Key Points
-
There are marked differences in the in vitro differentiation capacity of pluripotent stem cell lines, including embryonic stem (ES) cells and induced pluripotent stem (iPS) cells. These differences currently limit their possible application in the clinic and as a research tool.
-
The molecular underpinnings of these functional differences are unclear. In some cases, the differences can be ameliorated by altering culture conditions by adding or removing specific growth factors and/or signalling molecules.
-
Pluripotent stem cells derived using the same method have highly similar gene expression profiles, yet can exhibit functional differences. This suggests that genetic or epigenetic changes are silent in the pluripotent state.
-
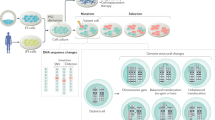
Most genetic differences between founder and reprogrammed cells seem to pre-exist in a minor population of the founder cells.
-
Epigenetic differences between ES cells and iPS cells are both residual (that is, they maintain the same epigenetic state as their respective founder cell type) and aberrant (that is, resembling neither ES cells nor founder cells), and both types of differences may have an impact on the in vitro differentiation process.
-
It remains unclear whether gene expression heterogeneity within a clonal pluripotent stem cell line is a crucial feature of pluripotency, and whether this gene expression heterogeneity is associated with functional differences between different pluripotent cell lines.
Abstract
Pluripotent stem cells constitute a platform to model disease and developmental processes and can potentially be used in regenerative medicine. However, not all pluripotent cell lines are equal in their capacity to differentiate into desired cell types in vitro. Genetic and epigenetic variations contribute to functional variability between cell lines and heterogeneity within clones. These genetic and epigenetic variations could 'lock' the pluripotency network resulting in residual pluripotent cells or alter the signalling response of developmental pathways leading to lineage bias. The molecular contributors to functional variability and heterogeneity in both embryonic stem (ES) cells and induced pluripotent stem (iPS) cells are only beginning to emerge, yet they are crucial to the future of the stem cell field.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Kahan, B. W. & Ephrussi, B. Developmental potentialities of clonal in vitro cultures of mouse testicular teratoma. J. Natl Cancer Inst. 44, 1015–1036 (1970).
Evans, M. J. The isolation and properties of a clonal tissue culture strain of pluripotent mouse teratoma cells. Development 28, 163–176 (1972).
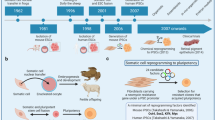
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006). A landmark paper that demonstrates transcription factor-based reprogramming to pluripotent-like cells. It initiated a race to reprogram human fibroblasts to pluripotency, leading to widespread accessibility to pluripotent stem cells.
Young, R. A. Control of the embryonic stem cell state. Cell 144, 940–954 (2011).
Nichols, J. & Smith, A. Pluripotency in the embryo and in culture. Cold Spring Harb. Persp.Biol. 4, a008128 (2012).
Solter, D. From teratocarcinomas to embryonic stem cells and beyond: a history of embryonic stem cell research. Nature Rev. Genet. 7, 319–327 (2006).
Wu, S. M. & Hochedlinger, K. Harnessing the potential of induced pluripotent stem cells for regenerative medicine. Nature Cell Biol. 13, 497–505 (2011).
Cohen, D. E. & Melton, D. Turning straw into gold: directing cell fate for regenerative medicine. Nature Rev. Genet. 12, 243–252 (2011).
Zhu, H., Lensch, M. W., Cahan, P. & Daley, G. Q. Investigating monogenic and complex diseases with pluripotent stem cells. Nature Rev. Genet. 12, 266–275 (2011).
Bellin, M., Marchetto, M. C., Gage, F. H. & Mummery, C. L. Induced pluripotent stem cells: the new patient? Nature Rev. Mol. Cell Biol. 13, 713–726 (2012).
Cherry, A. B. C. & Daley, G. Q. Reprogramming cellular identity for regenerative medicine. Cell 148, 1110–1122 (2012).
