Abstract
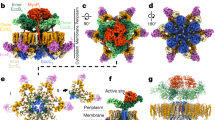
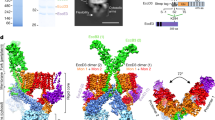
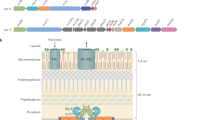
Mycobacteria are characterized by their impermeable outer membrane, which is rich in mycolic acids1. To transport substrates across this complex cell envelope, mycobacteria rely on type VII (also known as ESX) secretion systems2. In Mycobacterium tuberculosis, these ESX systems are essential for growth and full virulence and therefore represent an attractive target for anti-tuberculosis drugs3. However, the molecular details underlying type VII secretion are largely unknown, due to a lack of structural information. Here, we report the molecular architecture of the ESX-5 membrane complex from Mycobacterium xenopi determined at 13 Å resolution by electron microscopy. The four core proteins of the ESX-5 complex (EccB5, EccC5, EccD5 and EccE5) assemble with equimolar stoichiometry into an oligomeric assembly that displays six-fold symmetry. This membrane-associated complex seems to be embedded exclusively in the inner membrane, which indicates that additional components are required to translocate substrates across the mycobacterial outer membrane. Furthermore, the extended cytosolic domains of the EccC ATPase, which interact with secretion effectors, are highly flexible, suggesting an as yet unseen mode of substrate interaction. Comparison of our results with known structures of other bacterial secretion systems demonstrates that the architecture of type VII secretion system is fundamentally different, suggesting an alternative secretion mechanism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hoffmann, C., Leis, A., Niederweis, M., Plitzko, J. M. & Engelhardt, H. Disclosure of the mycobacterial outer membrane: cryo-electron tomography and vitreous sections reveal the lipid bilayer structure. Proc. Natl Acad. Sci. USA 105, 3963–3967 (2008).
Houben, E. N. G., Korotkov, K. V. & Bitter, W. Take five—type VII secretion systems of mycobacteria. Biochim. Biophys. Acta 1843, 1707–1716 (2014).
Bitter, W. & Kuijl, C. Targeting bacterial virulence. The coming out of type VII secretion inhibitors. Cell Host Microbe 16, 430–432 (2014).
Gröschel, M. I., Sayes, F., Simeone, R., Majlessi, L. & Brosch, R. ESX secretion systems mycobacterial evolution to counter host immunity. Nat. Rev. Microbiol. 14, 677–691 (2016).
Pym, A. S., Brodin, P., Brosch, R., Huerre, M. & Cole, S. T. Loss of RD1 contributed to the attenuation of the live tuberculosis vaccines Mycobacterium bovis BCG and Mycobacterium microti. Mol. Microbiol. 46, 709–717 (2002).
Stanley, S. A., Raghavan, S., Hwang, W. W. & Cox, J. S. Acute infection and macrophage subversion by Mycobacterium tuberculosis require a specialized secretion system. Proc. Natl Acad. Sci. USA 100, 13001–13006 (2003).
Hsu, T. et al. The primary mechanism of attenuation of bacillus Calmette-Guérin is a loss of secreted lytic function required for invasion of lung interstitial tissue. Proc. Natl Acad. Sci. USA 100, 12420–12425 (2003).
Ates, L. S. et al. Essential role of the ESX-5 secretion system in outer membrane permeability of pathogenic mycobacteria. PLoS Genet. 11, e1005190 (2015).
Fishbein, S., van Wyk, N., Warren, R. M. & Sampson, S. L. Phylogeny to function PE/PPE protein evolution and impact on Mycobacterium tuberculosis pathogenicity. Mol. Microbiol. 96, 901–916 (2015).
Houben, E. N. G. et al. Composition of the type VII secretion system membrane complex. Mol. Microbiol. 86, 472–484 (2012).
Sani, M. et al. Direct visualization by cryo-EM of the mycobacterial capsular layer: a labile structure containing ESX-1-secreted proteins. PLoS Pathog. 6, e1000794 (2010).
Rosenberg, O. S. et al. Substrates control multimerization and activation of the multi-domain ATPase motor of type VII secretion. Cell 161, 501–512 (2015).
Daleke, M. H. et al. General secretion signal for the mycobacterial type VII secretion pathway. Proc. Natl Acad. Sci. USA 109, 11342–11347 (2012).
Korotkova, N. et al. Structure of the Mycobacterium tuberculosis type VII secretion system chaperone EspG5 in complex with PE25–PPE41 dimer. Mol. Microbiol. 94, 367–382 (2014).
Houben, E. N. G. et al. Composition of the type VII secretion system membrane complex. Mol. Microbiol. 86, 472–484 (2012).
Poulsen, C., Panjikar, S., Holton, S. J., Wilmanns, M. & Song, Y.-H. WXG100 protein superfamily consists of three subfamilies and exhibits an α-helical C-terminal conserved residue pattern. PLoS ONE 9, e89313 (2014).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinform. 47, 5.6.1–5.6.32 (2014).
Zhang, X.-L. et al. Core component EccB1 of the Mycobacterium tuberculosis type VII secretion system is a periplasmic ATPase. FASEB J. 29, 4804–4814 (2015).
Pintilie, G. D., Zhang, J., Goddard, T. D., Chiu, W. & Gossard, D. C. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 170, 427–438 (2010).
Bitter, W. et al. Systematic genetic nomenclature for type VII secretion systems. PLoS Pathog. 5, e1000507 (2009).
