Abstract
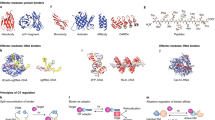
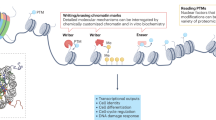
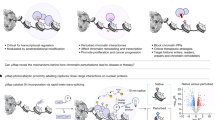
Epigenetic gene regulation is a dynamic process orchestrated by chromatin-modifying enzymes. Many of these master regulators exert their function through covalent modification of DNA and histone proteins. Aberrant epigenetic processes have been implicated in the pathophysiology of multiple human diseases. Small-molecule inhibitors have been essential to advancing our understanding of the underlying molecular mechanisms of epigenetic processes. However, the resolution offered by small molecules is often insufficient to manipulate epigenetic processes with high spatiotemporal control. Here we present a generalizable approach, referred to as 'chemo-optical modulation of epigenetically regulated transcription' (COMET), enabling high-resolution, optical control of epigenetic mechanisms based on photochromic inhibitors of human histone deacetylases using visible light. COMET probes may be translated into new therapeutic strategies for diseases where conditional and selective epigenome modulation is required.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Haggarty, S.J. & Tsai, L.H. Probing the role of HDACs and mechanisms of chromatin-mediated neuroplasticity. Neurobiol. Learn. Mem. 96, 41–52 (2011).
Orozco-Solis, R. & Sassone-Corsi, P. Epigenetic control and the circadian clock: linking metabolism to neuronal responses. Neuroscience 264, 76–87 (2014).
Goldberg, A.D., Allis, C.D. & Bernstein, E. Epigenetics: a landscape takes shape. Cell 128, 635–638 (2007).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Arrowsmith, C.H., Bountra, C., Fish, P.V., Lee, K. & Schapira, M. Epigenetic protein families: a new frontier for drug discovery. Nat. Rev. Drug Discov. 11, 384–400 (2012).
Knight, Z.A. & Shokat, K.M. Chemical genetics: where genetics and pharmacology meet. Cell 128, 425–430 (2007).
Stockwell, B.R. Exploring biology with small organic molecules. Nature 432, 846–854 (2004).
Fenno, L., Yizhar, O. & Deisseroth, K. The development and application of optogenetics. Annu. Rev. Neurosci. 34, 389–412 (2011).
Konermann, S. et al. Optical control of mammalian endogenous transcription and epigenetic states. Nature 500, 472–476 (2013).
Beharry, A.A. & Woolley, G.A. Azobenzene photoswitches for biomolecules. Chem. Soc. Rev. 40, 4422–4437 (2011).
Fehrentz, T., Schönberger, M. & Trauner, D. Optochemical genetics. Angew. Chem. Int. Edn Engl. 50, 12156–12182 (2011).
Kramer, R.H., Mourot, A. & Adesnik, H. Optogenetic pharmacology for control of native neuronal signaling proteins. Nat. Neurosci. 16, 816–823 (2013).
Schönberger, M., Althaus, M., Fronius, M., Clauss, W. & Trauner, D. Controlling epithelial sodium channels with light using photoswitchable amilorides. Nat. Chem. 6, 712–719 (2014).
Frank, J.A. et al. Photoswitchable fatty acids enable optical control of TRPV1. Nat. Commun. 6, 7118 (2015).
Brieke, C., Rohrbach, F., Gottschalk, A., Mayer, G. & Heckel, A. Light-controlled tools. Angew. Chem. Int. Edn Engl. 51, 8446–8476 (2012).
Copeland, R.A., Pompliano, D.L. & Meek, T.D. Drug-target residence time and its implications for lead optimization. Nat. Rev. Drug Discov. 5, 730–739 (2006).
Niino, H. et al. Stabilization of a labile cis-azobenzene derivative with amphiphilic cyclodextrins. Chem. Lett. 17, 1227–1230 (1988).
Shinkai, S., Minami, T., Kusano, Y. & Manabe, O. Photoresponsive crown ethers. 8. Azobenzenophane-type switched-on crown ethers which exhibit an all-or-nothing change in ion-binding ability. J. Am. Chem. Soc. 105, 1851–1856 (1983).
Bressi, J.C. et al. Exploration of the HDAC2 foot pocket: Synthesis and SAR of substituted N-(2-aminophenyl)benzamides. Bioorg. Med. Chem. Lett. 20, 3142–3145 (2010).
Bradner, J.E. et al. Chemical phylogenetics of histone deacetylases. Nat. Chem. Biol. 6, 238–243 (2010).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Andrews, D.M. et al. Design and campaign synthesis of piperidine- and thiazole-based histone deacetylase inhibitors. Bioorg. Med. Chem. Lett. 18, 2580–2584 (2008).
Lauffer, B.E. et al. Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J. Biol. Chem. 288, 26926–26943 (2013).
Garcia-Amorós, J., Martínez, M., Finkelmann, H. & Velasco, D. Kinetico-mechanistic study of the thermal cis-to-trans isomerization of 4,4′-dialkoxyazoderivatives in nematic liquid crystals. J. Phys. Chem. B 114, 1287–1293 (2010).