Robinton, D. A. & Daley, G. Q. The promise of induced pluripotent stem cells in research and therapy. Nature 481, 295–305 (2012).
Keller, G. Embryonic stem cell differentiation: emergence of a new era in biology and medicine. Genes Dev. 19, 1129–1155 (2005).
Niwa, H. Mouse ES cell culture system as a model of development. Dev. Growth Differ. 52, 275–283 (2010).
Grskovic, M., Javaherian, A., Strulovici, B. & Daley, G. Q. Induced pluripotent stem cells — opportunities for disease modelling and drug discovery. Nature Rev. Drug Discov. 10, 915–929 (2011).
Banito, A. & Gil, J. Induced pluripotent stem cells and senescence: learning the biology to improve the technology. EMBO Rep. 11, 353–359 (2010).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Loh, Y.-H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nature Genet. 38, 431–440 (2006).
Thomson, M. et al. Pluripotency factors in embryonic stem cells regulate differentiation into germ layers. Cell 145, 875–889 (2011).
Teo, A. K. K. et al. Pluripotency factors regulate definitive endoderm specification through eomesodermin. Genes Dev. 25, 238–250 (2011).
Yuan, H., Corbi, N., Basilico, C. & Dailey, L. Developmental-specific activity of the FGF-4 enhancer requires the synergistic action of Sox2 and Oct-3. Genes Dev. 9, 2635–2645 (1995).
Marks, H. et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 149, 590–604 (2012).
Kunath, T. et al. FGF stimulation of the Erk1/2 signalling cascade triggers transition of pluripotent embryonic stem cells from self-renewal to lineage commitment. Development 134, 2895–2902 (2007).
Niwa, H., Burdon, T., Chambers, I. & Smith, A. Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev. 12, 2048–2060 (1998).
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003).
Kim, J., Chu, J., Shen, X., Wang, J. & Orkin, S. H. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008).
Lowell, S., Benchoua, A., Heavey, B. & Smith, A. G. Notch promotes neural lineage entry by pluripotent embryonic stem cells. PLoS Biol. 4, e121 (2006).
Lindsley, R. C., Gill, J. G., Kyba, M., Murphy, T. L. & Murphy, K. M. Canonical Wnt signaling is required for development of embryonic stem cell-derived mesoderm. Development 133, 3787–3796 (2006).
Ogawa, K., Nishinakamura, R., Iwamatsu, Y., Shimosato, D. & Niwa, H. Synergistic action of Wnt and LIF in maintaining pluripotency of mouse ES cells. Biochem. Biophys. Res. Commun. 343, 159–166 (2006).
Berge, ten, D. et al. Embryonic stem cells require Wnt proteins to prevent differentiation to epiblast stem cells. Nature Cell Biol. 13, 1070–1075 (2011).
Yi, F. et al. Opposing effects of Tcf3 and Tcf1 control Wnt stimulation of embryonic stem cell self-renewal. Nature Cell Biol. 13, 762–770 (2011).
Wray, J. et al. Inhibition of glycogen synthase kinase-3 alleviates Tcf3 repression of the pluripotency network and increases embryonic stem cell resistance to differentiation. Nature Cell Biol. 13, 838–845 (2011).
Marson, A. et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134, 521–533 (2008).
Sinkkonen, L. et al. MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nature Struct. Mol. Biol. 15, 259–267 (2008).
Delaloy, C. et al. microRNA-9 coordinates proliferation and migration of human embryonic stem cell-derived neural progenitors. Cell Stem Cell 6, 323–335 (2010).
Wang, Y., Medvid, R., Melton, C., Jaenisch, R. & Blelloch, R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nature Genet. 39, 380–385 (2007).
Kanellopoulou, C. et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev. 19, 489–501 (2005).
Kaji, K. et al. The NuRD component Mbd3 is required for pluripotency of embryonic stem cells. Nature Cell Biol. 8, 285–292 (2006).
