Abstract
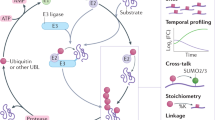
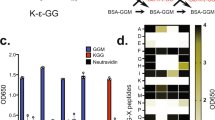
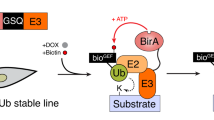
The protein called 'small ubiquitin-like modifier' (SUMO) is post-translationally linked to target proteins at the ɛ-amino group of lysine residues. This 'SUMOylation' alters the behavior of the target protein, a change that is utilized to regulate diverse cellular processes. Understanding the target-specific consequences of SUMO modification requires knowledge of the location of conjugation sites, and we have developed a straightforward protocol for the proteome-wide identification of SUMO modification sites using mass spectrometry (MS). The approach described herein requires the expression of a mutant form of SUMO, in which the residue preceding the C-terminal Gly-Gly (diGly) is replaced with a lysine (SUMOKGG). Digestion of SUMOKGG protein conjugates with endoproteinase Lys-C yields a diGly motif attached to target lysines. Peptides containing this adduct are enriched using a diGly-Lys (K-ɛ-GG)-specific antibody and identified by MS. This diGly signature is characteristic of SUMOKGG conjugation alone, as no other ubiquitin-like protein (Ubl) yields this adduct upon Lys-C digestion. We have demonstrated the utility of the approach in SUMOylation studies, but, in principle, it may be adapted for the site-specific identification of proteins modified by any Ubl. Starting from cell lysis, this protocol can be completed in ∼5 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Geiss-Friedlander, R. & Melchior, F. Concepts in sumoylation: a decade on. Nat. Rev. Mol. Cell Biol. 8, 947–956 (2007).
Hay, R.T. SUMO: a history of modification. Mol. Cell 18, 1–12 (2005).
Hay, R.T. SUMO-specific proteases: a twist in the tail. Trends Cell Biol. 17, 370–376 (2007).
Filosa, G., Barabino, S.M. & Bachi, A. Proteomics strategies to identify SUMO targets and acceptor sites: a survey of RNA-binding proteins SUMOylation. Neuromolecular Med. 15, 661–676 (2013).
Vertegaal, A.C. et al. Distinct and overlapping sets of SUMO-1 and SUMO-2 target proteins revealed by quantitative proteomics. Mol. Cell. Proteomics 5, 2298–2310 (2006).
Golebiowski, F. et al. System-wide changes to SUMO modifications in response to heat shock. Sci. Signal. 2, ra24 (2009).
Tatham, M.H., Matic, I., Mann, M. & Hay, R.T. Comparative proteomic analysis identifies a role for SUMO in protein quality control. Sci. Signal. 4, rs4 (2011).
Bruderer, R. et al. Purification and identification of endogenous polySUMO conjugates. EMBO Rep. 12, 142–148 (2011).
Becker, J. et al. Detecting endogenous SUMO targets in mammalian cells and tissues. Nat. Struct. Mol. Biol. 20, 525–531 (2013).
Olsen, J.V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325, 834–840 (2009).
Xu, G., Paige, J.S. & Jaffrey, S.R. Global analysis of lysine ubiquitination by ubiquitin remnant immunoaffinity profiling. Nat. Biotechnol. 28, 868–873 (2010).
Steen, H. & Mann, M. The ABC's (and XYZ's) of peptide sequencing. Nat. Rev. Mol. Cell Biol. 5, 699–711 (2004).
Olsen, J.V., Ong, S.E. & Mann, M. Trypsin cleaves exclusively C-terminal to arginine and lysine residues. Mol. Cell. Proteomics 3, 608–614 (2004).
Pedrioli, P.G. et al. Automated identification of SUMOylation sites using mass spectrometry and SUMmOn pattern recognition software. Nat. Methods 3, 533–539 (2006).
Matic, I. et al. In vivo identification of human small ubiquitin-like modifier polymerization sites by high accuracy mass spectrometry and an in vitro to in vivo strategy. Mol. Cell. Proteomics 7, 132–144 (2008).
