Abstract
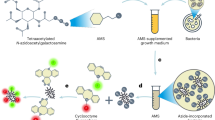
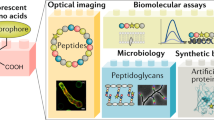
Fluorescent D-amino acids (FDAAs) are efficiently incorporated into the peptidoglycans (PGs) of diverse bacterial species at the sites of PG biosynthesis, allowing specific and covalent probing of bacterial growth with minimal perturbation. Here we provide a protocol for the synthesis of four FDAAs emitting light in blue (HCC-amino-D-alanine, HADA), green (NBD-amino-D-alanine, NADA, and fluorescein-D-lysine, FDL) or red (TAMRA-D-lysine, TDL) and for their use in PG labeling of live bacteria. Our modular synthesis protocol gives easy access to a library of different FDAAs made with commercially available fluorophores and diamino acid starting materials. Molecules can be synthesized in a typical chemistry laboratory in 2–3 d using standard chemical transformations. The simple labeling procedure involves the addition of the FDAAs to a bacterial sample for the desired labeling duration and stopping further label incorporation by fixing the cells with cold 70% (vol/vol) ethanol or by washing away excess dye. We discuss several scenarios for the use of these labels in fluorescence microscopy applications, including short or long labeling durations, and the combination of different labels in pure culture (e.g., for 'virtual time-lapse' microscopy) or in situ labeling of complex environmental samples. Depending on the experiment, FDAA labeling can take as little as 30 s for a rapidly growing species such as Escherichia coli.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Schneider, T. & Sahl, H.-G. An oldie but a goodie–cell wall biosynthesis as an antibiotic target pathway. Int. J. Med. Microbiol. 300, 2010 (2010).
Gould, I.M. Coping with antibiotic resistance: the impending crisis. Int. J. Antimicrob. Agents 36, S1–S2 (2010).
Neu, H.C. The crisis in antibiotic resistance. Science 257, 1064–1073 (1992).
van Dam, V., Olrichs, N. & Breukink, E. Specific labeling of peptidoglycan precursors as a tool for bacterial cell wall studies. Chem. Bio. Chem. 10, 617–624 (2009).
van der Donk, W.A. Lighting up the nascent cell wall. ACS Chem. Biol. 1, 425–428 (2006).
Kuru, E. et al. In situ probing of newly synthesized peptidoglycan in live bacteria with fluorescent D-amino acids. Angew. Chem. Int. Ed. 51, 12519–12523 (2012).
Pilhofer, M. et al. Discovery of chlamydial peptidoglycan reveals bacteria with murein sacculi but without FtsZ. Nat. Commun. 4, 2856 (2013).
Pinho, M.G., Kjos, M. & Veening, J.-V. How to get (a)round: mechanisms controlling growth and division of coccoid bacteria. Nat. Rev. Microbiol. 11, 601–614 (2013).
Ranjit, D.K. & Young, K.D. The Rcs stress response and accessory envelope proteins are required for de novo generation of cell shape in Escherichia coli. J. Bacteriol. 195, 2452–2462 (2013).
Tocheva, E.I. et al. Peptidoglycan transformations during Bacillus subtilis sporulation. Mol. Microbiol. 88, 673–686 (2013).
Daniel, R.A. & Errington, J. Control of cell morphogenesis in bacteria: two distinct ways to make a rod-shaped cell. Cell 113, 767–776 (2003).
Tiyanont, K. et al. Imaging peptidoglycan biosynthesis in Bacillus subtilis with fluorescent antibiotics. Proc. Natl. Acad. Sci. USA 103, 11033–11038 (2006).
de Pedro, M.A., Quintela, J., Holtje, J. & Schwarz, H. Murein segregation in Escherichia coli. J. Bacteriol. 179, 2823–2834 (1997).
Olrichs, N.K. et al. A novel in vivo cell-wall labeling approach sheds new light on peptidoglycan synthesis in Escherichia coli. Chembiochem 12, 1124–1133 (2011).
Sadamoto, R., Niikura, K., Monde, K. & Nishimura, S.-I. Cell wall engineering of living bacteria through biosynthesis. Methods Enzymol. 362, 273–286 (2003).
Sadamoto, R. et al. Cell-wall engineering of living bacteria. J. Am. Chem. Soc. 124, 9018–9019 (2002).
Sadamoto, R. et al. Control of bacterial adhesion by cell-wall engineering. J. Am. Chem. Soc. 126, 3755–3761 (2004).
Liechti, G.W. et al. A new metabolic cell-wall labelling method reveals peptidoglycan in Chlamydia trachomatis. Nature 506, 507–510 (2013).
Schouten, J.A. et al. Fluorescent reagents for in vitro studies of lipid-linked steps of bacterial peptidoglycan biosynthesis: derivatives of UDPMurNAc-pentapeptide containing D-cysteine at position 4 or 5. Mol. BioSyst. 2, 484–491 (2006).
de Pedro, M.A., Young, K.D., Höltje, J.-V. & Schwarz, H. Branching of Eschereichia coli cells arises from multiple sites of inert peptidoglycan. J. Bacteriol. 185, 1147–1152 (2003).
Brown, P.J.B. et al. Polar growth in the alphaproteobacterial order rhizobiales. Proc. Natl. Acad. Sci. USA 109, 1697–1701 (2012).
Cava, F., Lam, H., de Pedro, M.A. & Waldor, M.K. Emerging knowledge of regulatory roles of D-amino acids in bacteria. Cell. Mol. Life Sci. 68, 817–831 (2011).
