Abstract
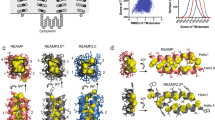
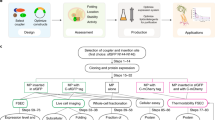
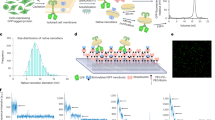
Finding a way to determine the structures of integral membrane proteins using solution nuclear magnetic resonance (NMR) spectroscopy has proved to be challenging. A residual-dipolar-coupling–based refinement approach can be used to resolve the structure of membrane proteins up to 40 kDa in size, but to do this you need a weak-alignment medium that is detergent-resistant and it has thus far been difficult to obtain such a medium suitable for weak alignment of membrane proteins. We describe here a protocol for robust, large-scale synthesis of detergent-resistant DNA nanotubes that can be assembled into dilute liquid crystals for application as weak-alignment media in solution NMR structure determination of membrane proteins in detergent micelles. The DNA nanotubes are heterodimers of 400-nm-long six-helix bundles, each self-assembled from a M13-based p7308 scaffold strand and >170 short oligonucleotide staple strands. Compatibility with proteins bearing considerable positive charge as well as modulation of molecular alignment, toward collection of linearly independent restraints, can be introduced by reducing the negative charge of DNA nanotubes using counter ions and small DNA-binding molecules. This detergent-resistant liquid-crystal medium offers a number of properties conducive for membrane protein alignment, including high-yield production, thermal stability, buffer compatibility and structural programmability. Production of sufficient nanotubes for four or five NMR experiments can be completed in 1 week by a single individual.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Seeman, N.C. Nucleic acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982).
Seeman, N.C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Rothemund, P.W.K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Douglas, S.M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Dietz, H., Douglas, S.M. & Shih, W.M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Douglas, S.M., Chou, J.J. & Shih, W.M. DNA-nanotube-induced alignment of membrane proteins for NMR structure determination. Proc. Natl. Acad. Sci. USA 104, 6644–6648 (2007).
Berardi, M.J., Shih, W.M., Harrison, S.C. & Chou, J.J. Mitochondrial uncoupling protein 2 structure determined by NMR molecular fragment searching. Nature 476, 109–113 (2011).
Wallin, E. & Heijne von, G. Genome-wide analysis of integral membrane proteins from eubacterial, archaean, and eukaryotic organisms. Protein Sci. 7, 1029–1038 (1998).
Goffeau, A. et al. Life with 6000 genes. Science 274 546, 563–567 (1996).
Landry, Y. & Gies, J. Drugs and their molecular targets: an updated overview. Fundam. Clin. Pharmacol. 22, 1–18 (2008).
Myers, J.K., Beihoffer, L.A. & Sanders, C.R. Phenotology of disease-linked proteins. Hum. Mutat. 25, 90–97 (2005).
Groom, C.R. & Hopkins, A.L. Protein kinase drugs–optimism doesn't wait on facts. Drug Discov. Today 7, 801–802 (2002).
Schnell, J.R. & Chou, J.J. Structure and mechanism of the M2 proton channel of influenza A virus. Nature 451, 591–595 (2008).
Hiller, S. et al. Solution structure of the integral human membrane protein VDAC-1 in detergent micelles. Science 321, 1206–1210 (2008).
Wang, J., Pielak, R.M., McClintock, M.A. & Chou, J.J. Solution structure and functional analysis of the influenza B proton channel. Nat. Struct. Mol. Biol. 16, 1267–1271 (2009).
Zhou, Y. et al. NMR solution structure of the integral membrane enzyme DsbB: functional insights into DsbB-catalyzed disulfide bond formation. Mol. Cell 31, 896–908 (2008).
Van Horn, W.D. et al. Solution nuclear magnetic resonance structure of membrane-integral diacylglycerol kinase. Science 324, 1726–1729 (2009).
Gautier, A., Mott, H.R., Bostock, M.J., Kirkpatrick, J.P. & Nietlispach, D. Structure determination of the seven-helix transmembrane receptor sensory rhodopsin II by solution NMR spectroscopy. Nat. Struct. Mol. Biol. 17, 768–774 (2010).
Kang, C. & Li, Q. Solution NMR study of integral membrane proteins. Curr. Opin. Chem. Biol. 15, 560–569 (2011).
Pervushin, K., Riek, R., Wider, G. & Wüthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl. Acad. Sci. USA 94, 12366–12371 (1997).
