Abstract
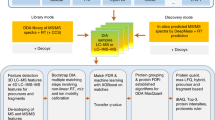
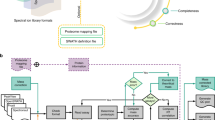
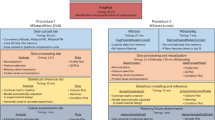
Liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) is the main method for high-throughput identification and quantification of peptides and inferred proteins. Within this field, data-independent acquisition (DIA) combined with peptide-centric scoring, as exemplified by the technique SWATH-MS, has emerged as a scalable method to achieve deep and consistent proteome coverage across large-scale data sets. We demonstrate that statistical concepts developed for discovery proteomics based on spectrum-centric scoring can be adapted to large-scale DIA experiments that have been analyzed with peptide-centric scoring strategies, and we provide guidance on their application. We show that optimal tradeoffs between sensitivity and specificity require careful considerations of the relationship between proteins in the samples and proteins represented in the spectral library. We propose the application of a global analyte constraint to prevent the accumulation of false positives across large-scale data sets. Furthermore, to increase the quality and reproducibility of published proteomic results, well-established confidence criteria should be reported for the detected peptide queries, peptides and inferred proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Domon, B. & Aebersold, R. Options and considerations when selecting a quantitative proteomics strategy. Nat. Biotechnol. 28, 710–721 (2010).
Chapman, J.D., Goodlett, D.R. & Masselon, C.D. Multiplexed and data-independent tandem mass spectrometry for global proteome profiling. Mass Spectrom. Rev. 33, 452–470 (2014).
Gillet, L.C., Leitner, A. & Aebersold, R. Mass spectrometry applied to bottom-up proteomics: entering the high-throughput era for hypothesis testing. Annu. Rev. Anal. Chem. (Palo Alto Calif.) 9, 449–472 (2016).
Ting, Y.S. et al. Peptide-centric proteome analysis: an alternative strategy for the analysis of tandem mass spectrometry data. Mol. Cell. Proteomics 14, 2301–2307 (2015).
Silva, J.C. et al. Quantitative proteomic analysis by accurate mass-retention-time pairs. Anal. Chem. 77, 2187–2200 (2005).
Tsou, C.-C. et al. DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics. Nat. Methods 12, 258–264 (2015).
Wang, J. et al. MSPLIT-DIA: sensitive peptide identification for data-independent acquisition. Nat. Methods 12, 1106–1108 (2015).
Li, Y. et al. Group-DIA: analyzing multiple data-independent acquisition mass spectrometry data files. Nat. Methods 12, 1105–1106 (2015).
Gillet, L.C. et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol. Cell. Proteomics 11, O111.016717 (2012).
Röst, H.L. et al. OpenSWATH enables automated, targeted analysis of data-independent acquisition MS data. Nat. Biotechnol. 32, 219–223 (2014).
Teleman, J. et al. DIANA—algorithmic improvements for analysis of data-independent acquisition MS data. Bioinformatics 31, 555–562 (2015).
MacLean, B. et al. Skyline: an open-source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Bruderer, R. et al. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol. Cell. Proteomics 14, 1400–1410 (2015).
Carr, S.A. et al. Targeted peptide measurements in biology and medicine: best practices for mass-spectrometry-based assay development using a fit-for-purpose approach. Mol. Cell. Proteomics 13, 907–917 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Stat. Methodol. 57, 289–300 (1995).
Keller, A., Nesvizhskii, A.I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Choi, H. & Nesvizhskii, A.I. Semi-supervised model-based validation of peptide identifications in mass-spectrometry-based proteomics. J. Proteome Res. 7, 254–265 (2008).
Käll, L., Storey, J.D., MacCoss, M.J. & Noble, W.S. Posterior error probabilities and false discovery rates: two sides of the same coin. J. Proteome Res. 7, 40–44 (2008).
Genovese, C. & Wasserman, L. Operating characteristics and extensions of the false discovery rate procedure. J. R. Stat. Soc. B Stat. Methodol. 64, 499–517 (2002).
