGRIP1L is a modifier of retinal degeneration in ciliopathies. Nat. Genet. 41, 739–745 (2009).
Deltas, C., Pierides, A. & Voskarides, K. Molecular genetics of familial hematuric diseases. Nephrol. Dial. Transplant. 28, 2946–2960 (2013).
MacArthur, D. G. et al. Guidelines for investigating causality of sequence variants in human disease. Nature 508, 469–476 (2014).
Bell, C. J. et al. Carrier testing for severe childhood recessive diseases by next-generation sequencing. Sci. Transl. Med. 3, 65ra64 (2011).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Sanna-Cherchi, S. et al. Mutations in DSTYK and dominant urinary tract malformations. N. Engl. J. Med. 369, 621–629 (2013).
Sethi, S. & Fervenza, F. C. Membranoproliferative glomerulonephritis — a new look at an old entity. N. Engl. J. Med. 366, 1119–1131 (2012).
Halbritter, J. et al. High-throughput mutation analysis in patients with a nephronophthisis-associated ciliopathy applying multiplexed barcoded array-based PCR amplification and next-generation sequencing. J. Med. Genet. 49, 756–767 (2012).
Halbritter, J. et al. Identification of 99 novel mutations in a worldwide cohort of 1,056 patients with a nephronophthisis-related ciliopathy. Hum. Genet. 132, 865–884 (2013).
George, J. N. & Nester, C. M. Syndromes of thrombotic microangiopathy. N. Engl. J. Med. 371, 654–666 (2014).
Sadowski, C. E. et al. A single-gene cause in 29.5% of cases of steroid-resistant nephrotic syndrome. J. Am. Soc. Nephrol. 26, 1279–1289 (2015).
Halbritter, J. et al. Fourteen monogenic genes account for 15% of nephrolithiasis/nephrocalcinosis. J. Am. Soc. Nephrol. 26, 543–551 (2014).
Braun, D. A. et al. Whole exome sequencing identifies causative mutations in the majority of consanguineous or familial cases with childhood-onset increased renal echogenicity. Kidney Int. http://dx.doi.org/10.1038/ki.2015.317 (2015).
Sanna-Cherchi, S. et al. Copy-number disorders are a common cause of congenital kidney malformations. Am. J. Hum. Genet. 91, 987–997 (2012).
Weber, S. et al. Mapping candidate regions and genes for congenital anomalies of the kidneys and urinary tract (CAKUT) by array-based comparative genomic hybridization. Nephrol. Dial. Transplant. 26, 136–143 (2011).
Ruf, R. G. et al. Patients with mutations in NPHS2 (podocin) do not respond to standard steroid treatment of nephrotic syndrome. J. Am. Soc. Nephrol. 15, 722–732 (2004).
Weber, S. et al. NPHS2 mutation analysis shows genetic heterogeneity of steroid-resistant nephrotic syndrome and low post-transplant recurrence. Kidney Int. 66, 571–579 (2004).
Chernin, G. et al. Genotype/phenotype correlation in nephrotic syndrome caused by WT1 mutations. Clin. J. Am. Soc. Nephrol. 5, 1655–1662 (2010).
Montini, G., Malaventura, C. & Salviati, L. Early coenzyme Q10 supplementation in primary coenzyme Q10 deficiency. N. Engl. J. Med. 358, 2849–2850 (2008).
Heeringa, S. F. et al. COQ6 mutations in human patients produce nephrotic syndrome with sensorineural deafness. J. Clin. Invest. 121, 2013–2024 (2011).
Ashraf, S. et al. ADCK4 mutations promote steroid-resistant nephrotic syndrome through CoQ10 biosynthesis disruption. J. Clin. Invest. 123, 5179–5189 (2013).
Gee, H. Y. et al. ARHGDIA mutations cause nephrotic syndrome via defective RHO GTPase signaling. J. Clin. Invest. 123, 3243–3253 (2013).
Gee, H. Y. et al. KANK deficiency leads to podocyte dysfunction and nephrotic syndrome. J. Clin. Invest. 125, 2375–2384 (2015).
Faul, C. et al. The actin cytoskeleton of kidney podocytes is a direct target of the antiproteinuric effect of cyclosporine A. Nat. Med. 14, 931–938 (2008).
Hinkes, B. et al. Positional cloning uncovers mutations in PLCE1 responsible for a nephrotic syndrome variant that may be reversible. Nat. Genet. 38, 1397–1405 (2006).
