Abstract
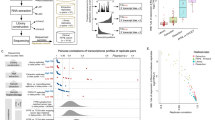
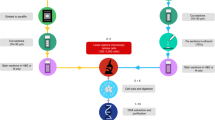
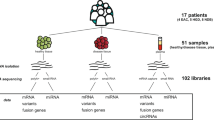
Genomic technologies have revolutionized our understanding of complex Mendelian diseases and cancer. Solid tumors present several challenges for genomic analyses, such as tumor heterogeneity and tumor contamination with surrounding stroma and infiltrating lymphocytes. We developed a protocol to (i) select tissues of high cellular purity on the basis of histological analyses of immediately flanking sections and (ii) simultaneously extract genomic DNA (gDNA), mRNA, noncoding RNA (ncRNA; enriched in miRNA) and protein from the same tissues. After tissue selection, about 12–16 extractions of DNA, RNA or protein can be obtained per day. Compared with other similar approaches, this fast and reliable methodology allowed us to identify mutations in tumors with remarkable sensitivity and to perform integrative analyses of whole-genome and exome data sets, DNA copy numbers (by single-nucleotide polymorphism (SNP) arrays), gene expression data (by transcriptome profiling and quantitative PCR (qPCR)) and protein levels (by western blotting and immunohistochemical analysis) from the same samples. Although we focused on renal cell carcinoma, this protocol may be adapted with minor changes to any human or animal tissue to obtain high-quality and high-yield nucleic acids and proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bamshad, M.J. et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat. Rev. Genet. 12, 745–755 (2011).
Mwenifumbo, J.C. & Marra, M.A. Cancer genome-sequencing study design. Nat. Rev. Genet. 14, 321–332 (2013).
Wheeler, D.A. & Wang, L. From human genome to cancer genome: The first decade. Genome Res. 23, 1054–1062 (2013).
Ley, T.J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, 66–72 (2008).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Espina, V. et al. Laser-capture microdissection. Nat. Protoc. 1, 586–603 (2006).
Baslan, T. et al. Genome-wide copy number analysis of single cells. Nat. Protoc. 7, 1024–1041 (2012).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Campbell, P.J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010).
Hahn, S.A. et al. Allelotype of pancreatic adenocarcinoma using xenograft enrichment. Cancer Res. 55, 4670–4675 (1995).
Peña-Llopis, S. et al. BAP1 loss defines a new class of renal cell carcinoma. Nat. Genet. 44, 751–759 (2012).
Sivanand, S. et al. A validated tumorgraft model reveals activity of dovitinib against renal cell carcinoma. Sci. Transl. Med. 4, 137ra75 (2012).
Sato, Y. et al. Integrated molecular analysis of clear-cell renal cell carcinoma. Nat. Genet. 45, 860–867 (2013).
Hakimi, A.A. et al. Adverse outcomes in clear cell renal cell carcinoma with mutations of 3p21 epigenetic regulators BAP1 and SETD2: a report by MSKCC and the KIRC TCGA research network. Clin. Cancer Res. 19, 3259–3267 (2013).
Creighton, C.J. et al. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49 (2013).
Dalgliesh, G.L. et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463, 360–363 (2010).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Hakimi, A.A. et al. Clinical and pathologic impact of select chromatin-modulating tumor suppressors in clear cell renal cell carcinoma. Eur. Urol. 63, 848–854 (2013).
Guo, G. et al. Frequent mutations of genes encoding ubiquitin-mediated proteolysis pathway components in clear cell renal cell carcinoma. Nat. Genet. 44, 17–19 (2012).
Nickerson, M.L. et al. Improved identification of von Hippel-Lindau gene alterations in clear cell renal tumors. Clin. Cancer Res. 14, 4726–4734 (2008).
Peña-Llopis, S., Christie, A., Xie, X.J. & Brugarolas, J. Cooperation and antagonism among cancer genes: the renal cancer paradigm. Cancer Res. 73, 4173–4179 (2013).
Boom, R. et al. Rapid and simple method for purification of nucleic acids. J. Clin. Microbiol. 28, 495–503 (1990).
Gross-Bellard, M., Oudet, P. & Chambon, P. Isolation of high-molecular-weight DNA from mammalian cells. Eur. J. Biochem. 36, 32–38 (1973).
Strauss, W.M. Preparation of genomic DNA from mammalian tissue. Curr. Protoc. Mol. Biol. 42, 2.2.1–2.2.3 (2001).
Chomczynski, P. & Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156–159 (1987).
Chomczynski, P. & Sacchi, N. The single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction: twenty-something years on. Nat. Protoc. 1, 581–585 (2006).