Zhu, D., Fang, J., Li, Y. & Zhang, J. Mbd3, a component of NuRD/Mi-2 complex, helps maintain pluripotency of mouse embryonic stem cells by repressing trophectoderm differentiation. PLoS ONE 4, e7684 (2009).
Berstine, E. G., Hooper, M. L., Grandchamp, S. & Ephrussi, B. Alkaline phosphatase activity in mouse teratoma. Proc. Natl Acad. Sci. 70, 3899–3903 (1973).
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Brons, I. G. M. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
De Los Angeles, A., Loh, Y.-H., Tesar, P. J. & Daley, G. Q. Accessing naïve human pluripotency. Curr. Opin. Genet. Dev. 22, 272–282 (2012).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009).
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Osafune, K. et al. Marked differences in differentiation propensity among human embryonic stem cell lines. Nature Biotechnol. 26, 313–315 (2008). The first paper to systematically compare the in vitro differentiation capacity of human ES cells. It demonstrates that despite the equivalence of human ES cells by teratoma formation, human ES cells can still exhibit great tendencies to differentiate into specific lineages in vitro.
Di Giorgio, F. P., Boulting, G. L., Bobrowicz, S. & Eggan, K. C. Human embryonic stem cell-derived motor neurons are sensitive to the toxic effect of glial cells carrying an ALS-causing mutation. Cell Stem Cell 3, 637–648 (2008).
Hu, B.-Y. et al. Neural differentiation of human induced pluripotent stem cells follows developmental principles but with variable potency. Proc. Natl Acad. Sci. USA 107, 4335–4340 (2010).
Boulting, G. L. et al. A functionally characterized test set of human induced pluripotent stem cells. Nature Biotechnol. 29, 279–286 (2011).
Grigoriadis, A. E. et al. Directed differentiation of hematopoietic precursors and functional osteoclasts from human ES and iPS cells. Blood 115, 2769–2776 (2010).
Chang, K.-H. et al. Diverse hematopoietic potentials of five human embryonic stem cell lines. Exp. Cell Res. 314, 2930–2940 (2008).
Kelly, D. L. & Rizzino, A. DNA microarray analyses of genes regulated during the differentiation of embryonic stem cells. Mol. Reprod. Dev. 56, 113–123 (2000).
Loring, J. F., Porter, J. G., Seilhammer, J., Kaser, M. R. & Wesselschmidt, R. A gene expression profile of embryonic stem cells and embryonic stem cell-derived neurons. Restor. Neurol. Neurosci. 18, 81–88 (2001).
Ramalho-Santos, M. 'Stemness': transcriptional profiling of embryonic and adult stem cells. Science 298, 597–600 (2002).
Ivanova, N. B. et al. A stem cell molecular signature. Science 298, 601–604 (2002).
Fortunel, N. O. et al. Comment on “'Stemness': transcriptional profiling of embryonic and adult stem cells” and “a stem cell molecular signature”. Science 302, 393 (2003).
D'Amour, K. A. & Gage, F. H. Genetic and functional differences between multipotent neural and pluripotent embryonic stem cells. Proc. Natl Acad. Sci. 100, 11866–11872 (2003).
Tanaka, T. S. Gene expression profiling of embryo-derived stem cells reveals candidate genes associated with pluripotency and lineage specificity. Genome Res. 12, 1921–1928 (2002).
Sato, N. et al. Molecular signature of human embryonic stem cells and its comparison with the mouse. Dev. Biol. 260, 404–413 (2003).
Richards, M., Tan, S. P., Tan, J. H., Chan, W. K. & Bongso, A. The transcriptome profile of human embryonic stem cells as defined by SAGE. Stem Cells 22, 51–64 (2004).
Sperger, J. M. et al. Gene expression patterns in human embryonic stem cells and human pluripotent germ cell tumors. Proc. Natl Acad. Sci. 100, 13350–13355 (2003).
Zeng, X. et al. Properties of pluripotent human embryonic stem cells BG01 and BG02. Stem Cells 22, 292–312 (2004).