Moriniere, V. et al. Improving mutation screening in familial hematuric nephropathies through next generation sequencing. J. Am. Soc. Nephrol. 25, 2740–2751 (2014).
Harris, P. C. & Rossetti, S. Molecular diagnostics for autosomal dominant polycystic kidney disease. Nat. Rev. Nephrology 6, 197–206 (2010).
Pearle, M. S. et al. Medical management of kidney stones: AUA guideline. J. Urol. 192, 316–324 (2014).
Staiger, K. et al. Hypomagnesemia and nephrocalcinosis in a patient with two heterozygous mutations in the CLDN16 gene. J. Nephrol. 20, 107–110 (2007).
Otto, E. A. et al. Candidate exome capture identifies mutation of SDCCAG8 as the cause of a retinal-renal ciliopathy. Nat. Genet. 42, 840–850 (2010).
Giglio, S. et al. Heterogeneous genetic alterations in sporadic nephrotic syndrome associate with resistance to immunosuppression. J. Am. Soc. Nephrol. 26, 230–236 (2015).
Boyer, O. et al. Mutational analysis of the PLCE1 gene in steroid resistant nephrotic syndrome. J. Med. Genet. 47, 445–452 (2010).
Lipska, B. S. et al. Genetic screening in adolescents with steroid-resistant nephrotic syndrome. Kidney Int. 84, 206–213 (2013).
Ross, L. F. et al. Technical report: ethical and policy issues in genetic testing and screening of children. Genet. Med. 15, 234–245 (2013).
Mardis, E. R. The impact of next-generation sequencing technology on genetics. Trends Genet. 24, 133–141 (2008).
Koboldt, D. C., Steinberg, K. M., Larson, D. E., Wilson, R. K. & Mardis, E. R. The next-generation sequencing revolution and its impact on genomics. Cell 155, 27–38 (2013).
Hildebrandt, F. et al. A systematic approach to mapping recessive disease genes in individuals from outbred populations. PLoS Genet. 5, e1000353 (2009).
Green, R. C. et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet. Med. 5, 565–574 (2013).
Yang, Y. et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA 312, 1870–1879 (2014).
Lee, H. et al. Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA 312, 1880–1887 (2014).
Boute, N. et al. NPHS2, encoding the glomerular protein podocin, is mutated in autosomal recessive steroid-resistant nephrotic syndrome. Nat. Genet. 24, 349–354 (2000).
Lowik, M. M. et al. Focal segmental glomerulosclerosis in a patient homozygous for a CD2AP mutation. Kidney Int. 72, 1198–1203 (2007).
Kaplan, J. M. et al. Mutations in ACTN4, encoding α-actinin-4, cause familial focal segmental glomerulosclerosis. Nat. Genet. 24, 251–256 (2000).
Brown, E. J. et al. Mutations in the formin gene INF2 cause focal segmental glomerulosclerosis. Nat. Genet. 42, 72–76 (2010).
Zenker, M. et al. Human laminin β2 deficiency causes congenital nephrosis with mesangial sclerosis and distinct eye abnormalities. Hum. Mol. Genet. 13, 2625–2632 (2004).
Gee, H. Y. et al. Mutations in EMP2 cause childhood-onset nephrotic syndrome. Am. J. Hum. Genet. 94, 884–890 (2014).
Diomedi-Camassei, F. et al. COQ2 nephropathy: a newly described inherited mitochondriopathy with primary renal involvement. J. Am. Soc. Nephrol. 18, 2773–2780 (2007).
Ozaltin, F. Primary coenzyme Q10 (CoQ10) deficiencies and related nephropathies. Pediatr. Nephrol. 29, 961–969 (2014).
Salviati, L. et al. Infantile encephalomyopathy and nephropathy with CoQ10 deficiency: a CoQ10-responsive condition. Neurology 65, 606–608 (2005).
Rotig, A. et al. Quinone-responsive multiple respiratory-chain dysfunction due to widespread coenzyme Q10 deficiency. Lancet 356, 391–395 (2000).
Saisawat, P. et al. Identification of two novel CAKUT-causing genes by massively parallel exon resequencing of candidate genes in patients with unilateral renal agenesis. Kidney Int. 81, 196–200 (2012).
Bettencourt-Dias, M., Hildebrandt, F., Pellman, D., Woods, G. & Godinho, S. A. Centrosomes and cilia in human disease. Trends Genet. 27, 307–315 (2011).
