Abstract
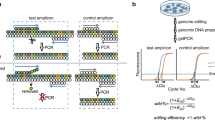
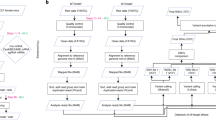
This protocol describes methods for increasing and evaluating the efficiency of genome editing based on the CRISPR–Cas9 (clustered regularly interspaced short palindromic repeats–CRISPR-associated 9) system, transcription activator-like effector nucleases (TALENs) or zinc-finger nucleases (ZFNs). First, Indel Detection by Amplicon Analysis (IDAA) determines the size and frequency of insertions and deletions elicited by nucleases in cells, tissues or embryos through analysis of fluorophore-labeled PCR amplicons covering the nuclease target site by capillary electrophoresis in a sequenator. Second, FACS enrichment of cells expressing nucleases linked to fluorescent proteins can be used to maximize knockout or knock-in editing efficiencies or to balance editing efficiency and toxic/off-target effects. The two methods can be combined to form a pipeline for cell-line editing that facilitates the testing of new nuclease reagents and the generation of edited cell pools or clonal cell lines, reducing the number of clones that need to be generated and increasing the ease with which they are screened. The pipeline shortens the time line, but it most prominently reduces the workload of cell-line editing, which may be completed within 4 weeks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Carroll, D. Genome engineering with targetable nucleases. Annu. Rev. Biochem. 83, 409–439 (2014).
Bae, S., Kweon, J., Kim, H.S. & Kim, J.-S. Microhomology-based choice of Cas9 nuclease target sites. Nat. Methods 11, 705–706 (2014).
Urnov, F.D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Moehle, E.A. et al. Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proc. Natl. Acad. Sci. USA 104, 3055–3060 (2007).
Chen, F. et al. High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases. Nat. Methods 8, 753–755 (2011).
Yang, L. et al. Optimization of scarless human stem cell genome editing. Nucleic Acids Res. 41, 9049–9061 (2013).
Richardson, C.D., Ray, G.J., DeWitt, M.A., Curie, G.L. & Corn, J.E. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat. Biotechnol. 34, 339–344 (2016).
Renaud, J.-B. et al. Improved genome editing efficiency and flexibility using modified oligonucleotides with TALEN and CRISPR-Cas9 nucleases. Cell Rep. 14, 2263–2272 (2016).
Kim, H. & Kim, J.-S. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 15, 321–334 (2014).
Mussolino, C., Mlambo, T. & Cathomen, T. Proven and novel strategies for efficient editing of the human genome. Curr. Opin. Pharmacol. 24, 105–112 (2015).
Maeder, M.L., Thibodeau-Beganny, S., Sander, J.D., Voytas, D.F. & Joung, J.K. Oligomerized pool engineering (OPEN): an 'open-source' protocol for making customized zinc-finger arrays. Nat. Protoc. 4, 1471–1501 (2009).
Kim, H.J., Lee, H.J., Kim, H., Cho, S.W. & Kim, J.-S. Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Res. 19, 1279–1288 (2009).
Sander, J.D., Maeder, M.L. & Joung, J.K. Engineering designer nucleases with customized cleavage specificities. Curr. Protoc. Mol. Biol. 96, 12.13 (2011).
Urnov, F.D., Rebar, E.J., Holmes, M.C., Zhang, H.S. & Gregory, P.D. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 11, 636–646 (2010).
Sanjana, N.E. et al. A transcription activator-like effector toolbox for genome engineering. Nat. Protoc. 7, 171–192 (2012).
Reyon, D. et al. FLASH assembly of TALENs for high-throughput genome editing. Nat. Biotechnol. 30, 460–465 (2012).
Sander, J.D. & Joung, J.K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Wright, A.V., Nuñez, J.K. & Doudna, J.A. Biology and applications of CRISPR systems: harnessing nature's toolbox for genome engineering. Cell 164, 29–44 (2016).
Tsai, S.Q. & Joung, J.K. Defining and improving the genome-wide specificities of CRISPR-Cas9 nucleases. Nat. Rev. Genet. 17, 300–312 (2016).
Yang, Z. et al. Fast and sensitive detection of indels induced by precise gene targeting. Nucleic Acids Res. 43, e59 (2015).