Bhattacharya, B. Gene expression in human embryonic stem cell lines: unique molecular signature. Blood 103, 2956–2964 (2004).
Abeyta, M. J. Unique gene expression signatures of independently-derived human embryonic stem cell lines. Hum. Mol. Genet. 13, 601–608 (2004).
Okita, K., Ichisaka, T. & Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 448, 313–317 (2007).
Wernig, M. et al. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 448, 318–324 (2007).
Maherali, N. et al. Directly reprogrammed fibroblasts show global epigenetic remodeling and widespread tissue contribution. Cell Stem Cell 1, 55–70 (2007).
Park, I.-H. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451, 141–146 (2008).
Chin, M. H. et al. Induced pluripotent stem cells and embryonic stem cells are distinguished by gene expression signatures. Cell Stem Cell 5, 111–123 (2009).
Marchetto, M. C. N. et al. Transcriptional signature and memory retention of human-induced pluripotent stem cells. PLoS ONE 4, e7076 (2009).
Ghosh, Z. et al. Persistent donor cell gene expression among human induced pluripotent stem cells contributes to differences with human embryonic stem cells. PLoS ONE 5, e8975 (2010).
Guenther, M. G. et al. Chromatin structure and gene expression programs of human embryonic and induced pluripotent stem cells. Cell Stem Cell 7, 249–257 (2010). A careful comparison of the expression profiles of human ES cells and human iPS cells, arguing that no set of genes consistently distinguishes these two cell types.
Newman, A. M. & Cooper, J. B. Lab-specific gene expression signatures in pluripotent stem cells. Cell Stem Cell 7, 258–262 (2010).
Ohi, Y. et al. Incomplete DNA methylation underlies a transcriptional memory of somatic cells in human iPS cells. Nature Cell Biol. 13, 541–549 (2011).
Bock, C. et al. Reference maps of human ES and iPS cell variation enable high-throughput characterization of pluripotent cell lines. Cell 144, 439–452 (2011).
Phanstiel, D. H. et al. Proteomic and phosphoproteomic comparison of human ES and iPS cells. Nature Methods 8, 821–827 (2011).
Ruiz, S. et al. Identification of a specific reprogramming-associated epigenetic signature in human induced pluripotent stem cells. Proc. Natl Acad. Sci. USA 109, 16196–16201 (2012).
Kim, H. et al. miR-371-3 expression predicts neural differentiation propensity in human pluripotent stem cells. Cell Stem Cell 8, 695–706 (2011).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Niwa, H., Ogawa, K., Shimosato, D. & Adachi, K. A parallel circuit of LIF signalling pathways maintains pluripotency of mouse ES cells. Nature 460, 118–122 (2009).
Toyooka, Y., Shimosato, D., Murakami, K., Takahashi, K. & Niwa, H. Identification and characterization of subpopulations in undifferentiated ES cell culture. Development 135, 909–918 (2008).
Narsinh, K. H. et al. Single cell transcriptional profiling reveals heterogeneity of human induced pluripotent stem cells. J. Clin. Invest. 121, 1217 (2011).
Kalmar, T. et al. Regulated fluctuations in Nanog expression mediate cell fate decisions in embryonic stem cells. PLoS Biol. 7, e1000149 (2009).
MacArthur, B. D. et al. Nanog-dependent feedback loops regulate murine embryonic stem cell heterogeneity. Nature Cell Biol. 14, 1139–1147 (2012).
Yamaji, M. et al. PRDM14 ensures naive pluripotency through dual regulation of signaling and epigenetic pathways in mouse embryonic stem cells. Cell Stem Cell 12, 368–382 (2013).
Gore, A. et al. Somatic coding mutations in human induced pluripotent stem cells. Nature 470, 63–67 (2011).
Gaspar-Maia, A., Alajem, A., Meshorer, E. & Ramalho-Santos, M. Open chromatin in pluripotency and reprogramming. Nature Rev. Mol. Cell Biol. 12, 36–47 (2011).