Hildebrandt, F., Attanasio, M. & Otto, E. Nephronophthisis: disease mechanisms of a ciliopathy. J. Am. Soc. Nephrol. 20, 23–35 (2009).
Lemaire, M. et al. Recessive mutations in DGKE cause atypical hemolytic-uremic syndrome. Nat. Genet. 45, 531–536 (2013).
Neumann, H. P. et al. Haemolytic uraemic syndrome and mutations of the factor H gene: a registry-based study of German speaking countries. J. Med. Genet. 40, 676–681 (2003).
Noris, M. et al. Relative role of genetic complement abnormalities in sporadic and familial aHUS and their impact on clinical phenotype. Clin. J. Am. Soc. Nephrol. 5, 1844–1859 (2010).
Westra, D. et al. Atypical hemolytic uremic syndrome and genetic aberrations in the complement factor H-related 5 gene. J. Hum. Genet. 57, 459–464 (2012).
Goldfarb, D. S. The search for monogenic causes of kidney stones. J. Am. Soc. Nephrol. 26, 507–510 (2015).
Weber, S. et al. SIX2 and BMP4 mutations associate with anomalous kidney development. J. Am. Soc. Nephrol. 19, 891–903 (2008).
Brockschmidt, A. et al. CHD1L: a new candidate gene for congenital anomalies of the kidneys and urinary tract (CAKUT). Nephrol. Dial. Transplant. 27, 2355–2364 (2012).
Abdelhak, S. et al. A human homologue of the Drosophila eyes absent gene underlies branchio-oto-renal (BOR) syndrome and identifies a novel gene family. Nat. Genet. 15, 157–164 (1997).
Pandolfi, P. P. et al. Targeted disruption of the GATA3 gene causes severe abnormalities in the nervous system and in fetal liver haematopoiesis. Nat. Genet. 11, 40–44 (1995).
Van Esch, H. et al. GATA3 haplo-insufficiency causes human HDR syndrome. Nature 406, 419–422 (2000).
Lindner, T. H. et al. A novel syndrome of diabetes mellitus, renal dysfunction and genital malformation associated with a partial deletion of the pseudo-POU domain of hepatocyte nuclear factor-1β. Hum. Mol. Genet. 8, 2001–2008 (1999).
Kirby, A. et al. Mutations causing medullary cystic kidney disease type 1 lie in a large VNTR in MUC1 missed by massively parallel sequencing. Nat. Genet. 45, 299–303 (2013).
Sanyanusin, P., McNoe, L. A., Sullivan, M. J., Weaver, R. G. & Eccles, M. R. Mutation of PAX2 in two siblings with renal-coloboma syndrome. Hum. Mol. Genet. 4, 2183–2184 (1995).
Skinner, M. A. et al. Renal aplasia in humans is associated with RET mutations.. Am. J. Hum. Genet. 82, 344–351 (2008).
Lu, W. et al. Disruption of ROBO2 is associated with urinary tract anomalies and confers risk of vesicoureteral reflux. Am. J. Hum. Genet. 80, 616–632 (2007).
Kohlhase, J., Wischermann, A., Reichenbach, H., Froster, U. & Engel, W. Mutations in the SALL1 putative transcription factor gene cause Townes−Brocks syndrome. Nat. Genet. 18, 81–83 (1998).
Ruf, R. G. et al. SIX1 mutations cause branchio-oto-renal syndrome by disruption of EYA1−SIX1−DNA complexes. Proc. Natl Acad. Sci. USA 101, 8090–8095 (2004).
Hoskins, B. E. et al. Transcription factor SIX5 is mutated in patients with branchio-oto-renal syndrome. Am. J. Hum. Genet. 80, 800–804 (2007).
Gimelli, S. et al. Mutations in SOX17 are associated with congenital anomalies of the kidney and the urinary tract. Hum. Mut. 31, 1352–1359 (2010).
Hwang, D. Y. et al. Mutations of the SLIT2−ROBO2 pathway genes SLIT2 and SRGAP1 confer risk for congenital anomalies of the kidney and urinary tract. Hum. Genet. 134, 905–916 (2015).
Gbadegesin, R. A. et al. TNXB mutations can cause vesicoureteral reflux. J. Am. Soc. Nephrol. 24, 1313–1322 (2013).
Hart, T. C. et al. Mutations of the UMOD gene are responsible for medullary cystic kidney disease 2 and familial juvenile hyperuricaemic nephropathy. J. Med. Genet. 39, 882–892 (2002).