Duda, K. et al. High-efficiency genome editing via 2A-coupled co-expression of fluorescent proteins and zinc finger nucleases or CRISPR/Cas9 nickase pairs. Nucleic Acids Res. 42, e84 (2014).
Yang, Z. et al. Engineered CHO cells for production of diverse, homogeneous glycoproteins. Nat. Biotechnol. 33, 842–844 (2015).
Carrington, B., Varshney, G.K., Burgess, S.M. & Sood, R. CRISPR-STAT: an easy and reliable PCR-based method to evaluate target-specific sgRNA activity. Nucleic Acids Res. 43, e157 (2015).
Certo, M.T. et al. Tracking genome engineering outcome at individual DNA breakpoints. Nat. Methods 8, 671–676 (2011).
Mandal, P.K. et al. Efficient ablation of genes in human hematopoietic stem and effector cells using CRISPR/Cas9. Cell Stem Cell 15, 643–652 (2014).
Ding, Q. et al. A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell 12, 238–251 (2013).
Kim, H. et al. Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nat. Methods 8, 941–943 (2011).
Kim, H. et al. Magnetic separation and antibiotics selection enable enrichment of cells with ZFN/TALEN-induced mutations. PLoS One 8, e56476 (2013).
Ramakrishna, S. et al. Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nuclease-induced mutations. Nat. Commun. 5, 3378 (2014).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Miller, J.C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat. Biotechnol. 25, 778–785 (2007).
Yeung, A.T., Hattangadi, D., Blakesley, L. & Nicolas, E. Enzymatic mutation detection technologies. BioTechniques 38, 749–758 (2005).
Vouillot, L., Thélie, A. & Pollet, N. Comparison of T7E1 and surveyor mismatch cleavage assays to detect mutations triggered by engineered nucleases. G3 (Bethesda) 5, 407–415 (2015).
Oleykowski, C.A., Bronson Mullins, C.R., Godwin, A.K. & Yeung, A.T. Mutation detection using a novel plant endonuclease. Nucleic Acids Res. 26, 4597–4602 (1998).
Till, B.J., Burtner, C., Comai, L. & Henikoff, S. Mismatch cleavage by single-strand specific nucleases. Nucleic Acids Res. 32, 2632–2641 (2004).
Cross, M.J., Waters, D.L.E., Lee, L.S. & Henry, R.J. Endonucleolytic mutation analysis by internal labeling (EMAIL). Electrophoresis 29, 1291–1301 (2008).
Tsuji, T. & Niida, Y. Development of a simple and highly sensitive mutation screening system by enzyme mismatch cleavage with optimized conditions for standard laboratories. Electrophoresis 29, 1473–1483 (2008).
Qiu, P. et al. Mutation detection using Surveyor™ nuclease. BioTechniques 36, 702–707 (2004).
Mashal, R.D., Koontz, J. & Sklar, J. Detection of mutations by cleavage of DNA heteroduplexes with bacteriophage resolvases. Nat. Genet. 9, 177–183 (1995).
Güell, M., Yang, L. & Church, G.M. Genome editing assessment using CRISPR Genome Analyzer (CRISPR-GA). Bioinformatics 30, 2968–2970 (2014).
Brinkman, E.K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Mock, U., Hauber, I. & Fehse, B. Digital PCR to assess gene-editing frequencies (GEF-dPCR) mediated by designer nucleases. Nat. Protoc. 11, 598–615 (2016).
Orlando, S.J. et al. Zinc-finger nuclease-driven targeted integration into mammalian genomes using donors with limited chromosomal homology. Nucleic Acids Res. 38, e152 (2010).
Maresca, M., Lin, V.G., Guo, N. & Yang, Y. Obligate Ligation-Gated Recombination (ObLiGaRe): custom-designed nuclease-mediated targeted integration through nonhomologous end joining. Genome Res. 23, 539–546 (2013).
Carlson, D.F. et al. Efficient TALEN-mediated gene knockout in livestock. Proc. Natl. Acad. Sci. USA 109, 17382–17387 (2012).
Frank, S., Skryabin, B.V. & Greber, B. A modified TALEN-based system for robust generation of knock-out human pluripotent stem cell lines and disease models. BMC Genomics 14, 1–9 (2013).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Shen, B. et al. Efficient genome modification by CRISPR-Cas9 nickase with minimal off-target effects. Nat. Methods 11, 399–402 (2014).
