Abstract
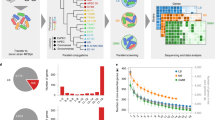
Insertional mutagenesis and depletion (iMAD) is a genetic screening strategy for dissecting complex interactions between two organisms. The simultaneous genetic manipulation of both organisms allows the identification of aggravating and alleviating genetic interactions between pairs of gene disruptions, one from each organism. Hierarchial clustering and genetic interaction networks are then used to identify common behavioral patterns among subsets of genes, which allow functional relationships between proteins and their component pathways to be elucidated. Here we present a protocol for dissecting the interaction between a pathogen (Legionella pneumophila) and its host (cultured Drosophila melanogaster cells) using bacterial mutagenesis and host RNAi. The key stages covered in the PROCEDURE include the design, execution and data analysis of an iMAD screen; details for determining the abundance of individual mutants by microarray analysis and next-generation sequencing are not included because detailed protocols have been published elsewhere. Adapting and optimizing iMAD to a specific experimental system can require 6–18 months. Once a bacterial mutant library, host cell factor depletion strategies and conditions to monitor the interaction are established, an iMAD screen can be completed in 4–8 weeks, depending on the organisms' growth rates, the duration of the interaction and the types of data analysis performed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Typas, A. et al. High-throughput, quantitative analyses of genetic interactions in E. coli. Nat. Methods 5, 781–787 (2008).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Horn, T. et al. Mapping of signaling networks through synthetic genetic interaction analysis by RNAi. Nat. Methods 8, 341–346 (2011).
O'Connor, T.J. et al. Aggravating genetic interactions allow a solution to redundancy in a bacterial pathogen. Science 338, 1440–1444 (2012).
Huang, L. et al. The E Block motif is associated with Legionella pneumophila translocated substrates. Cell. Microbiol. 13, 227–245 (2011).
Zhu, W. et al. Comprehensive identification of protein substrates of the Dot/Icm type IV transporter of Legionella pneumophila. PLoS ONE 6, e17638 (2011).
Dorer, M.S. et al. RNA interference analysis of Legionella in Drosophila cells: exploitation of early secretory apparatus dynamics. PLoS Pathog. 2, e34 (2006).
Sassetti, C.M. et al. Comprehensive identification of conditionally essential genes in mycobacteria. Proc. Natl. Acad. Sci. USA 98, 12712–12717 (2001).
van Opijnen, T. et al. Tn-seq: high-throughput parallel sequencing for fitness and genetic interaction studies in microorganisms. Nat. Methods 6, 767–772 (2009).
Goodman, A.L. et al. Identifying genetic determinants needed to establish a human gut symbiont in its habitat. Cell Host Microbe 6, 279–289 (2009).
Langridge, G.C. et al. Simultaneous assay of every Salmonella typhi gene using one million transposon mutants. Genome Res. 19, 2308–2316 (2009).
Gawronski, J.D. et al. Tracking insertion mutants within libraries by deep sequencing and a genome-wide screen for Haemophilus genes required in the lung. Proc. Natl. Acad. Sci. USA 106, 16422–16427 (2009).
De Jesus, D.A. et al. Analysis of Legionella infection using RNA interference in Drosophila cells. Methods Mol. Biol. 954, 251–264 (2013).
Wong, S.M. et al. High-throughput insertion tracking by deep sequencing for the analysis of bacterial mutants. Methods Mol. Biol. 733, 209–222 (2011).
Murry, J.P. et al. Transposon site hybridization in Mycobacterium tuberculosis. Methods Mol. Biol. 416, 45–59 (2008).
Goodman, A.L. et al. Identifying microbial fitness determinants by insertion sequencing using genome-wide transposon mutant libraries. Nat. Protoc. 6, 1969–1980 (2011).
Lampe, D.J. et al. A purified mariner transposase is sufficient to mediate transposition in vitro. EMBO J. 15, 5470–5479 (1996).
O'Connor, T.J. et al. Minimization of the Legionella pneumophila genome reveals chromosomal regions involved in host range expansion. Proc. Natl. Acad. Sci. USA 108, 14733–14740 (2011).
Chiang, S.U. et al. Construction of a mariner-based transposon for epitope-tagging and genomic targeting. Gene 296, 179–185 (2002).
Crimmins, G.T. et al. Identification of MrtAB, an ABC transporter specifically required for Yersinia pseudotuberculosis to colonize the mesenteric lymph nodes. PLoS Pathog. 8, e1002828 (2012).
Bassik, M.C. et al. A systematic mammalian genetic interaction map reveals pathways underlying ricin susceptibility. Cell 152, 909–922 (2013).
Kim, H.S. et al. Systematic identification of molecular subtype-selective vulnerabilities in non-small-cell lung cancer. Cell 155, 552–566 (2013).
Roguev, A. et al. Quantitative genetic-interaction mapping in mammalian cells. Nat. Methods 10, 432–437 (2013).
Carette, J.E. et al. Haploid genetic screens in human cells identify host factors used by pathogens. Science 326, 1231–1235 (2009).
Xu, M. et al. Down-regulation of ribosomal protein S15A mRNA with a short hairpin RNA inhibits human hepatic cancer cell growth in vitro. Gene 536, 84–89 (2014).
Tang, F.C. et al. Stable suppression of gene expression in murine embryonic stem cells by RNAi directed from DNA vector-based short hairpin RNA. Stem Cells 22, 93–99 (2004).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(–ΔΔCT) method. Methods 25, 402–408 (2001).
US Centers for Disease Control and Prevention. Biosafety in Microbiological and Biomedical Laboratories (BMBL), 5th edn. http://www.cdc.gov/biosafety/publications/bmbl5/ (2009).
van Opijnen, T. & Camilli, A. Transposon insertion sequencing: a new tool for systems-level analysis of microorganisms. Nat. Rev. Microbiol. 11, 435–442 (2013).
Drees, B.L. et al. Derivation of genetic interaction networks from quantitative phenotype data. Genome Biol. 6, R38 (2005).
St. Onge, R.P. et al. Systematic pathway analysis using high-resolution fitness profiling of combinatorial gene deletions. Nat. Genet. 39, 199–206 (2007).
Eisen, M.B. et al. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Shannon, P. Cytoscape: a software environment for integrate models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Acknowledgements
This work was supported by the Howard Hughes Medical Institute and a Natalie Zucker Fellowship to T.J.O. R.R.I. is a Howard Hughes Investigator.
Author information
Authors and Affiliations
Contributions
Both authors wrote the paper and designed the experiments. T.J.O. conducted the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
O'Connor, T., Isberg, R. iMAD, a genetic screening strategy for dissecting complex interactions between a pathogen and its host. Nat Protoc 9, 1916–1930 (2014). https://doi.org/10.1038/nprot.2014.133
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.133
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.