Jenkins, D. et al. De novo Uroplakin IIIa heterozygous mutations cause human renal adysplasia leading to severe kidney failure. J. Am. Soc. Nephrol. 16, 2141–2149 (2005).
Biason-Lauber, A., Konrad, D., Navratil, F. & Schoenle, E. J. A WNT4 mutation associated with Müllerian-duct regression and virilization in a 46,XX woman. N. Engl. J. Med. 351, 792–798 (2004).
Mandel, H. et al. SERKAL syndrome: an autosomal-recessive disorder caused by a loss-of-function mutation in WNT4. Am. J. Hum. Genet. 82, 39–47 (2008).
Vivante, A. et al. Renal hypodysplasia associates with a WNT4 variant that causes aberrant canonical Wnt signaling. J. Am. Soc. Nephrol. 24, 550–558 (2013).
Gribouval, O. et al. Mutations in genes in the renin–angiotensin system are associated with autosomal recessive renal tubular dysgenesis. Nat. Genet. 37, 964–968 (2005).
Weber, S. et al. Muscarinic acetylcholine receptor M3 mutation causes urinary bladder disease and a prune-belly-like syndrome. Am. J. Hum. Genet. 89, 668–674 (2011).
Barak, H. et al. FGF9 and FGF20 maintain the stemness of nephron progenitors in mice and man. Dev. Cell 22, 1191–1207 (2012).
McGregor, L. et al. Fraser syndrome and mouse blebbed phenotype caused by mutations in FRAS1/Fras1 encoding a putative extracellular matrix protein. Nat. Genet. 34, 203–208 (2003).
Bulum, B. et al. HPSE2 mutations in urofacial syndrome, non-neurogenic neurogenic bladder and lower urinary tract dysfunction. Nephron 130, 54–58 (2015).
Humbert, C. et al. Integrin alpha 8 recessive mutations are responsible for bilateral renal agenesis in humans. Am. J. Hum. Genet. 94, 288–294 (2014).
Stuart, H. M. et al. LRIG2 mutations cause urofacial syndrome. Am. J. Hum. Genet. 92, 259–264 (2013).
Hardelin, J. P. et al. X chromosome-linked Kallmann syndrome: stop mutations validate the candidate gene. Proc. Natl Acad. Sci. USA 89, 8190–8194 (1992).
Shih, N. Y. et al. Congenital nephrotic syndrome in mice lacking CD2-associated protein. Science 286, 312–315 (1999).
Sethi, S., Fervenza, F. C., Zhang, Y. & Smith, R. J. Secondary focal and segmental glomerulosclerosis associated with single-nucleotide polymorphisms in the genes encoding complement factor H and C3. Am. J. Kidney Dis. 60, 316–321 (2012).
Ovunc, B. et al. Exome sequencing reveals cubilin mutation as a single-gene cause of proteinuria. J. Am. Soc. Nephrol. 22, 1815–1820 (2011).
Ozaltin, F. et al. DGKE variants cause a glomerular microangiopathy that mimics membranoproliferative GN. J. Am. Soc. Nephrol. 24, 377–384 (2013).
Has, C. et al. Integrin α3 mutations with kidney, lung, and skin disease. N. Engl. J. Med. 366, 1508–1514 (2012).
Kambham, N. et al. Congenital focal segmental glomerulosclerosis associated with beta4 integrin mutation and epidermolysis bullosa. Am. J. Kidney Dis. 36, 190–196 (2000).
Yasukawa, T. et al. Modification defect at anticodon wobble nucleotide of mitochondrial tRNAs(Leu)(UUR) with pathogenic mutations of mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. J. Biol. Chem. 275, 4251–4257 (2000).
Mele, C. et al. MYO1E mutations and childhood familial focal segmental glomerulosclerosis. N. Engl. J. Med. 365, 295–306 (2011).
Miyake, N. et al. Biallelic mutations in nuclear pore complex subunit nup107 cause early-childhood-onset steroid-resistant nephrotic syndrome. Am. J. Hum. Genet. 97, 555–566 (2015).
Lopez, L. C. et al. Leigh syndrome with nephropathy and CoQ10 deficiency due to decaprenyl diphosphate synthase subunit 2 (PDSS2) mutations. Am. J. Hum. Genet. 79, 1125–1129 (2006).
Ozaltin, F. et al. Disruption of PTPRO causes childhood-onset nephrotic syndrome. Am. J. Hum. Genet. 89, 139–147 (2011).