Cho, S.W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014).
Kim, S., Kim, D., Cho, S.W., Kim, J. & Kim, J.-S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Zuris, J.A. et al. Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat. Biotechnol. 33, 73–80 (2015).
Liang, X. et al. Rapid and highly efficient mammalian cell engineering via Cas9 protein transfection. J. Biotechnol. 208, 44–53 (2015).
Yu, X. et al. Improved delivery of Cas9 protein/gRNA complexes using lipofectamine CRISPRMAX. Biotechnol. Lett. 38, 919–929 (2016).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Dieffenbach, C.W., Lowe, T.M. & Dveksler, G.S. General concepts for PCR primer design. Genome Res. 3, S30–S37 (1993).
Kibbe, W.A. OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res. 35, W43–W46 (2007).
Korbie, D.J. & Mattick, J.S. Touchdown PCR for increased specificity and sensitivity in PCR amplification. Nat. Protoc. 3, 1452–1456 (2008).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Frock, R.L. et al. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat. Biotechnol. 33, 179–186 (2015).
Wang, X. et al. Unbiased detection of off-target cleavage by CRISPR-Cas9 and TALENs using integrase-defective lentiviral vectors. Nat. Biotechnol. 33, 175–178 (2015).
Fu, Y., Sander, J.D., Reyon, D., Cascio, V.M. & Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Kim, D. et al. Digenome-seq: genome-wide profiling of CRISPR-Cas9 off-target effects in human cells. Nat. Methods 12, 237–243 (2015).
Tsai, S.Q. et al. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat. Biotechnol. 32, 569–576 (2014).
Guilinger, J.P., Thompson, D.B. & Liu, D.R. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat. Biotechnol. 32, 577–582 (2014).
Kleinstiver, B.P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Slaymaker, I.M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2015).
Muller, M. et al. Streptococcus thermophilus CRISPR-Cas9 systems enable specific editing of the human genome. Mol. Ther. 24, 636–644 (2016).
Bolukbasi, M.F., Gupta, A. & Wolfe, S.A. Creating and evaluating accurate CRISPR-Cas9 scalpels for genomic surgery. Nat. Methods 13, 41–50 (2016).
Koo, T., Lee, J. & Kim, J.-S. Measuring and reducing off-target activities of programmable nucleases including CRISPR-Cas9. Mol. Cells 38, 475–481 (2015).
Doyon, Y. et al. Enhancing zinc-finger-nuclease activity with improved obligate heterodimeric architectures. Nat. Methods 8, 74–79 (2011).
Nakade, S. et al. Microhomology-mediated end-joining-dependent integration of donor DNA in cells and animals using TALENs and CRISPR/Cas9. Nat. Commun. 5, 5560 (2014).
Sakuma, T., Nakade, S., Sakane, Y., Suzuki, K.-I.T. & Yamamoto, T. MMEJ-assisted gene knock-in using TALENs and CRISPR-Cas9 with the PITCh systems. Nat. Protoc. 11, 118–133 (2016).
Yoshimi, K. et al. ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat. Commun. 7, 10431 (2016).
Chu, V.T. et al. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat. Biotechnol. 33, 543–548 (2015).
Park, J., Bae, S. & Kim, J.-S. Cas-Designer: a web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 31, 4014–4016 (2015).
Sander, J.D. et al. ZiFiT (Zinc Finger Targeter): an updated zinc finger engineering tool. Nucleic Acids Res. 38, W462–W468 (2010).
Heigwer, F., Kerr, G. & Boutros, M. E-CRISP: fast CRISPR target site identification. Nat Methods 11, 122–123 (2014).
Hodgkins, A. et al. WGE: a CRISPR database for genome engineering. Bioinformatics 31, 3078–3080 (2015).
Moreno-Mateos, M.A. et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat. Methods 12, 982–988 (2015).
Singh, R., Kuscu, C., Quinlan, A., Qi, Y. & Adli, M. Cas9-chromatin binding information enables more accurate CRISPR off-target prediction. Nucleic Acids Res. 43, e118 (2015).
Lee, C.M., Cradick, T.J., Fine, E.J. & Bao, G. Nuclease target site selection for maximizing on-target activity and minimizing off-target effects in genome editing. Mol. Ther. 24, 475–487 (2016).
