Abstract
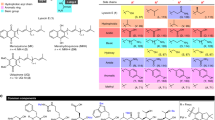
The majority of bacterial proteins are dispensable for growth in the laboratory but nevertheless have important physiological roles. There are no systematic approaches to identify cell-permeable small-molecule inhibitors of these proteins. We demonstrate a strategy to identify such inhibitors that exploits synthetic lethal relationships both for small-molecule discovery and for target identification. Applying this strategy in Staphylococcus aureus, we have identified a compound that inhibits DltB, a component of the teichoic acid D-alanylation machinery that has been implicated in virulence. This D-alanylation inhibitor sensitizes S. aureus to aminoglycosides and cationic peptides and is lethal in combination with a wall teichoic acid inhibitor. We conclude that DltB is a druggable target in the D-alanylation pathway. More broadly, the work described demonstrates a systematic method to identify biologically active inhibitors of major bacterial processes that can be adapted to numerous organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hung, D.T. & Rubin, E.J. Chemical biology and bacteria: not simply a matter of life or death. Curr. Opin. Chem. Biol. 10, 321–326 (2006).
Mitchison, T.J. Towards a pharmacological genetics. Chem. Biol. 1, 3–6 (1994).
Chung, H.S. et al. Rapid β-lactam–induced lysis requires successful assembly of the cell division machinery. Proc. Natl. Acad. Sci. USA 106, 21872–21877 (2009).
Kitano, K. & Tomasz, A. Triggering of autolytic cell wall degradation in Escherichia coli by β-lactam antibiotics. Antimicrob. Agents Chemother. 16, 838–848 (1979).
Cho, H., Uehara, T. & Bernhardt, T.G. β-lactam antibiotics induce a lethal malfunctioning of the bacterial cell wall synthesis machinery. Cell 159, 1300–1311 (2014).
Garner, E.C. et al. Coupled, circumferential motions of the cell wall synthesis machinery and MreB filaments in B. subtilis. Science 333, 222–225 (2011).
Schirner, K. et al. Lipid-linked cell wall precursors regulate membrane association of bacterial actin MreB. Nat. Chem. Biol. 11, 38–45 (2015).
van Teeffelen, S. et al. The bacterial actin MreB rotates, and rotation depends on cell-wall assembly. Proc. Natl. Acad. Sci. USA 108, 15822–15827 (2011).
Domínguez-Escobar, J. et al. Processive movement of MreB-associated cell wall biosynthetic complexes in bacteria. Science 333, 225–228 (2011).
Wright, G.D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 5, 175–186 (2007).
Le Meur, N. & Gentleman, R. Modeling synthetic lethality. Genome Biol. 9, R135 (2008).
Typas, A. et al. A tool-kit for high-throughput, quantitative analyses of genetic interactions in E. coli. Nat. Methods 5, 781–787 (2008).
Butland, G. et al. eSGA: E. coli synthetic genetic array analysis. Nat. Methods 5, 789–795 (2008).
Tong, A.H.Y. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Santa Maria, J.P. et al. Compound-gene interaction mapping reveals distinct roles for Staphylococcus aureus teichoic acids. Proc. Natl. Acad. Sci. USA 111, 12510–12515 (2014).
McLornan, D.P., List, A. & Mufti, G.J. Applying synthetic lethality for the selective targeting of cancer. N. Engl. J. Med. 371, 1725–1735 (2014).
Brown, S., Santa Maria, J.P. & Walker, S. Wall teichoic acids of gram-positive bacteria. Annu. Rev. Microbiol. 67, 313–336 (2013).
Schlag, M. et al. Role of staphylococcal wall teichoic acid in targeting the major autolysin Atl. Mol. Microbiol. 75, 864–873 (2010).
Atilano, M.L. et al. Teichoic acids are temporal and spatial regulators of peptidoglycan cross-linking in Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 107, 18991–18996 (2010).
Campbell, J. et al. Synthetic lethal compound combinations reveal a fundamental connection between wall teichoic acid and peptidoglycan biosyntheses in Staphylococcus aureus. ACS Chem. Biol. 6, 106–116 (2011).