Berkovic, S. F. et al. Array-based gene discovery with three unrelated subjects shows SCARB2/LIMP-2 deficiency causes myoclonus epilepsy and glomerulosclerosis. Am. J. Hum. Genet. 82, 673–684 (2008).
Boerkoel, C. F. et al. Mutant chromatin remodeling protein SMARCAL1 causes Schimke immuno-osseous dysplasia. Nat. Genet. 30, 215–220 (2002).
Colin, E. et al. Loss-of-function mutations in WDR73 are responsible for microcephaly and steroid-resistant nephrotic syndrome: Galloway-Mowat syndrome. Am. J. Hum. Genet. 95, 637–648 (2014).
Vodopiutz, J. et al. WDR73 mutations cause infantile neurodegeneration and variable glomerular kidney disease. Hum. Mut. 36, 1021–1028 (2015).
Jinks, R. N. et al. Recessive nephrocerebellar syndrome on the Galloway-Mowat syndrome spectrum is caused by homozygous protein-truncating mutations of WDR73. Brain 138, 2173–2190 (2015).
Gbadegesin, R. A. et al. Mutations in the gene that encodes the F-actin binding protein anillin cause FSGS. J. Am. Soc. Nephrol. 25, 1991–2002 (2014).
Akilesh, S. et al. Arhgap24 inactivates Rac1 in mouse podocytes, and a mutant form is associated with familial focal segmental glomerulosclerosis. J. Clin. Invest. 121, 4127–4137 (2011).
McIntosh I. et al. Mutation analysis of LMX1B gene in nail-patella syndrome patients. Am. J. Hum. Genet. 63, 1651–1658 (1998).
Heath, K. E. et al. Nonmuscle myosin heavy chain IIA mutations define a spectrum of autosomal dominant macrothrombocytopenias: May-Hegglin anomaly and Fechtner, Sebastian, Epstein, and Alport-like syndromes. Am. J. Hum. Genet. 69, 1033–1045 (2001).
Winn, M. P. et al. A mutation in the TRPC6 cation channel causes familial focal segmental glomerulosclerosis. Science 308, 1801–1804 (2005).
Reiser, J. et al. TRPC6 is a glomerular slit diaphragm-associated channel required for normal renal function. Nat. Genet. 37, 739–744 (2005).
Jeanpierre, C. et al. Identification of constitutional WT1 mutations, in patients with isolated diffuse mesangial sclerosis, and analysis of genotype/phenotype correlations by use of a computerized mutation database. Am. J. Hum. Genet. 62, 824–833 (1998).
Fremeaux-Bacchi, V. et al. Complement factor I: a susceptibility gene for atypical haemolytic uraemic syndrome. J. Med. Genet. 41, e84 (2004).
McRae, J. L. et al. Human factor H-related protein 5 (FHR-5). A new complement-associated protein. J. Biol. Chem. 276, 6747–6754 (2001).
Gale, D. P. et al. Identification of a mutation in complement factor H-related protein 5 in patients of Cypriot origin with glomerulonephritis. Lancet 376, 794–801 (2010).
Athanasiou, Y. et al. Familial C3 glomerulopathy associated with CFHR5 mutations: clinical characteristics of 91 patients in 16 pedigrees. Clin. J. Am. Soc. Nephrol.: CJASN 6, 1436–1446 (2011).
Castelletti, F. et al. Mutations in FN1 cause glomerulopathy with fibronectin deposits. Proc. Natl Acad. Sci. USA 105, 2538–2543 (2008).
Mochizuki, T. et al. Identification of mutations in the α3(IV) and α4(IV) collagen genes in autosomal recessive Alport syndrome. Nat. Genet. 8, 77–81 (1994).
Marcocci, E. et al. Autosomal dominant Alport syndrome: molecular analysis of the COL4A4 gene and clinical outcome. Nephrol. Dial. Transplant. 24, 1464–1471 (2009).
Noris, M. et al. Familial haemolytic uraemic syndrome and an MCP mutation. Lancet 362, 1542–1547 (2003).
Knebelmann, B. et al. Spectrum of mutations in the COL4A5 collagen gene in X-linked Alport syndrome. Am. J. Hum. Genet. 59, 1221–1232 (1996).
Anker, M. C. et al. Alport syndrome with diffuse leiomyomatosis. Am. J. Med. Genet. 119A, 381–385 (2003).
Hildebrandt, F. et al. A novel gene encoding an SH3 domain protein is mutated in nephronophthisis type 1. Nat. Genet. 17, 149–153 (1997).