Iyer, V. & Sarkar, S. An adaptive single-step FDR procedure with applications to DNA microarray analysis. Biom. J. 49, 127–135 (2007).
Storey, J.D. The positive false discovery rate: a Bayesian interpretation and the q-value. Ann. Stat. 31, 2013–2035 (2003).
Nesvizhskii, A.I. A survey of computational methods and error-rate estimation procedures for peptide and protein identification in shotgun proteomics. J. Proteomics 73, 2092–2123 (2010).
Käll, L., Canterbury, J.D., Weston, J., Noble, W.S. & MacCoss, M.J. Semi-supervised learning for peptide identification from shotgun proteomics data sets. Nat. Methods 4, 923–925 (2007).
Serang, O. & Noble, W. A review of statistical methods for protein identification using tandem mass spectrometry. Stat. Interface 5, 3–20 (2012).
The, M., Tasnim, A. & Käll, L. How to talk about protein-level false discovery rates in shotgun proteomics. Proteomics 16, 2461–2469 (2016).
Shteynberg, D. et al. iProphet: multilevel integrative analysis of shotgun proteomic data improves peptide and protein identification rates and error estimates. Mol. Cell. Proteomics 10, M111.007690 (2011).
Reiter, L. et al. Protein identification false discovery rates for very large proteomics data sets generated by tandem mass spectrometry. Mol. Cell. Proteomics 8, 2405–2417 (2009).
Savitski, M.M., Wilhelm, M., Hahne, H., Kuster, B. & Bantscheff, M. A scalable approach for protein false discovery rate estimation in large proteomic data sets. Mol. Cell. Proteomics 14, 2394–2404 (2015).
The, M., MacCoss, M.J., Noble, W.S. & Käll, L. Fast and accurate protein false discovery rates on large-scale proteomics data sets with Percolator 3.0. J. Am. Soc. Mass Spectrom. 27, 1719–1727 (2016).
Choi, H., Ghosh, D. & Nesvizhskii, A.I. Statistical validation of peptide identifications in large-scale proteomics using the target-decoy database search strategy and flexible mixture modeling. J. Proteome Res. 7, 286–292 (2008).
Ahrens, C.H., Brunner, E., Qeli, E., Basler, K. & Aebersold, R. Generating and navigating proteome maps using mass spectrometry. Nat. Rev. Mol. Cell Biol. 11, 789–801 (2010).
Reiter, L. et al. mProphet: automated data processing and statistical validation for large-scale SRM experiments. Nat. Methods 8, 430–435 (2011).
Karlsson, C., Malmström, L., Aebersold, R. & Malmström, J. Proteome-wide selected reaction monitoring assays for the human pathogen Streptococcus pyogenes. Nat. Commun. 3, 1301 (2012).
Schubert, O.T. et al. The Mtb proteome library: a resource of assays to quantify the complete proteome of Mycobacterium tuberculosis. Cell Host Microbe 13, 602–612 (2013).
Picotti, P. et al. A complete mass spectrometric map of the yeast proteome applied to quantitative trait analysis. Nature 494, 266–270 (2013).
Rosenberger, G. et al. A repository of assays to quantify 10,000 human proteins by SWATH-MS. Sci. Data 1, 140031 (2014).
Collins, B.C. et al. Multi-laboratory assessment of reproducibility, qualitative and quantitative performance of SWATH–mass spectrometry. Nat. Commun. 8, DOI: 10.1038/s41467-017-00249-5 (2017).
Liu, Y. et al. Quantitative variability of 342 plasma proteins in a human twin population. Mol. Syst. Biol. 11, 786 (2015).
Selevsek, N. et al. Reproducible and consistent quantification of the Saccharomyces cerevisiae proteome by SWATH-MS. Mol. Cell. Proteomics 14, 739–749 (2015).
Guo, T. et al. Rapid mass spectrometric conversion of tissue biopsy samples into permanent quantitative digital proteome maps. Nat. Med. 21, 407–413 (2015).