Weidenmaier, C. et al. Role of teichoic acids in Staphylococcus aureus nasal colonization, a major risk factor in nosocomial infections. Nat. Med. 10, 243–245 (2004).
Neuhaus, F.C. & Baddiley, J. A continuum of anionic charge: structures and functions of D-alanyl-teichoic acids in Gram-positive bacteria. Microbiol. Mol. Biol. Rev. 67, 686–723 (2003).
Collins, L.V. et al. Staphylococcus aureus strains lacking D-alanine modifications of teichoic acids are highly susceptible to human neutrophil killing and are virulence attenuated in mice. J. Infect. Dis. 186, 214–219 (2002).
Santiago, M. et al. A new platform for ultra-high density Staphylococcus aureus transposon libraries. BMC Genomics 16, 252 (2015).
van Opijnen, T. & Camilli, A. Transposon insertion sequencing: a new tool for systems-level analysis of microorganisms. Nat. Rev. Microbiol. 11, 435–442 (2013).
Percy, M.G. & Gründling, A. Lipoteichoic acid synthesis and function in gram-positive bacteria. Annu. Rev. Microbiol. 68, 81–100 (2014).
Falord, M., Karimova, G., Hiron, A. & Msadek, T. GraXSR proteins interact with the VraFG ABC transporter to form a five-component system required for cationic antimicrobial peptide sensing and resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 56, 1047–1058 (2012).
Fridman, M. et al. Two unique phosphorylation-driven signaling pathways crosstalk in Staphylococcus aureus to modulate the cell-wall charge: Stk1/Stp1 meets GraSR. Biochemistry 52, 7975–7986 (2013).
Ketron, A.C., Denny, W.A., Graves, D.E. & Osheroff, N. Amsacrine as a topoisomerase II poison: importance of drug-DNA interactions. Biochemistry 1, 1730–1739 (2012).
Nelson, E.M., Tewey, K.M. & Liu, L.F. Mechanism of antitumor drug action: poisoning of mammalian DNA topoisomerase II on DNA by 4′-(9-acridinylamino)-methanesulfon-m-anisidide. Proc. Natl. Acad. Sci. USA 81, 1361–1365 (1984).
Fey, P.D. et al. A genetic resource for rapid and comprehensive phenotype screening of nonessential Staphylococcus aureus genes. MBio 4, e00537–e12 (2013).
Poyart, C. et al. Attenuated virulence of Streptococcus agalactiae deficient in D-alanyl–lipoteichoic acid is due to an increased susceptibility to defensins and phagocytic cells. Mol. Microbiol. 49, 1615–1625 (2003).
Peschel, A. et al. Inactivation of the dlt operon in Staphylococcus aureus confers sensitivity to defensins, protegrins, and other antimicrobial peptides. J. Biol. Chem. 274, 8405–8410 (1999).
Hunt, C.L., Nauseef, W.M. & Weiss, J.P. Effect of D-alanylation of (lipo)teichoic acids of Staphylococcus aureus on host secretory phospholipase A2 action before and after phagocytosis by human neutrophils. J. Immunol. 176, 4987–4994 (2006).
Peschel, A., Vuong, C., Otto, M. & Götz, F. The D-alanine residues of Staphylococcus aureus teichoic acids alter the susceptibility to vancomycin and the activity of autolytic enzymes. Antimicrob. Agents Chemother. 44, 2845–2847 (2000).
May, J.J. et al. Inhibition of the D-alanine: D-alanyl carrier protein ligase from Bacillus subtilis increases the bacterium's susceptibility to antibiotics that target the cell wall. FEBS J. 272, 2993–3003 (2005).
Reichmann, N.T., Picarra Cassona, C. & Gründling, A. Revised mechanism of D-alanine incorporation into cell wall polymers in Gram-positive bacteria. Microbiology 159, 1868–1877 (2013).
Taylor, M.S. et al. Architectural organization of the metabolic regulatory enzyme ghrelin O-acyltransferase. J. Biol. Chem. 288, 32211–32228 (2013).
Ruiz, N., Falcone, B., Kahne, D. & Silhavy, T.J. Chemical conditionality: a genetic strategy to probe organelle assembly. Cell 121, 307–317 (2005).