Magnuson, V.L. et al. Substrate nucleotide-determined non-templated addition of adenine by Taq DNA polymerase: implications for PCR-based genotyping and cloning. BioTechniques 21, 700–709 (1996).
Brownstein, M.J., Carpten, J.D. & Smith, J.R. Modulation of non-templated nucleotide addition by Taq DNA polymerase: primer modifications that facilitate genotyping. BioTechniques 20, 1004–1006 (1996).
Acknowledgements
We thank A. Fossum (BRIC), C.K. Beier Holden, C. Andersen, and S. Narimatsu (Copenhagen Center for Glycomics), and A. Dürr (the Danish Stem Cell Center) for excellent technical assistance. We thank S. Miller, Washington University, for critical reading of the manuscript. This work was supported by the Danish Cancer Society (R124-A7632-15-S2 to M.F.), the Danish Council for Independent Research (0602-02368B to M.F.), the University of Copenhagen Excellence Programme for Interdisciplinary Research—CDO2016 (to M.F., E.P.B. and H.H.W.), the Novo Nordisk Foundation (to M.F., Y.N. and Z.Y., and NNF15CC0018344 to E.A.O.), the Lundbeck Foundation (R165-2013-15743 to F.N.), the National Institutes of Health (R21CA184656 to S.H.H.), the Medical Research Council UK (U117581329 to E.A.O.) and the Danish National Research Foundation (DNRF107 to E.P.B., H.H.W., Y.N., Z.Y. and H.C.).
Author information
Authors and Affiliations
Contributions
All authors were involved in designing and/or performing experiments underlying the manuscript or the development of the methods described. L.A.L., E.P.B. and M.F. wrote the manuscript with comments from the other authors.
Corresponding authors
Ethics declarations
Competing interests
E.P.B. declares competing financial interests (see the HTML version of this article for details). E.P.B. possesses the rights to a patent application describing the IDAA methodology.
Integrated supplementary information
Supplementary Figure 1 Sensitivity and quantitative performance of IDAA performed in an ABI 3500 instrument is comparable to Next-Generation Sequencing (MiSeq).
Indel analysis by (a) IDAA or (b) MiSeq of PCR amplicons from the Cosmc locus edited by CRISPR/Cas9 in a CHO cell pool. On the x-axis, indel sizes (bp) are indicated for some of the amplicons. In the zoom-in panels, a stippled line indicates the background signal level, defined based on the signals from 45-55 bp insertions that were considered background, since CRISPR/Cas9 rarely elicits insertions of such sizes. (c) Graphs of frequencies of indels a-m from the IDAA and MiSeq analyses. Note the high degree of agreement between the two analyses.
Supplementary Figure 2 IDAA can generate useful indel profiles from minute amounts of IDAA PCR amplicons.
The Trp53 locus was targeted with CRISPR/Cas9 in a mouse Neuro2A cell pool and the target site was amplified by IDAA PCR. The indicated amounts of the IDAA PCR were analyzed by (a) agarose gel electrophoresis (an arrow indicates the IDAA PCR amplicon) or (b) IDAA capillary electrophoresis. The size and frequency of selected indels are indicated. IDAA was performed in an ABI 3500 instrument.
Note that the indel profiles are almost identical across the range of IDAA amplicons analyzed and that a high-quality profile can be generated from amplicon amounts barely visible or undetectable by agarose gel analysis.
Supplementary Figure 3 Upper and lower detection levels for IDAA amplicons in ABI 3130 and 3500 instruments.
(a) Various amounts of a wt IDAA amplicon from a non-edited control sample were analyzed in ABI 3130 and 3500 instruments. When the loaded amplicons exceed the dynamic range, a smaller peak artifact often appears 20-30 bp ahead of the true signal (indicated by asterisk in the upper 3130 panel). (b) Zoom in on the peaks boxed in (a). The lower 3130 panel illustrates that also below the dynamic range, a specific signal can be discriminated from background signals.
Supplementary Figure 4 IDAA reveals the indel signature of a given gRNA, TALEN or ZFN.
IDAA profiles for two independent experiments using the same construct of FP-linked CRISPR/Cas9 (a,b), TALENs (c) or ZFNs (d), targeting various loci in K562 or HEK293 cells, as indicated. Prior to analysis, the nuclease-transfected cells were subjected to bulk FACS for the top 30% most fluorescent cells. The size and frequency of selected indels are indicated. IDAA profiles were generated in an ABI 3130 instrument.