Saunier, S. et al. A novel gene that encodes a protein with a putative src homology 3 domain is a candidate gene for familial juvenile nephronophthisis. Hum. Mol. Genet. 6, 2317–2323 (1997).
Otto, E. A. et al. Mutations in INVS encoding inversin cause nephronophthisis type 2, linking renal cystic disease to the function of primary cilia and left–right axis determination. Nat. Genet. 34, 413–420 (2003).
Olbrich, H. et al. Mutations in a novel gene, NPHP3, cause adolescent nephronophthisis, tapeto-retinal degeneration and hepatic fibrosis. Nat. Genet. 34, 455–459 (2003).
Otto, E. et al. A gene mutated in nephronophthisis and retinitis pigmentosa encodes a novel protein, nephroretinin, conserved in evolution. Am. J. Hum. Genet. 71, 1167–1171 (2002).
Delous, M. et al. Nephrocystin-1 and nephrocystin-4 are required for epithelial morphogenesis and associate with PALS1/PATJ and Par6. Hum. Mol. Genet. 18, 4711–4723 (2009).
Otto, E. A. et al. Nephrocystin-5, a ciliary IQ domain protein, is mutated in Senior–Løken syndrome and interacts with RPGR and calmodulin. Nat. Genet. 37, 282–288 (2005).
Sayer, J. A. et al. The centrosomal protein nephrocystin-6 is mutated in Joubert syndrome and activates transcription factor ATF4. Nat. Genet. 38, 674–681 (2006).
Attanasio, M. et al. Loss of GLIS2 causes nephronophthisis in humans and mice by increased apoptosis and fibrosis. Nat. Genet. 39, 1018–1024 (2007).
Delous, M. et al. The ciliary gene RPGRIP1L is mutated in cerebello-oculo-renal syndrome (Joubert syndrome type B) and Meckel syndrome. Nat. Genet. 39, 875–881 (2007).
Otto, E. A. et al. NEK8 mutations affect ciliary and centrosomal localization and may cause nephronophthisis. J. Am. Soc. Nephrol. 19, 587–592 (2008).
Otto, E. A. et al. Hypomorphic mutations in meckelin (MKS3/TMEM67) cause nephronophthisis with liver fibrosis (NPHP11). J. Med. Genet. 46, 663–670 (2009).
Davis, E. E. et al. TTC21B contributes both causal and modifying alleles across the ciliopathy spectrum. Nat. Genet. 43, 189–196 (2011).
Coussa, R. G. et al. WDR19: an ancient, retrograde, intraflagellar ciliary protein is mutated in autosomal recessive retinitis pigmentosa and in Senior–Løken syndrome. Clin. Genet. 84, 150–159 (2013).
Chaki, M. et al. Exome capture reveals ZNF423 and CEP164 mutations, linking renal ciliopathies to DNA damage response signaling. Cell 150, 533–548 (2012).
Hoff, S. et al. ANKS6 is a central component of a nephronophthisis module linking NEK8 to INVS and NPHP3. Nat. Genet. 45, 951–956 (2013).
Reed, B. Y. et al. Identification and characterization of a gene with base substitutions associated with the absorptive hypercalciuria phenotype and low spinal bone density. J. Clin. Endocrinol. Metab. 87, 1476–1485 (2002).
Purdue, P. E., Allsop, J., Isaya, G., Rosenberg, L. E. & Danpure, C. J. Mistargeting of peroxisomal l-alanine:glyoxylate aminotransferase to mitochondria in primary hyperoxaluria patients depends upon activation of a cryptic mitochondrial targeting sequence by a point mutation. Proc. Natl Acad. Sci. USA 88, 10900–10904 (1991).
Hidaka, Y., Palella, T. D., O'Toole, T. E., Tarle, S. A. & Kelley, W. N. Human adenine phosphoribosyltransferase. Identification of allelic mutations at the nucleotide level as a cause of complete deficiency of the enzyme. J. Clin. Invest. 80, 1409–1415 (1987).
Stover, E. H. et al. Novel ATP6V1B1 and ATP6V0A4 mutations in autosomal recessive distal renal tubular acidosis with new evidence for hearing loss. J. Med. Genet. 39, 796–803 (2002).
Karet, F. E. et al. Mutations in the gene encoding B1 subunit of H+-ATPase cause renal tubular acidosis with sensorineural deafness. Nat. Genet. 21, 84–90 (1999).