Prestegard, J.H. New techniques in structural NMR–anisotropic interactions. Nat. Struct. Biol. 5 (suppl.), 517–522 (1998).
Tjandra, N. & Bax, A. Direct measurement of distances and angles in biomolecules by NMR in a dilute liquid crystalline medium. Science 278, 1111–1114 (1997).
Bax, A., Kontaxis, G. & Tjandra, N. Dipolar couplings in macromolecular structure determination. Meth. Enzymol. 339, 127–174 (2001).
Tolman, J.R., Flanagan, J.M., Kennedy, M.A. & Prestegard, J.H. Nuclear magnetic dipole interactions in field-oriented proteins: information for structure determination in solution. Proc. Natl. Acad. Sci. USA 92, 9279–9283 (1995).
Hansen, M.R., Mueller, L. & Pardi, A. Tunable alignment of macromolecules by filamentous phage yields dipolar coupling interactions. Nat. Struct. Biol. 5, 1065–1074 (1998).
Rückert, M. & Otting, G. Alignment of biological macromolecules in novel nonionic liquid crystalline media for NMR experiments. J. Am. Chem. Soc. 122, 7793–7797 (2000).
Prosser, R.S., Losonczi, J.A. & Shiyanovskaya, I.V. Use of a novel aqueous liquid crystalline medium for high-resolution NMR of macromolecules in solution. J. Am. Chem. Soc. 120, 11010–11011 (1998).
Fleming, K., Gray, D., Prasannan, S. & Matthews, S. Cellulose crystallites: a new and robust liquid crystalline medium for the measurement of residual dipolar couplings. J. Am. Chem. Soc. 122, 5224–5225 (2000).
Tycko, R., Blanco, F.J. & Ishii, Y. Alignment of biopolymers in strained gels: a new way to create detectable dipoledipole couplings in high-resolution biomolecular NMR. J. Am. Chem. Soc. 122, 9340–9341 (2000).
Jones, D.H. & Opella, S.J. Weak alignment of membrane proteins in stressed polyacrylamide gels. J. Magn. Reson. 171, 258–269 (2004).
Oxenoid, K. & Chou, J.J. The structure of phospholamban pentamer reveals a channel-like architecture in membranes. Proc. Natl. Acad. Sci. USA 102, 10870–10875 (2005).
Chou, J.J., Kaufman, J.D., Stahl, S.J., Wingfield, P.T. & Bax, A. Micelle-induced curvature in a water-insoluble HIV-1 Env peptide revealed by NMR dipolar coupling measurement in stretched polyacrylamide gel. J. Am. Chem. Soc. 124, 2450–2451 (2002).
Chou, J.J., Gaemers, S., Howder, B., Louis, J.M. & Bax, A. A simple apparatus for generating stretched polyacrylamide gels, yielding uniform alignment of proteins and detergent micelles. J. Biomol. NMR 21, 377–382 (2001).
Park, S.H., Son, W.S., Mukhopadhyay, R., Valafar, H. & Opella, S.J. J. Am. Chem. Soc. 131, 14140–14141 (2009).
Ma, J., Goldberg, G.I. & Tjandra, N. Weak alignment of biomacromolecules in collagen gels: an alternative way to yield residual dipolar couplings for NMR measurements. J. Am. Chem. Soc. 130, 16148–16149 (2008).
Lorieau, J., Yao, L. & Bax, A. Liquid crystalline phase of G-tetrad DNA for NMR study of detergent-solubilized proteins. J. Am. Chem. Soc. 130, 7536–7537 (2008).
Marvin, D.A. Filamentous phage structure, infection and assembly. Curr. Opin. Struct. Biol. 8, 150–158 (1998).
Call, M.E., Wucherpfennig, K.W. & Chou, J.J. The structural basis for intramembrane assembly of an activating immunoreceptor complex. Nat. Immunol. 11, 1023–1029 (2010).
Zweckstetter, M. NMR: prediction of molecular alignment from structure using the PALES software. Nat. Protoc. 3, 679–690 (2008).
Zweckstetter, M. & Bax, A. Characterization of molecular alignment in aqueous suspensions of Pf1 bacteriophage. J. Biomol. NMR 20, 365–377 (2001).
Flynn, P.F., Mattiello, D.L., Hill, H.D.W. & Wand, A.J. Optimal use of cryogenic probe technology in NMR studies of proteins. J. Am. Chem. Soc. 122, 4823–4824 (2000).