Schubert, O.T. et al. Absolute proteome composition and dynamics during dormancy and resuscitation of Mycobacterium tuberculosis. Cell Host Microbe 18, 96–108 (2015).
Schubert, O.T. et al. Building high-quality assay libraries for targeted analysis of SWATH-MS data. Nat. Protoc. 10, 426–441 (2015).
Storey, J.D. & Tibshirani, R. Statistical significance for genome-wide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Serang, O. & Käll, L. Solution to statistical challenges in proteomics is more statistics, not less. J. Proteome Res. 14, 4099–4103 (2015).
Blattmann, P., Heusel, M. & Aebersold, R. SWATH2stats: an R/Bioconductor package to process and convert quantitative SWATH-MS proteomics data for downstream analysis tools. PLoS One 11, e0153160 (2016).
Tsou, C.-C., Tsai, C.F., Teo, G.C., Chen, Y.J. & Nesvizhskii, A.I. Untargeted, spectral library-free analysis of data-independent acquisition proteomics data generated using Orbitrap mass spectrometers. Proteomics 16, 2257–2271 (2016).
Keller, A., Bader, S.L., Shteynberg, D., Hood, L. & Moritz, R.L. Automated validation of results and removal of fragment ion interferences in targeted analysis of data-independent acquisition mass spectrometry (MS) using SWATHProphet. Mol. Cell. Proteomics 14, 1411–1418 (2015).
Gupta, N. & Pevzner, P.A. False discovery rates of protein identifications: a strike against the two-peptide rule. J. Proteome Res. 8, 4173–4181 (2009).
Muntel, J. et al. Advancing urinary protein biomarker discovery by data-independent acquisition on a quadrupole-orbitrap mass spectrometer. J. Proteome Res. 14, 4752–4762 (2015).
Vizcaíno, J.A. et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 41, D1063–D1069 (2013).
Acknowledgements
Please note that M.H., C.L.H., Y.L., M.J.M., B.X.M., A.I.N., P.G.A.P., L.R., H.L.R., S.T. and Y.S.T. were added to the author list in alphabetical order. We thank the authors of the SWATH-MS interlaboratory study and of the human blood plasma data set for providing the data to conduct this study. We also thank the Scientific IT Support (ID SIS) and the high-performance computing (HPC) teams of ETH Zurich for support and maintenance of the computing infrastructure. M.H. was supported by a grant from the Institut Mérieux; A.I.N. was funded by the US National Institutes of Health (NIH; grant R01GM094231); H.L.R. was funded by the Swiss National Science Foundation (SNSF; grant P2EZP3 162268); B.C.C. was supported by a SNSF Ambizione grant (PZ00P3_161435); and R.A. was supported by ERC Proteomics v3.0 (AdG-233226 Proteomics v.3.0) and AdG-670821 Proteomics 4D), the PhosphonetX project of SystemsX.ch and the Swiss National Science Foundation (SNSF) grant 31003A_166435.
Author information
Authors and Affiliations
Contributions
G.R., I.B. and R.A. wrote the paper with feedback from all authors; G.R. and B.C.C. developed the methods; I.B. analyzed the data set; U.S. and G.R. implemented the PyProphet extension; M.H., C.L.H., Y.L., M.J.M., B.X.M., A.I.N., P.G.A.P., L.R., H.L.R., S.T. and Y.S.T. provided critical input on the project; and B.C.C. and R.A. designed and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
C.L.H. and S.T. are employees of SCIEX, which operates in the field of quantitative proteomics by data-independent acquisition covered by the article. M.J.M. is a paid consultant for Thermo Fisher Scientific, which operates in the field of quantitative proteomics by data-independent acquisition covered by the article. L.R. is employee of Biognosys AG, which operates in the field of quantitative proteomics by data-independent acquisition covered by the article. R.A. holds shares of Biognosys AG.
Integrated supplementary information
Supplementary Figure 1 Estimation of q-values at the peptide-query level, the peptide level and the protein level.
