Abstract
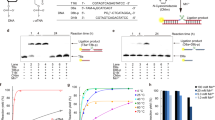
The PCR amplification of oligonucleotides enables the evolution of sequences called aptamers that bind specific targets with antibody-like affinity. However, in many applications the use of these aptamers is limited by nuclease-mediated degradation. In contrast, oligonucleotides that are modified at their sugar C2ʹ positions with methoxy or fluorine substituents are stable to nucleases, but they cannot be synthesized by natural polymerases. Here we report the development of a polymerase-evolution system and its use to evolve thermostable polymerases that efficiently interconvert C2ʹ-OMe-modified oligonucleotides and their DNA counterparts via ‘transcription’ and ‘reverse transcription’ or, more importantly, that PCR-amplify partially C2ʹ-OMe- or C2ʹ-F-modified oligonucleotides. A mechanistic analysis demonstrates that the ability to amplify the modified oligonucleotides evolved by optimizing interdomain interactions that stabilize the catalytically competent closed conformation of the polymerase. The evolved polymerases should find practical applications and the developed evolution system should be a powerful tool for tailoring polymerases to have other types of novel function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Joyce, G. F. Directed evolution of nucleic acid enzymes. Annu. Rev. Biochem. 73, 791–836 (2004).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Ng, E. W. et al. Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nature Rev. Drug Discov. 5, 123–132 (2006).
Metelev, V., Lisziewicz, J. & Agrawal, S. Study of antisense oligonucleotide phosphorothioates containing segments of oligodeoxynucleotides and 2ʹ-O- methyloligoribonucleotides. Bioorg. Med. Chem. Lett. 4, 2929–2934 (1994).
Agrawal, S. et al. Mixed-backbone oligonucleotides as second generation antisense oligonucleotides: in vitro and in vivo studies. Proc. Natl Acad. Sci. USA 94, 2620–2625 (1997).
Heemskerk, H. et al. Preclinical PK and PD studies on 2ʹ-O-methyl-phosphorothioate RNA antisense oligonucleotides in the mdx mouse model. Mol. Ther. 18, 1210–1217 (2010).
Goemans, N. M. et al. Systemic administration of PRO051 in Duchenne's muscular dystrophy. N. Engl. J. Med. 364, 1513–1522 (2011).
Manoharan, M. et al. Unique gene-silencing and structural properties of 2ʹ-fluoro-modified siRNAs. Angew. Chem. Int. Ed. 50, 2284–2288 (2011).
Rigo, F. et al. Synthetic oligonucleotides recruit ILF2/3 to RNA transcripts to modulate splicing. Nature Chem. Biol. 8, 555–561 (2012).
Kojima, T. et al. PCR amplification of 4ʹ-thioDNA using 2ʹ-deoxy-4ʹ-thionucleoside 5ʹ-triphosphates. ACS Synth. Biol. 2, 529–536 (2013).
Jones, G. D., Lesnik, E. A., Owens, S. R., Risen, L. M. & Walker, R. T. Investigation of some properties of oligodeoxynucleotides containing 4ʹ-thio-2ʹ-deoxynucleotides: duplex hybridization and nuclease sensitivity. Nucleic Acids Res. 24, 4117–4122 (1996).
Zhou, J. et al. Cell-specific RNA aptamer against human CCR5 specifically targets HIV-1 susceptible cells and inhibits HIV-1 infectivity. Chem. Biol. 22, 379–390 (2015).
Taylor, A. I. et al. Catalysts from synthetic genetic polymers. Nature 518, 427–430 (2015).
Lin, Y., Qiu, Q., Gill, S. C. & Jayasena, S. D. Modified RNA sequence pools for in vitro selection. Nucleic Acids Res. 22, 5229–5234 (1994).
Fa, M., Radeghieri, A., Henry, A. A. & Romesberg, F. E. Expanding the substrate repertoire of a DNA polymerase by directed evolution. J. Am. Chem. Soc. 126, 1748–1754 (2004).
Chen, T. & Romesberg, F. E. Directed polymerase evolution. FEBS Lett. 588, 219–229 (2014).
Chelliserrykattil, J. & Ellington, A. D. Evolution of a T7 RNA polymerase variant that transcribes 2ʹ-O-methyl RNA. Nature Biotechnol. 22, 1155–1160 (2004).
Ibach, J. et al. Identification of a T7 RNA polymerase variant that permits the enzymatic synthesis of fully 2ʹ-O-methyl-modified RNA. J. Biotechnol. 167, 287–295 (2013).