Venta, P. J., Welty, R. J., Johnson, T. M., Sly, W. S. & Tashian, R. E. Carbonic anhydrase II deficiency syndrome in a Belgian family is caused by a point mutation at an invariant histidine residue (107 His→Tyr): complete structure of the normal human CA II gene. Am. J. Hum. Genet. 49, 1082–1090 (1991).
Pearce, S. H. et al. A familial syndrome of hypocalcemia with hypercalciuria due to mutations in the calcium-sensing receptor. N. Engl. J. Med. 335, 1115–1122 (1996).
Lloyd, S. E. et al. A common molecular basis for three inherited kidney stone diseases. Nature 379, 445–449 (1996).
Simon, D. B. et al. Mutations in the chloride channel gene, CLCNKB, cause Bartter's syndrome type III. Nat. Genet. 17, 171–178 (1997).
Simon, D. B. et al. Paracellin-1, a renal tight junction protein required for paracellular Mg2+ resorption. Science 285, 103–106 (1999).
Konrad, M. et al. Mutations in the tight-junction gene claudin 19 (CLDN19) are associated with renal magnesium wasting, renal failure, and severe ocular involvement. Am. J. Hum. Genet. 79, 949–957 (2006).
Schlingmann, K. P. et al. Mutations in CYP24A1 and idiopathic infantile hypercalcemia. N. Engl. J. Med. 365, 410–421 (2011).
Jaureguiberry, G. et al. Nephrocalcinosis (enamel renal syndrome) caused by autosomal recessive FAM20A mutations. Nephron Physiol. 122, 1–6 (2012).
Cramer, S. D., Ferree, P. M., Lin, K., Milliner, D. S. & Holmes, R. P. The gene encoding hydroxypyruvate reductase (GRHPR) is mutated in patients with primary hyperoxaluria type II. Hum. Mol. Genet. 8, 2063–2069 (1999).
Hamilton, A. J. et al. The HNF4A R76W mutation causes atypical dominant Fanconi syndrome in addition to a β cell phenotype. J. Med. Genet. 51, 165–169 (2014).
Belostotsky, R. et al. Mutations in DHDPSL are responsible for primary hyperoxaluria type III. Am. J. Hum. Genet. 87, 392–399 (2010).
Davidson, B. L. et al. Identification of 17 independent mutations responsible for human hypoxanthine-guanine phosphoribosyltransferase (HPRT) deficiency. Am. J. Hum. Genet. 48, 951–958 (1991).
Simon, D. B. et al. Genetic heterogeneity of Bartter's syndrome revealed by mutations in the K+ channel, ROMK. Nat. Genet. 14, 152–156 (1996).
Reilly, D. S., Lewis, R. A., Ledbetter, D. H. & Nussbaum, R. L. Tightly linked flanking markers for the Lowe oculocerebrorenal syndrome, with application to carrier assessment. Am. J. Hum. Genet. 42, 748–755 (1988).
Simon, D. B. et al. Bartter's syndrome, hypokalaemic alkalosis with hypercalciuria, is caused by mutations in the Na–K–2Cl cotransporter NKCC2. Nat. Genet. 13, 183–188 (1996).
Enomoto, A. et al. Molecular identification of a renal urate anion exchanger that regulates blood urate levels. Nature 417, 447–452 (2002).
Matsuo, H. et al. Mutations in glucose transporter 9 gene SLC2A9 cause renal hypouricemia. Am. J. Hum. Genet. 83, 744–751 (2008).
Prie, D. et al. Nephrolithiasis and osteoporosis associated with hypophosphatemia caused by mutations in the type 2a sodium-phosphate cotransporter. N. Engl. J. Med. 347, 983–991 (2002).
Lorenz-Depiereux, B. et al. Hereditary hypophosphatemic rickets with hypercalciuria is caused by mutations in the sodium-phosphate cotransporter gene SLC34A3. Am. J. Hum. Genet. 78, 193–201 (2006).
Calonge, M. J. et al. Cystinuria caused by mutations in rBAT, a gene involved in the transport of cystine. Nat. Genet. 6, 420–425 (1994).
Bruce, L. J. et al. Familial distal renal tubular acidosis is associated with mutations in the red cell anion exchanger (Band 3, AE1) gene. J. Clin. Invest. 100, 1693–1707 (1997).
Feliubadalo, L. et al. Non-type I cystinuria caused by mutations in SLC7A9, encoding a subunit (bo,+AT) of rBAT. Nat. Genet. 23, 52–57 (1999).