Lou, Y., Rybniker, J., Sala, C. & Cole, S. T. EspC forms a filamentous structure in the cell envelope of Mycobacterium tuberculosis and impacts ESX-1 secretion. Mol. Microbiol. 103, 1–39 (2016).
Peña, A. et al. The hexameric structure of a conjugative VirB4 protein ATPase provides new insights for a functional and phylogenetic relationship with DNA translocases. J. Biol. Chem. 287, 39925–39932 (2012).
van Winden, V. J. C. et al. Mycosins are required for the stabilization of the ESX-1 and ESX-5 type VII secretion membrane complexes. mBio 7, e01471–16 (2016).
Labidi, A., Mardis, E., Roe, B. A. & Wallace, R. J. Cloning and DNA sequence of the Mycobacterium fortuitum var fortuitum plasmid pAL5000. Plasmid 27, 130–140 (1992).
Zhang, Y., Werling, U. & Edelmann, W. SLiCE: a novel bacterial cell extract-based DNA cloning method. Nucleic Acids Res. 40, e55 (2012).
Noens, E. E. et al. Improved mycobacterial protein production using a Mycobacterium smegmatis groEL1ΔC expression strain. BMC Microbiol. 11, 27 (2011).
Hughes, C. S. et al. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 10, 757 (2014).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Cox, J. & Mann, M. Maxquant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R. & Wolfson, H. J. Patchdock and SymmDock servers for rigid and symmetric docking. Nucleic Acids Res. 33, W363–W367 (2005).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J. & Svergun, D. I. PRIMUS: a Windows PC-based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003).
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Tria, G., Mertens, H. D. T., Kachala, M. & Svergun, D. I. Advanced ensemble modelling of flexible macromolecules using X-ray solution scattering. IUCrJ. 2, 207–217 (2015).
Bernado, P., Mylonas, E., Petoukhov, M. V., Blackledge, M. & Svergun, D. I. Structural characterization of flexible proteins using small-angle X-ray scattering. J. Am. Chem. Soc. 129, 5656–5664 (2007).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Du, D. et al. Structure of the AcrAB-TolC multidrug efflux pump. Nature 509, 512–515 (2014).
Schraidt, O. & Marlovits, T. C. Three-dimensional model of Salmonella's needle complex at subnanometer resolution. Science 331, 1192–1195 (2011).
Low, H. H. et al. Structure of a type IV secretion system. Nature 508, 550–553 (2014).
Durand, E. et al. Biogenesis and structure of a type VI secretion membrane core complex. Nature 523, 555–560 (2015).
Acknowledgements
This work was generously supported through the Behörde für Wissenschaft, Forschung und Gleichstellung of the city of Hamburg (to T.C.M.) and funded by a VIDI grant from the Netherlands Organization of Scientific Research (NWO) (to E.N.G.H. and C.M.B.). K.S.H.B. was funded by a postdoctoral fellowship from the EMBL Interdisciplinary Postdoc Program (EIPOD) under Marie Curie COFUND actions. The authors thank the EMBL P12 beamline and synchrotron facilities at PETRA III (DESY, Hamburg). The authors thank M. Jeske for assistance with the ATPase assays (EMBL, Heidelberg), S. Bhushan (Nanyang Technological University, Singapore) for initial EM analysis of ESX-5 particles and F. Schluenzen for DESY high-performance cluster support. Samples were recorded at the EM facility of the Vienna Biocenter Core Facilities GmbH (VBCF), Austria. The authors thank S.J. Ludtke (Baylor College of Medicine) for comments on resolution estimation and A. Jakobi and C. Sachse (EMBL, Heidelberg) for providing feedback on the manuscript.
Author information
Authors and Affiliations
Contributions
A.H.A.P., E.N.G.H., W.B., T.C.M. and M.W. supervised and supported the project. K.S.H.B., A.H.A.P., E.N.G.H., C.M.B. and L.C. designed experiments. E.N.G.H., R.U., C.M.B. and K.S.H.B. generated the constructs. K.S.H.B., C.M.B. and E.N.G.H. purified the ESX-5 complexes for EM analysis. K.S.H.B. performed gold labelling and ATPase assays. C.M.B. performed secretion analysis and disulfide bond characterization. L.C. and J.M. performed electron microscopy. L.C. collected and processed the negative stain electron microscopy data together with W.L. K.S.H.B. and H.D.T.M. collected and analysed SAXS data with the support of D.I.S. M.M.S. and M.R. conducted the mass-spectrometry analysis. K.S.H.B., A.H.A.P., E.N.G.H., C.M.B., W.B., T.C.M., L.C. and M.W. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–9, Supplementary Tables 1–3, Supplementary References. (PDF 3399 kb)
Rights and permissions
About this article
Cite this article
Beckham, K., Ciccarelli, L., Bunduc, C. et al. Structure of the mycobacterial ESX-5 type VII secretion system membrane complex by single-particle analysis. Nat Microbiol 2, 17047 (2017). https://doi.org/10.1038/nmicrobiol.2017.47
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.47
This article is cited by
-
Type VII secretion systems: structure, functions and transport models
Nature Reviews Microbiology (2021)
-
Toxin secretion and trafficking by Mycobacterium tuberculosis
Nature Communications (2021)
-
Structure and dynamics of a mycobacterial type VII secretion system
Nature (2021)
-
Non-tuberculous mycobacteria and the rise of Mycobacterium abscessus
Nature Reviews Microbiology (2020)
-
Structural insights into substrate recognition by the type VII secretion system
Protein & Cell (2020)