Pinheiro, V. B. et al. Synthetic genetic polymers capable of heredity and evolution. Science 336, 341–344 (2012).
Schultz, H. J. et al. Taq DNA polymerase mutants and 2′-modified sugar recognition. Biochemistry 54, 5999–6008 (2015).
Xia, G. et al. Directed evolution of novel polymerase activities: mutation of a DNA polymerase into an efficient RNA polymerase. Proc. Natl Acad. Sci. USA 99, 6597–6602 (2002).
Leconte, A. M. et al. Directed evolution of DNA polymerases for next-generation sequencing. Angew. Chem. Int. Ed. 49, 5921–5924 (2010).
Young, T. S., Ahmad, I., Yin, J. A. & Schultz, P. G. An enhanced system for unnatural amino acid mutagenesis in E. coli. J. Mol. Biol. 395, 361–374 (2010).
Marks, I. S. et al. Strain-promoted ‘click’ chemistry for terminal labeling of DNA. Bioconjug. Chem. 22, 1259–1263 (2011).
Ong, J. L., Loakes, D., Jaroslawski, S., Too, K. & Holliger, P. Directed evolution of DNA polymerase, RNA polymerase and reverse transcriptase activity in a single polypeptide. J. Mol. Biol. 361, 537–550 (2006).
Vichier-Guerre, S., Ferris, S., Auberger, N., Mahiddine, K. & Jestin, J.-L. A population of thermostable reverse transcriptases evolved from Thermus aquaticus DNA polymerase I by phage display. Angew. Chem. Int. Ed. 45, 6133–6137 (2006).
Li, Y., Korolev, S. & Waksman, G. Crystal structures of open and closed forms of binary and ternary complexes of the large fragment of Thermus aquaticus DNA polymerase I: structural basis for nucleotide incorporation. EMBO J. 17, 7514–7525 (1998).
Taq DNA polymerase with standard Taq buffer New England Biolabs Inc.; www.neb.com/products/m0273-taq-dna-polymerase-with-standard-taq-buffer
Davis, M. et al. Taq DNA polymerases having an amino acid substitution at E681 and homologs thereof exhibiting improved salt tolerance. Canadian Patent 2,381,206.
Brandis, J., Bloom, C. & Richards, J. DNA polymerases having improved labeled nucleotide incorporation properties. US Patent 6,265,193.
Brandis, J., Bloom, C., & Richards, J. DNA polymerases having improved labeled nucleotide incorporation properties. US Patent 7,897,738.
Holzberger, B., Obeid, S., Welte, W., Diederichs, K. & Marx, A. Structural insights into the potential of 4-fluoroproline to modulate biophysical properties of proteins. Chem. Sci. 3, 2924–2931 (2012).
Ghadessy, F. J. et al. Generic expansion of the substrate spectrum of a DNA polymerase by directed evolution. Nature Biotechnol. 22, 755–759 (2004).
Santangelo, T. J. & Artsimovitch, I. Termination and antitermination: RNA polymerase runs a stop sign. Nature Rev. Microbiol. 9, 319–329 (2011).
Acknowledgements
We thank the National Institutes of Health (GM097489) and the Defense Advanced Research Projects Agency (DARPA; N66001-14-2-4052) for supporting this work. We thank P. Schultz for the plasmid pEVOL-pAzF.
Author information
Authors and Affiliations
Contributions
T.C. and F.E.R. conceived and designed the experiments. T.C., N.H., Z.L, R.A. and S.S.T. performed experiments. T.C., N.H., Z.L. and F.E.R. analysed the data. T.C. and F.E.R. co-wrote the paper. F.E.R. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 9913 kb)
Rights and permissions
About this article
Cite this article
Chen, T., Hongdilokkul, N., Liu, Z. et al. Evolution of thermophilic DNA polymerases for the recognition and amplification of C2ʹ-modified DNA. Nature Chem 8, 556–562 (2016). https://doi.org/10.1038/nchem.2493
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2493
This article is cited by
-
A two-residue nascent-strand steric gate controls synthesis of 2′-O-methyl- and 2′-O-(2-methoxyethyl)-RNA
Nature Chemistry (2023)
-
Systematic bio-fabrication of aptamers and their applications in engineering biology
Systems Microbiology and Biomanufacturing (2023)
-
Biocatalysis
Nature Reviews Methods Primers (2021)
-
Modified nucleic acids: replication, evolution, and next-generation therapeutics
BMC Biology (2020)
-
Discovery and evolution of RNA and XNA reverse transcriptase function and fidelity
Nature Chemistry (2020)