Kelly, A.E., Ou, H.D., Withers, R. & Dötsch, V. Low-conductivity buffers for high-sensitivity NMR measurements. J. Am. Chem. Soc. 124, 12013–12019 (2002).
Robosky, L.C., Reily, M.D. & Avizonis, D. Improving NMR sensitivity by use of salt-tolerant cryogenically cooled probes. Anal. Bioanal. Chem. 387, 529–532 (2007).
Voehler, M.W., Collier, G., Young, J.K., Stone, M.P. & Germann, M.W. Performance of cryogenic probes as a function of ionic strength and sample tube geometry. J. Magn. Reson. 183, 102–109 (2006).
Zweckstetter, M., Hummer, G. & Bax, A. Prediction of charge-induced molecular alignment of biomolecules dissolved in dilute liquid-crystalline phases. Biophys. J. 86, 3444–3460 (2004).
Neto, B.A.D. & Lapis, A.A.M. Recent developments in the chemistry of deoxyribonucleic acid (DNA) intercalators: principles, design, synthesis, applications and trends. Molecules 14, 1725–1746 (2009).
Nafisi, S., Saboury, A.A., Keramat, N., Neault, J. & Tajmir-Riahi, H. Stability and structural features of DNA intercalation with ethidium bromide, acridine orange and methylene blue. J. Mol. Struct. 827, 35–43 (2007).
Furse, K.E. & Corcelli, S.A. The dynamics of water at DNA interfaces: computational studies of Hoechst 33258 bound to DNA. J. Am. Chem. Soc. 130, 13103–13109 (2008).
Banerjee, D. & Pal, S.K. Ultrafast charge transfer and solvation of DNA minor groove binder: Hoechst 33258 in restricted environments. Chem. Phys. Lett. 432, 257–262 (2006).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Zweckstetter, M. & Bax, A. Prediction of sterically induced alignment in a dilute liquid crystalline phase: aid to protein structure determination by NMR. J. Am. Chem. Soc. 122, 3791–3792 (2000).
Cornilescu, G., Marquardt, J.L., Ottiger, M. & Bax, A. Validation of protein structure from anisotropic carbonyl chemical shifts in a dilute liquid crystalline phase. J. Am. Chem. Soc. 120, 6836–6837 (1998).
Acknowledgements
We thank I. Ayala and J. Boisbouvier (CNRS, Institut de Biologie Structurale) for the gift of the ubiquitin vector and R. Sounier (Harvard Medical School) for helpful discussions. This work was supported by the US National Institutes of Health (NIH) grants 1U54GM094608 to J.J.C., and 1DP2OD004641 and 1U54GM094608 to W.M.S.
Author information
Authors and Affiliations
Contributions
B.G. and M.A.M. used, developed and troubleshot the technique. B.G. optimized procedures for NMR experiments, data processing and analysis. W.M.S. supervised the projects and J.J.C. participated in technology design and discussions. B.G. and W.M.S. wrote the manuscript, and J.J.C and M.A.M. revised the paper.
Corresponding author
Ethics declarations
Competing interests
J.J.C. and W.M.S. declare competing financial interests. A patent (US Patent application no. 13/090892) entitled “Nucleic-acid-nanotube liquid crystals and use for NMR structure determination of detergent-solubilized membrane proteins” was filed in 2007 on behalf of the Dana-Farber Cancer Institute and Harvard Medical School by Edwards Angell Palmer & Dodge LLP, listing J.J.C. and W.M.S. as two of the co-inventors.
Supplementary information
Supplementary Table 1
DNA oligonucleotide staple sequences (PDF 436 kb)
Rights and permissions
About this article
Cite this article
Bellot, G., McClintock, M., Chou, J. et al. DNA nanotubes for NMR structure determination of membrane proteins. Nat Protoc 8, 755–770 (2013). https://doi.org/10.1038/nprot.2013.037
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.037
This article is cited by
-
Design and operation of reconfigurable two-dimensional DNA molecular arrays
Nature Protocols (2018)
-
Structure refinement and membrane positioning of selectively labeled OmpX in phospholipid nanodiscs
Journal of Biomolecular NMR (2015)
-
DNA origami–based standards for quantitative fluorescence microscopy
Nature Protocols (2014)
-
Amphipols for Each Season
The Journal of Membrane Biology (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.