Kim, K. et al. Epigenetic memory in induced pluripotent stem cells. Nature 467, 285–290 (2010). One of the first and most comprehensive studies to investigate the extent to which DNA methylation patterns of diverse founder cell types are detectable in reprogrammed mouse iPS cells.
Kim, K. et al. Donor cell type can influence the epigenome and differentiation potential of human induced pluripotent stem cells. Nature Biotechnol. 29, 1117–1119 (2011).
Polo, J. M. et al. Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nature Biotechnol. 28, 848–855 (2010).
Bar-Nur, O., Russ, H. A., Efrat, S. & Benvenisty, N. Epigenetic memory and preferential lineage-specific differentiation in induced pluripotent stem cells derived from human pancreatic islet β-cells. Cell Stem Cell 9, 17–23 (2011).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011).
Stadtfeld, M. et al. Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature 465, 175–181 (2010).
Carey, B. W. et al. Reprogramming factor stoichiometry influences the epigenetic state and biological properties of induced pluripotent stem cells. Cell Stem Cell 9, 588–598 (2011).
Kajiwara, M. et al. Donor-dependent variations in hepatic differentiation from human-induced pluripotent stem cells. Proc. Natl Acad. Sci. USA 109, 12538–12543 (2012).
Laurent, L. C. et al. Dynamic changes in the copy number of pluripotency and cell proliferation genes in human ESCs and iPSCs during reprogramming and time in culture. Cell Stem Cell 8, 106–118 (2011).
Hussein, S. M. et al. Copy number variation and selection during reprogramming to pluripotency. Nature 471, 58–62 (2011).
Martins-Taylor, K. et al. Recurrent copy number variations in human induced pluripotent stem cells. Nature Biotechnol. 29, 488–491 (2011).
Cheng, L. et al. Low incidence of DNA sequence variation in human induced pluripotent stem cells generated by nonintegrating plasmid expression. Cell Stem Cell 10, 337–344 (2012).
Ji, J. et al. Elevated coding mutation rate during the reprogramming of human somatic cells into induced pluripotent stem cells. Stem Cells 30, 435–440 (2012).
Young, M. A. et al. Background mutations in parental cells account for most of the genetic heterogeneity of induced pluripotent stem cells. Cell Stem Cell 10, 570–582 (2012).
Abyzov, A. et al. Somatic copy number mosaicism in human skin revealed by induced pluripotent stem cells. Nature 492, 438–442 (2012). A sequence-based mapping of DNA copy number differences between founder cells and reprogrammed human iPS cells. Rather than being induced by reprogramming, most CNVs are already present as rare alleles in the founder population.
Ruiz, S. et al. Analysis of protein-coding mutations in hiPSCs and their possible role during somatic cell reprogramming. Nature Commun. 4, 1382 (2013).
International Stem Cell Initiative. Screening ethnically diverse human embryonic stem cells identifies a chromosome 20 minimal amplicon conferring growth advantage. Nature Biotechnol. 29, 1132–1144 (2011).
Stevens, L. C. The development of transplantable teratocarcinomas from intratesticular grafts of pre- and postimplantation mouse embryos. Dev. Biol. 21, 364–382 (1970).
Rosenthal, M. D., Wishnow, R. M. & Sato, G. H. In vitro growth and differetiation of clonal populations of multipotential mouse clls derived from a transplantable testicular teratocarcinoma. J. Natl Cancer Inst. 44, 1001–1014 (1970).
Mintz, B. & Illmensee, K. Normal genetically mosaic mice produced from malignant teratocarcinoma cells. Proc. Natl Acad. Sci. 72, 3585–3589 (1975).
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl Acad. Sci. 78, 7634–7638 (1981).
Thomson, J. A. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Park, I.-H. et al. Disease-specific induced pluripotent stem cells. Cell 134, 877–886 (2008).
Sneddon, J. B., Borowiak, M. & Melton, D. A. Self-renewal of embryonic-stem-cell-derived progenitors by organ-matched mesenchyme. Nature 491, 765–768 (2012).