The peptide-query-level (left), peptide-level (middle) and protein-level (right) discriminant score density plots (a) and p-value histograms (b) for one DIA run of the SWATH-MS interlaboratory study that was analyzed with the combined human assay library (CAL) are shown. a) The distributions indicate a large (false target)/(total target) ratio (π0 ≍ 0.6) on the peptide-query level. The q-value estimation was adapted for peptide and protein level by using the best scoring peak group per peptide or protein across all samples for both targets and decoys. The (false target)/(total target) ratio decreases slightly on peptide level and more on protein level (π0 ≍ 0.5), compared to the peptide-query level. b) On the peptide-query level and the peptide level, the estimation of π0 is anticonservative, indicated by a lower density of p-values after the p-value threshold of λ=0.4. On the protein level, the estimation of π0 is more accurate with a consistent density of p-values.
Supplementary Figure 2 Influence of protein length on the peptide-query-level and protein-level q-value estimation.
a) Protein length distribution of all proteins in the combined human assay library (CAL), all proteins inferred at 1% peptide-query-level FDR in the global context of all 229 DIA runs of the SWATH-MS interlaboratory comparison study, and all proteins inferred at 1% global protein FDR respectively. b) Histogram of protein length distribution for the differently filtered protein subsets of the CAL. The distributions show that there is no bias for protein length when selecting the best peak group as proxy for protein-level q-value estimation.
Supplementary Figure 3 Decoy accumulation across multiple runs.
The number of cumulatively detected peak group decoys (a), peptide decoys (b) and protein decoys (c) is shown for 229 DIA runs of the SWATH-MS interlaboratory study data set.
Supplementary Figure 4 Analyte accumulation across multiple runs (5% FDR).
The number of cumulatively detected peak groups (a), peptides (b) and proteins (c) is shown for 229 DIA runs of the SWATH-MS interlaboratory study data set.
Supplementary Figure 5 Decoy accumulation across multiple runs (5% FDR).
The number of cumulatively detected peak group decoys (a), peptide decoys (b) and protein decoys (c) is shown for 229 DIA runs of the SWATH-MS interlaboratory study data set.
Supplementary Figure 6 Combined human and M. tuberculosis spectral library analysis.
a) The peptide-level discriminant score density of human targets, human decoys, M. tuberculosis (Mtb) targets, and Mtb decoys is shown for global analysis of the 229 DIA runs of the SWATH-MS interlaboratory study data set applying the combined human and Mtb spectral library. The Mtb targets and decoys show a similar distribution compared to the human decoys and the fraction of false human targets. The number of cumulatively detected peptides is shown for human targets (b), human decoys (c), Mtb targets (d), and Mtb decoys (e) from the combined human and Mtb spectral library with different error rate control strategies. The Mtb decoy to target ratio is 0.82, explaining the absolute higher number of the accumulated Mtb targets.
Supplementary Figure 7 Analyte accumulation across multiple runs in the plasma data set (1% FDR).
The number of cumulatively detected peak groups (a), peptides (b) and proteins (c) is shown for the 246 DIA runs of the plasma data set analyzed with the nonparametric model for q-value estimation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1 and Supplementary Notes 1–6. (PDF 1409 kb)
Rights and permissions
About this article
Cite this article
Rosenberger, G., Bludau, I., Schmitt, U. et al. Statistical control of peptide and protein error rates in large-scale targeted data-independent acquisition analyses. Nat Methods 14, 921–927 (2017). https://doi.org/10.1038/nmeth.4398
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4398
This article is cited by
-
AlphaPept: a modern and open framework for MS-based proteomics
Nature Communications (2024)
-
Prediction of glycopeptide fragment mass spectra by deep learning
Nature Communications (2024)
-
Network-based elucidation of colon cancer drug resistance mechanisms by phosphoproteomic time-series analysis
Nature Communications (2024)
-
Achieving quantitative reproducibility in label-free multisite DIA experiments through multirun alignment
Communications Biology (2023)
-
Data-independent acquisition boosts quantitative metaproteomics for deep characterization of gut microbiota
npj Biofilms and Microbiomes (2023)