Supplementary Figure 5 Using IDAA to estimate probability of germline transmission of indels from F0 mosaic fish and to identify F1 mutant fish in Danio rerio genome editing.
(a) In zebrafish genome editing, a major challenge is determination of the nature and degree of indel mosaicism and hence, probability of germline transmission of indels from F0 fish derived from one-cell embryos injected with Cas9 and gRNA. IDAA enables easy and rapid evaluation of these variables through analysis of F0 fin DNA, allowing optimal set-up of F0 breeding pairs. IDAA also enables easy identification of indels in subsequent generations through fin DNA analysis. (b) Somatic IDAA profiles of the bambi locus in fin DNA of F0 fish targeted at the one-cell embryo stage via injection of Cas9 and gRNA. F0 male #1 and F0 female #4 were chosen for mating due to the presence of predominant indels indicated by open or closed circles, suggesting high probability of germline transmission of these mutations. (c) IDAA profiles of 200 downstream F1 fish, showing that 4 out of 5 fish harbored the predominant indels identified in the F0 breeding pair, of which 3 were in the biallelic state (F1 #1, #3, #4) and one homozygous for an indel (F1 #4). In this example, IDAA was performed on embryos, but IDAA could also have been performed on fin DNA. The size and frequency of selected indels are indicated. IDAA profiles were generated in an ABI 3130 instrument.
Supplementary Figure 6 QCgRNA amplicon expression cassettes.
(a) Schematic showing the various elements of the QCgRNA amplicon primers QCgFwd, QCgX and QCgRev. Sequences annealing to U6 promotor template are shown in orange; gRNA design in red; tracr elements in blue; restriction enzyme sites for sub-cloning to pEPB104 in italics. Note that the gRNA design is incorporated into QCgX as the complementary sequence to the target, which is shown in green. The nucleotide (c) is only included in the QCgX primer, if the gRNA (=target) does not contain a G as the first (5´) nucleotide, which is the case in the present example. (b) Agarose gel (2%) electrophoresis check for the formation of full-length (f) products (arrow) by QCgRNA amplicon tri-primer PCRs, as compared to control (c) PCRs containing only QCgFwd and QCgX primers (Step 3 in Procedure). Various amounts of MassRuler Low Range DNA ladder (M) are run alongside to enable quantitation of the QCgRNA amplicons. (c) The indel profiles elicited by a gRNA design in a QCgRNA amplicon and a plasmid vector are identical, as illustrated by targeting GALNT10 in HEK293 cells. The GALNT10 QCgRNA amplicon was subcloned into pEPB104 plasmid (Addgene #68369; Supplementary Sequence 1) using EcoRI and KpnI restriction endonuclease sites present in both constructs. (d) A QCgRNA design found non-functional in one cell type typically remains non-functional when tested in other cell types, as illustrated with a POMT2 QCgRNA. IDAA profiles were generated in an (c) ABI 3500 or and (d) ABI 3130 instrument.
Supplementary information
Supplementary Figures and Text
Supplementary Figures 1–6, Supplementary Tables 1–3, the Supplementary Note, the Supplementary Data and the Supplementary Manual (PDF 5743 kb)
Rights and permissions
About this article
Cite this article
Lonowski, L., Narimatsu, Y., Riaz, A. et al. Genome editing using FACS enrichment of nuclease-expressing cells and indel detection by amplicon analysis. Nat Protoc 12, 581–603 (2017). https://doi.org/10.1038/nprot.2016.165
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.165
This article is cited by
-
ADAM12 expression is upregulated in cancer cells upon radiation and constitutes a prognostic factor in rectal cancer patients following radiotherapy
Cancer Gene Therapy (2023)
-
ST3GalIV drives SLeX biosynthesis in gastrointestinal cancer cells and associates with cancer cell motility
Glycoconjugate Journal (2023)
-
Identification of global inhibitors of cellular glycosylation
Nature Communications (2023)
-
A roadmap for translational cancer glycoimmunology at single cell resolution
Journal of Experimental & Clinical Cancer Research (2022)
-
CRISPR/Cas9-mediated knockout of clinically relevant alloantigenes in human primary T cells
BMC Biotechnology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.