Suda, Y., Suzuki, M., Ikawa, Y. & Aizawa, S. Mouse embryonic stem cells exhibit indefinite proliferative potential. J. Cell. Physiol. 133, 197–201 (1987).
Nagy, A. et al. Embryonic stem cells alone are able to support fetal development in the mouse. Development 110, 815–821 (1990).
Acknowledgements
G.Q.D. is supported by grants from the US National Institutes of Health (NIH) (UO1-HL100001 Progenitor cell biology consortium, R24-DK092760, P50HG005550 and special funds from the ARRA stimulus package- RC2-HL102815, RC4DK 090913), the Roche Foundation for Anemia Research, Alex's Lemonade Stand Foundation, Doris Duke Charitable Foundation and the Ellison Medical Foundation. G.Q.D. is an affiliate member of the Broad Institute and an investigator of the Manton Center for Orphan Disease Research and the Howard Hughes Medical Institute. P.C. is supported by grants T32HL007623 and 2T32HL66987-11 from the National Heart, Lung, and Blood Institute (NHLBI). The authors would like to thank R. Zhao and A. De Los Angeles for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
G.Q.D. is a member of the scientific advisory boards of the following companies: Johnson & Johnson, Verastem, Epizyme, Inc., iPierian, Inc., Solasia K.K. and MPM Capital, LLP. P.C. declares no competing financial interests.
Related links
Glossary
- Teratomas
-
Localized tumours in which germ layer derivatives are apparent, often in highly organized patterns resembling normal tissues.
- Trophectoderm
-
The first cells to differentiate from the fertilized egg and that form the outer layer of the blastocyst. They give rise to extra-embryonic tissues, including the placenta.
- Unsupervised analysis
-
Unsupervised analysis methods, such as clustering, discover groups in data without being informed of group labels.This is in contrast to supervised analysis methods, such as the analysis of variance (ANOVA), which find features that distinguish groups.
- Seminomas
-
Malignant but highly treatable germ cell tumours of the testis.
- Episome method
-
A method to force the ectopic expression of reprogramming factors. This method relies on transient transfection of plasmids that encode reprogramming factors.
- Embryoid bodies
-
Three-dimensional, semi-organized aggregates of differentiating pluripotent stem cells.
- CpG bisulphite sequencing
-
Identification of CpG methylation by targeted sequencing of bisulphite-treated genomic DNA. Bisulphite treatment causes the conversion of cytosine to uracil. Because uracil is not a genomic residue, it can be used to infer cytosine methylation.
- Dlk1–Dio3 locus
-
An imprinted region of mouse chromosome 12qF1 containing multiple dosage-sensitive, protein-coding and non-coding genes.
- Genetic mosaicism
-
Genetic heterogeneity of a cellular population that has arisen due to the presence of distinct founder clones. Can occur at the tissue, organ or organism level, or in the dish.
- Exome
-
The nucleotide sequence of the protein-coding portion of a genome as determined by targeted next-generation sequencing.
Rights and permissions
About this article
Cite this article
Cahan, P., Daley, G. Origins and implications of pluripotent stem cell variability and heterogeneity. Nat Rev Mol Cell Biol 14, 357–368 (2013). https://doi.org/10.1038/nrm3584
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3584
This article is cited by
-
Potential Role of Nrf2, HER2, and ALDH in Cancer Stem Cells: A Narrative Review
The Journal of Membrane Biology (2024)
-
CDiP technology for reverse engineering of sporadic Alzheimer’s disease
Journal of Human Genetics (2023)
-
Changes in PRC1 activity during interphase modulate lineage transition in pluripotent cells
Nature Communications (2023)
-
Stem cell-derived synthetic embryos self-assemble by exploiting cadherin codes and cortical tension
Nature Cell Biology (2022)
-
Generation of high-performance human cardiomyocytes and engineered heart tissues from extended pluripotent stem cells
Cell Discovery (2022)