Santa Maria, J.P. et al. Compound-gene interaction mapping reveals distinct roles for Staphylococcus aureus teichoic acids. Proc. Natl. Acad. Sci. USA 111, 12510–12515 (2014).
Cain, B.F., Seelye, R.N. & Atwell, G.J. Potential antitumor agents 14. Acridylmethanesulfonanilides. J. Med. Chem. 17, 922–930 (1974).
Santiago, M. et al. A new platform for ultra-high density Staphylococcus aureus transposon libraries. BMC Genomics 16, 252 (2015).
Goecks, J., Nekrutenko, A. & Taylor, J. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, R86 (2010).
Blankenberg, D. et al. Galaxy: a web-based genome analysis tool for experimentalists. Curr. Protoc. Mol. Biol. 89, 19.10.1–19.10.21 (2010).
Giardine, B. et al. Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 15, 1451–1455 (2005).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Campbell, J. et al. Synthetic lethal compound combinations reveal a fundamental connection between wall teichoic acid and peptidoglycan biosyntheses in Staphylococcus aureus. ACS Chem. Biol. 6, 106–116 (2011).
Takatsuki, A., Arima, K. & Tamura, G. Tunicamycin, a new antibiotic. I. Isolation and characterization of tunicamycin. J. Antibiot. (Tokyo) 24, 215–223 (1971).
Pan, X.S. & Fisher, L.M. Streptococcus pneumoniae DNA gyrase and topoisomerase IV: overexpression, purification, and differential inhibition by fluoroquinolones. Antimicrob. Agents Chemother. 43, 1129–1136 (1999).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Corrigan, R.M., Abbott, J.C., Burhenne, H., Kaever, V. & Gründling, A. c-di-AMP is a new second messenger in Staphylococcus aureus with a role in controlling cell size and envelope stress. PLoS Pathog. 7, e1002217 (2011).
Acknowledgements
The authors thank T. Pang (Department of Microbiology and Immunobiology, Harvard Medical School) for providing plasmids and other reagents; C. Shamu, J. Smith and the staff at the ICCB-Longwood Screening Facility for compound screening; and the Harvard Medical School Biopolymers Facility and the Tufts University Core Facility for sequencing. This work was funded by U.S. National Institutes of Health grants U19AI109764 to S.W. and T.C.M., U54AI057159 to the ICCB-L Screening Facility, P01AI083214 and R01AI099144 to S.W. and F32AI118160 to S.H.M.
Author information
Authors and Affiliations
Contributions
L.P. and J.P.S.M. performed the chemical screen; T.C.M. and M.S. prepared the transposon library; L.P., L.M.M., T.C.M. and M.S. performed the Tn-seq experiments; W.L. constructed the ΔypfP mutant used for validation; L.P. selected for resistance mutations and analyzed whole genome sequences; B.M.W. performed LTA D-alanylation and gyrase assays; S.H.M. constructed complementation mutants; L.M.M. carried out MIC and spot dilution assays; and S.E.S.M. synthesized o-AMSA. S.W. designed and supervised the project, and figure design, preparation, and writing was primarily done by L.P. and S.W. with important contributions from L.M.M., J.P.S.M. and B.M.W. All of the authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1–12. (PDF 1486 kb)
Rights and permissions
About this article
Cite this article
Pasquina, L., Santa Maria, J., McKay Wood, B. et al. A synthetic lethal approach for compound and target identification in Staphylococcus aureus. Nat Chem Biol 12, 40–45 (2016). https://doi.org/10.1038/nchembio.1967
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1967
This article is cited by
-
Structural insights into the transporting and catalyzing mechanism of DltB in LTA D-alanylation
Nature Communications (2024)
-
Systematic analysis of drug combinations against Gram-positive bacteria
Nature Microbiology (2023)
-
Antibacterial activities and action mode of anti-hyperlipidemic lomitapide against Staphylococcus aureus
BMC Microbiology (2022)
-
Construction of high-density transposon mutant library of Staphylococcus aureus using bacteriophage ϕ11
Journal of Microbiology (2022)
-
Genome-wide mutant profiling predicts the mechanism of a Lipid II binding antibiotic
Nature Chemical Biology (2018)