Hsiao, H.H., Meulmeester, E., Frank, B.T., Melchior, F. & Urlaub, H. 'ChopNSpice,' a mass spectrometric approach that allows identification of endogenous small ubiquitin-like modifier-conjugated peptides. Mol. Cell. Proteomics 8, 2664–2675 (2009).
Tammsalu, T. et al. Proteome-wide identification of SUMO2 modification sites. Sci. Signal. 7, rs2 (2014).
Raijmakers, R., Neerincx, P., Mohammed, S. & Heck, A.J. Cleavage specificities of the brother and sister proteases Lys-C and Lys-N. Chem. Commun. (Camb) 46, 8827–8829 (2010).
Drapeau, G.R., Boily, Y. & Houmard, J. Purification and properties of an extracellular protease of Staphylococcus aureus. J. Biol. Chem. 247, 6720–6726 (1972).
Impens, F., Radoshevich, L., Cossart, P. & Ribet, D. Mapping of SUMO sites and analysis of SUMOylation changes induced by external stimuli. Proc. Natl. Acad. Sci. USA 111, 12432–12437 (2014).
Hendriks, I.A. et al. Uncovering global SUMOylation signaling networks in a site-specific manner. Nat. Struct. Mol. Biol. 21, 927–936 (2014).
Lamoliatte, F. et al. Large-scale analysis of lysine SUMOylation by SUMO remnant immunoaffinity profiling. Nat. Commun. 5, 5409 (2014).
Hobbs, S., Jitrapakdee, S. & Wallace, J.C. Development of a bicistronic vector driven by the human polypeptide chain elongation factor 1alpha promoter for creation of stable mammalian cell lines that express very high levels of recombinant proteins. Biochem. Biophys. Res. Commun. 252, 368–372 (1998).
Udeshi, N.D. et al. Refined preparation and use of anti-diglycine remnant (K-epsilon-GG) antibody enables routine quantification of 10,000s of ubiquitination sites in single proteomics experiments. Mol. Cell. Proteomics 12, 825–831 (2013).
Nielsen, M.L. et al. Iodoacetamide-induced artifact mimics ubiquitination in mass spectrometry. Nat. Methods 5, 459–460 (2008).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Acknowledgements
T.T. is funded through the EU Seventh Framework Programme (FP7A-PEOPLE-2011-ITN). I.M. was supported by a Sir Henry Wellcome Fellowship (Wellcome Trust 088957/Z/09/Z) and the DFG (Deutsche Forschungsgemeinschaft; Cologne Cluster of Excellence in Cellular Stress Responses in Aging-Associated Diseases). E.G.J. and M.H.T. are funded through a Cancer Research UK programme grant (C434/A13067). R.T.H. holds a Wellcome Trust Senior Investigator Award (098391/Z/12/Z).
Author information
Authors and Affiliations
Contributions
T.T. optimized sample processing and conducted sample preparation, MS analysis, data analysis and bioinformatic analysis. R.T.H. and I.M. conceived the enrichment and sample processing strategy. I.M. developed the workflow for the MS analysis and edited the manuscript. E.G.J. purified recombinant proteins and conducted preliminary in vitro assays. A.F.M.I. created the stable cell lines. M.H.T. consulted at multiple stages. T.T., R.T.H. and M.H.T. co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Tammsalu, T., Matic, I., Jaffray, E. et al. Proteome-wide identification of SUMO modification sites by mass spectrometry. Nat Protoc 10, 1374–1388 (2015). https://doi.org/10.1038/nprot.2015.095
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.095
This article is cited by
-
SUMOylation of AnxA6 facilitates EGFR-PKCα complex formation to suppress epithelial cancer growth
Cell Communication and Signaling (2023)
-
The scaffold protein IQGAP1 links heat-induced stress signals to alternative splicing regulation in gastric cancer cells
Oncogene (2021)
-
The in vivo ISGylome links ISG15 to metabolic pathways and autophagy upon Listeria monocytogenes infection
Nature Communications (2019)
-
An E2-ubiquitin thioester-driven approach to identify substrates modified with ubiquitin and ubiquitin-like molecules
Nature Communications (2018)
-
Proteome-wide identification of ubiquitin interactions using UbIA-MS
Nature Protocols (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.