Lam, H. et al. d-Amino acids govern stationary phase cell wall remodeling in bacteria. Science 325, 1552–1555 (2009).
Lupoli, T.J. et al. Transpeptidase-mediated incorporation of D-amino acids into bacterial peptidoglycan. J. Am. Chem. Soc. 133, 10748–10751 (2011).
Cava, F., de Pedro, M.A., Lam, H., Davis, B.M. & Waldor, M.K. Distinct pathways for modification of the bacterial cell wall by non-canonical D-amino acids. EMBO J. 30, 3442–3453 (2011).
Siegrist, M.S. et al. d-Amino acid chemical reporters reveal peptidoglycan dynamics of an intracellular pathogen. ACS Chem. Biol. 8, 500–505 (2013).
Zapun, A. et al. In vitro reconstitution of peptidoglycan assembly from the Gram-positive pathogen Streptococcus pneumoniae. ACS Chem. Biol. 8, 2688–2696 (2013).
Bugg, T.D.H. & Walsh, C.T. lntracellular steps of bacterial cell wall peptidoglycan biosynthesis: enzymology, antibiotics, and antibiotic resistance. Nat. Prod. Rep. 9, 199–215 (1992).
Höltje, J.-V. Growth of the stress-bearing and shape-maintaining murein sacculus of Escherichia coli. Microbiol. Mol. Biol. Rev. 62, 181–203 (1998).
Magnet, S., Dubost, L., Marie, A., Arthur, M. & Guttmann, L. Identification of the L,D-transpeptidases for peptidoglycan cross-linking in Escherichia coli. J. Bacteriol. 190, 4782–4785 (2008).
Vollmer, W., Blanot, D. & de Pedro, M.A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Kolb, H.C., Finn, M.G. & Sharpless, K.B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. 40, 2004–2021 (2001).
Prescher, J.A. & Bertozzi, C.R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–20 (2005).
Fura, J.M., Sabulski, M.J. & Pires, M.M. Amino acid mediated recruitment of endogenous antibodies to bacterial surfaces. ACS Chem. Biol. 9, 1480–1489 (2014).
Kerr, C.H., Culham, D.E., Marom, D. & Wood, J.M. Salinity-dependent impacts of ProQ, Prc, and Spr deficiencies on Escherichia coli cell structure. J. Bacteriol. 196, 1286–1296 (2014).
Ursell, T.S. et al. Rod-like bacterial shape is maintained by feedback between cell curvature and cytoskeletal localization. Proc. Natl. Acad. Sci. USA 111, E1025–E1034 (2014).
Jiang, C., Brown, P.J., Ducret, A. & Brun, Y.V. Sequential evolution of bacterial morphology by co-option of a developmental regulator. Nature 506, 489–493 (2014).
Eun, Y.J. et al. Divin: a small molecule inhibitor of bacterial divisome assembly. J. Am. Chem. Soc. 135, 9768–9776 (2013).
Fenton, A.K. & Gerdes, K. Direct interaction of FtsZ and MreB is required for septum synthesis and cell division in Escherichia coli. EMBO J. 32, 1953–1965 (2013).
Cava, F., Kuru, E., Brun, Y.V. & de Pedro, M.A. Modes of cell wall growth differentiation in rod-shaped bacteria. Curr. Opin. Microbiol. 16, 731–737 (2013).
Lupoli, T.J. et al. Lipoprotein activators stimulate Escherichia coli penicillin-binding proteins by different mechanisms. J. Am. Chem. Soc. 136, 52–55 (2014).
Shieh, P., Siegrist, M.S., Cullen, A.J. & Bertozzi, C.R. Imaging bacterial peptidoglycan with near-infrared fluorogenic azide probes. Proc. Natl. Acad. Sci. USA 111, 5456–5461 (2014).
Takacs, C.N. et al. Growth medium-dependent glycine incorporation into the peptidoglycan of Caulobacter crescentus. PLoS ONE 8, e57579 (2013).
Fleurie, A. et al. Interplay of the serine/threonine-kinase StkP and the paralogs DivIVA and GpsB in pneumococcal cell elongation and division. PLoS Genet. 10, e1004275 (2014).
Acknowledgements
This work was supported by grants from the National Institutes of Health (AI 059327 to M.S.V. and GM051986 to Y.V.B.). We thank E. Garner, K.C. Huang, P. Brown, V. Hughes, A. Ducret, S. Carmody and B. Turner for their helpful discussions.
Author information
Authors and Affiliations
Contributions
E.K., M.S.V. and Y.V.B. designed the study; E.K. designed, and S.T. and E.H. synthesized FDAAs; E.K. designed and conducted experiments involving microscopy; and E.K., S.T., E.H., Y.V.B. and M.S.V. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kuru, E., Tekkam, S., Hall, E. et al. Synthesis of fluorescent D-amino acids and their use for probing peptidoglycan synthesis and bacterial growth in situ. Nat Protoc 10, 33–52 (2015). https://doi.org/10.1038/nprot.2014.197
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.197
This article is cited by
-
Two broadly conserved families of polyprenyl-phosphate transporters
Nature (2023)
-
Spatial transcriptome uncovers rich coordination of metabolism in E. coli K12 biofilm
Nature Chemical Biology (2023)
-
Hyphal compartmentalization and sporulation in Streptomyces require the conserved cell division protein SepX
Nature Communications (2022)
-
Dynamics of plasmid-mediated niche invasion, immunity to invasion, and pheromone-inducible conjugation in the murine gastrointestinal tract
Nature Communications (2022)
-
Magnesium rescues the morphology of Bacillus subtilis mreB mutants through its inhibitory effect on peptidoglycan hydrolases
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.