Zheng, Y., Thomas, P.M. & Kelleher, N.L. Measurement of acetylation turnover at distinct lysines in human histones identifies long-lived acetylation sites. Nat. Commun. 4, 2203 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Montojo, J. et al. GeneMANIA Cytoscape plugin: fast gene function predictions on the desktop. Bioinformatics 26, 2927–2928 (2010).
Beckers, T. et al. Distinct pharmacological properties of second generation HDAC inhibitors with the benzamide or hydroxamate head group. Int. J. Cancer 121, 1138–1148 (2007).
Gräff, J. & Tsai, L.H. The potential of HDAC inhibitors as cognitive enhancers. Annu. Rev. Pharmacol. Toxicol. 53, 311–330 (2013).
Gräff, J. et al. An epigenetic blockade of cognitive functions in the neurodegenerating brain. Nature 483, 222–226 (2012).
Schroeder, F.A. et al. A selective HDAC 1/2 inhibitor modulates chromatin and gene expression in brain and alters mouse behavior in two mood-related tests. PLoS One 8, e71323 (2013).
Gräff, J. et al. Epigenetic priming of memory updating during reconsolidation to attenuate remote fear memories. Cell 156, 261–276 (2014).
Bolden, J.E., Peart, M.J. & Johnstone, R.W. Anticancer activities of histone deacetylase inhibitors. Nat. Rev. Drug Discov. 5, 769–784 (2006).
Peters, J.U. Polypharmacology - foe or friend? J. Med. Chem. 56, 8955–8971 (2013).
Banks, J.L. et al. Integrated Modeling Program, Applied Chemical Theory (IMPACT). J. Comput. Chem. 26, 1752–1780 (2005).
Lee, C., Yang, W. & Parr, R.G. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B Condens. Matter 37, 785–789 (1988).
Marenich, A.V., Cramer, C.J. & Truhlar, D.G. Universal solvation model based on solute electron density and on a continuum model of the solvent defined by the bulk dielectric constant and atomic surface tensions. J. Phys. Chem. B 113, 6378–6396 (2009).
Chirlian, L.E. & Francl, M.M. Atomic charges derived from electrostatic potentials: a detailed study. J. Comput. Chem. 8, 894–905 (1987).
Acknowledgements
We thank members of the Haggarty and Mazitschek laboratories and T. Pezeril for their constructive feedback throughout the project. We acknowledge financial support from the US National Institutes of Health (R01NS088209 R.M. and S.J.H., P50CA086355 R.M., R01DA028301 S.J.H., T32-CA079443 J.A.H.). S.J.H. is supported through funding from the Tau Consortium, Bluefield Consortium for Frontotemporal Dementia and Pitt-Hopkins Research Foundation. The computations in this paper were run on the Odyssey cluster supported by the Faculty of Arts and Sciences, Division of Science, Research Computing Group at Harvard University. We thank S. Johnston for acquiring the high-resolution mass spectra, and members of the Arduino development team and the open-source Maker Movement.
Author information
Authors and Affiliations
Contributions
R.M. and S.J.H. conceived of the idea for the study; designed, directed and interpreted experiments, and wrote the manuscript; R.M. designed, synthesized and characterized inhibitors, and designed and built LED array; S.A.R. planned and performed cell-based assays, high-content image analysis, analyzed data and helped prepare the manuscript; B.G. synthesized inhibitors, planned and performed cell-based assays, analyzed data and helped prepare the manuscript; J.A.H. and D.M.S.-K. performed biochemical assays and analyzed data; L.T. performed DFT calculation studies and analyzed data; K.N.R. analyzed gene expression profiling data; J.L., W.R.-B. and B.Z. performed gene expression analysis and analyzed data; H.W. and C.S. designed experimental set-up for relaxation experiments, performed relaxation experiments and analyzed data.
Corresponding authors
Ethics declarations
Competing interests
R.M. has financial interests in SHAPE Pharmaceuticals and Acetylon Pharmaceuticals, and is the inventor on IP licensed to these two entities. S.J.H. has financial interests in Rodin Therapeutics and is an inventor on IP licensed to this entity. None of these entities were involved in the present study and the licensed IP does not include any of the work presented here. R.M., B.G., J.A.H., S.A.R. and S.J.H. have filed a patent application on the reported invention (WO 2014160221).
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1–16. (PDF 28097 kb)
Supplementary Data Set
Gene expression SNR scores; BG14, CI-994, and C60 response gene IDs (XLSX 301 kb)
Supplementary Note
Supplementary Note 1 (PDF 791 kb)
LED array
Video shows footprint of LED array in an incubator and exposure of 470 nm light (8.5 mW/cm2) modulating at 1 Hz (1 s on/7 s off) per row both with and without a 96-well plate. (MOV 21364 kb)
Rights and permissions
About this article
Cite this article
Reis, S., Ghosh, B., Hendricks, J. et al. Light-controlled modulation of gene expression by chemical optoepigenetic probes. Nat Chem Biol 12, 317–323 (2016). https://doi.org/10.1038/nchembio.2042
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2042
This article is cited by
-
NIR-light-mediated spatially selective triggering of anti-tumor immunity via upconversion nanoparticle-based immunodevices
Nature Communications (2019)
-
Lighting the way
Nature Chemical Biology (2016)
-
Optical control of epigenetics
Nature Reviews Genetics (2016)
-
Light-responsive enzyme inhibitors
Nature Methods (2016)