Karim, Z. et al. NHERF1 mutations and responsiveness of renal parathyroid hormone. N. Engl. J. Med. 359, 1128–1135 (2008).
Scott, P. et al. Suggestive evidence for a susceptibility gene near the vitamin D receptor locus in idiopathic calcium stone formation. J. Am. Soc. Nephrol. 10, 1007–1013 (1999).
Ichida, K. et al. Identification of two mutations in human xanthine dehydrogenase gene responsible for classical type I xanthinuria. J. Clin. Invest. 99, 2391–2397 (1997).
Wainwright, C. E. et al. Lumacaftor-ivacaftor in patients with cystic fibrosis homozygous for Phe508del CFTR. N. Engl. J. Med. 373, 220–231 (2015).
Acknowledgements
The authors' work described in this Review was supported by grants from the NIH (R01-DK088767 to F.H.) and by the March of Dimes Foundation (6FY11-241). A.V. is a recipient of the Fulbright postdoctoral scholar award for 2013 and is also supported by grants from the Manton Center Fellowship Program, Boston Children's Hospital, Boston, Massachusetts, USA, and the Mallinckrodt Research Fellowship Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
F.H. receives royalties for a mutation analysis panel that is licensed to Claritas Genomics. A.V. declares no competing interests.
Glossary
- Penetrance
-
The proportion of individuals who express a certain phenotype in relation to the number of individuals that carry the pathogenic variant(s). Incomplete penetrance refers to the observation that some individuals with the mutation do not develop the disease phenotype.
- Next-generation sequencing
-
A DNA sequencing method that enables simultaneous sequencing of multiple DNA segments in a high-throughput manner. Also known as massively parallel sequencing.
- Whole exome sequencing
-
Targeted capture and sequencing of the exome using next-generation sequencing. This method offers a powerful approach towards the identification of monogenic disease-causing genes.
- Variant
-
A difference in a DNA sequence compared to a 'normal' reference sequence. A variant can be benign (for example, a single nucleotide polymorphism) or disease-causing (for example, a mutation).
- Allele
-
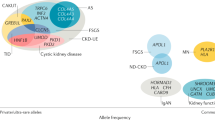
Specific DNA sequence variant in a given gene. Alleles can be designated according to their frequency as common or rare alleles.
- Phenotype
-
The observable characteristics of an individual as a morphological, clinical or biochemical trait. A phenotype can also be the presence or absence of a disease.
- Genotype
-
The set of alleles (variants of genes) that structure an individual's genetic makeup.
- Expressivity
-
Variation of the expression of the phenotype among affected individuals with the same genotype. Variable expressivity refers to different degrees of severity and/or organ involvement in different affected individuals that carry an identical mutation.
- Exon
-
The protein coding part of a gene. Exons are spliced together following gene transcription to form mRNA, which is translated into protein.
- Exome
-
The protein coding sequences of the entire genome (comprising ∼1% of the human genome).
- Variant filtering
-
The process of excluding variants as disease-causing. For instance, very common variants and variants that do not alter the protein sequence are excluded.
- Homozygosity mapping
-
A technique in which the homozygous regions across the genome are identified. This strategy is effective for the discovery of autosomal recessive monogenic disease genes in consanguineous families.
- Phenocopies
-
A variation in a phenotype of given trait that can mimick a different trait.
Rights and permissions
About this article
Cite this article
Vivante, A., Hildebrandt, F. Exploring the genetic basis of early-onset chronic kidney disease. Nat Rev Nephrol 12, 133–146 (2016). https://doi.org/10.1038/nrneph.2015.205
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrneph.2015.205
This article is cited by
-
A step-by-step, multidisciplinary strategy to maximize the yield of genetic testing in pediatric patients with chronic kidney diseases
Pediatric Nephrology (2024)
-
Diagnostic accuracy of shear wave elastography in evaluating renal fibrosis in children with chronic kidney disease: a comparative study with nuclear scan
Egyptian Journal of Radiology and Nuclear Medicine (2023)
-
Hidden genetics behind glomerular scars: an opportunity to understand the heterogeneity of focal segmental glomerulosclerosis?
Pediatric Nephrology (2023)
-
Genetische Diagnostik bei Nierenerkrankungen im Erwachsenenalter
Die Nephrologie (2023)
-
Kidney biopsy diagnosis in childhood in the Norwegian Kidney Biopsy Registry and the long-term risk of kidney replacement therapy: a 25-year follow-up
Pediatric Nephrology (2023)