Chomczynski, P. A reagent for the single-step simultaneous isolation of RNA, DNA and proteins from cell and tissue samples. Biotechniques 15, 532–534, 536-537 (1993).
Eddy, S.R. Non-coding RNA genes and the modern RNA world. Nat. Rev. Genet. 2, 919–929 (2001).
Siomi, M.C., Sato, K., Pezic, D. & Aravin, A.A. PIWI-interacting small RNAs: the vanguard of genome defence. Nat. Rev. Mol. Cell Biol. 12, 246–258 (2011).
Vega-Rubin-de-Celis, S. et al. Structural analysis and functional implications of the negative mTORC1 regulator REDD1. Biochemistry 49, 2491–2501 (2010).
Kucejova, B. et al. Interplay between pVHL and mTORC1 pathways in clear-cell renal cell carcinoma. Mol. Cancer Res. 9, 1255–1265 (2011).
Peña-Llopis, S. et al. Regulation of TFEB and V-ATPases by mTORC1. EMBO J. 30, 3242–3258 (2011).
Clark, S.J., Statham, A., Stirzaker, C., Molloy, P.L. & Frommer, M. DNA methylation: bisulphite modification and analysis. Nat. Protoc. 1, 2353–2364 (2006).
Taiwo, O. et al. Methylome analysis using MeDIP-seq with low DNA concentrations. Nat. Protoc. 7, 617–636 (2012).
Southern, E. Southern blotting. Nat. Protoc. 1, 518–525 (2006).
Wilhelm, B.T., Marguerat, S., Goodhead, I. & Bahler, J. Defining transcribed regions using RNA-seq. Nat. Protoc. 5, 255–266 (2010).
Streit, S., Michalski, C.W., Erkan, M., Kleeff, J. & Friess, H. Northern blot analysis for detection and quantification of RNA in pancreatic cancer cells and tissues. Nat. Protoc. 4, 37–43 (2009).
Pall, G.S. & Hamilton, A.J. Improved northern blot method for enhanced detection of small RNA. Nat. Protoc. 3, 1077–1084 (2008).
Liu, C.G., Calin, G.A., Volinia, S. & Croce, C.M. MicroRNA expression profiling using microarrays. Nat. Protoc. 3, 563–578 (2008).
Carrette, O., Burkhard, P.R., Sanchez, J.C. & Hochstrasser, D.F. State-of-the-art two-dimensional gel electrophoresis: a key tool of proteomics research. Nat. Protoc. 1, 812–823 (2006).
Shiio, Y. & Aebersold, R. Quantitative proteome analysis using isotope-coded affinity tags and mass spectrometry. Nat. Protoc. 1, 139–145 (2006).
de Planell-Saguer, M., Rodicio, M.C. & Mourelatos, Z. Rapid in situ codetection of noncoding RNAs and proteins in cells and formalin-fixed paraffin-embedded tissue sections without protease treatment. Nat. Protoc. 5, 1061–1073 (2010).
Acknowledgements
We thank S. Vega-Rubín-de-Celis and A. Pavía-Jiménez for critically reviewing the manuscript. This work was supported by a Postdoctoral Fellowship of Excellence from Generalitat Valenciana (Spain) (no. BPOSTDOC06/004) to S.P.-L. and the following grants to J.B.: a grant from the Cancer Prevention and Research Institute of Texas (no. RP101075) and an American Cancer Society Research Scholar grant (no. 55927). J.B. is a Virginia Murchison Linthicum Endowed Scholar in Medical Research. The tissue management shared resource was supported in part by the US National Cancer Institute (no. 1P30CA142543).
Author information
Authors and Affiliations
Contributions
S.P.-L. developed the protocols and performed all the experiments. S.P.-L. and J.B. designed the experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Peña-Llopis, S., Brugarolas, J. Simultaneous isolation of high-quality DNA, RNA, miRNA and proteins from tissues for genomic applications. Nat Protoc 8, 2240–2255 (2013). https://doi.org/10.1038/nprot.2013.141
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.141
This article is cited by
-
Germline and sporadic mTOR pathway mutations in low-grade oncocytic tumor of the kidney
Modern Pathology (2022)
-
The analysis of N6-methyladenosine regulators impacting the immune infiltration in clear cell renal cell carcinoma
Medical Oncology (2022)
-
Determinants of renal cell carcinoma invasion and metastatic competence
Nature Communications (2021)
-
Cold seawater induces early sexual developmental stages in the BPG axis of European eel males
BMC Genomics (2019)
-
Dual DNA-protein extraction from human archeological remains
Archaeological and Anthropological Sciences (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.