Abstract
The ability of cells to adhere and sense differences in tissue stiffness is crucial for organ development and function. The central mechanisms by which adherent cells detect extracellular matrix compliance, however, are still unknown. Using two single-molecule-calibrated biosensors that allow the analysis of a previously inaccessible but physiologically highly relevant force regime in cells, we demonstrate that the integrin activator talin establishes mechanical linkages following cell adhesion, which are indispensable for cells to probe tissue stiffness. Talin linkages are exposed to a range of piconewton forces and bear, on average, 7–10 pN during cell adhesion depending on their association with F-actin and vinculin. Disruption of talin’s mechanical engagement does not impair integrin activation and initial cell adhesion but prevents focal adhesion reinforcement and thus extracellular rigidity sensing. Intriguingly, talin mechanics are isoform specific so that expression of either talin-1 or talin-2 modulates extracellular rigidity sensing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Heisenberg, C. P. & Bellaiche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Wozniak, M. A. & Chen, C. S. Mechanotransduction in development: a growing role for contractility. Nat. Rev. Mol. Cell Biol. 10, 34–43 (2009).
Franze, K., Janmey, P. A. & Guck, J. Mechanics in neuronal development and repair. Annu. Rev. Biomed. Eng. 15, 227–251 (2013).
Ingber, D. E. Mechanobiology and diseases of mechanotransduction. Ann. Med. 35, 564–577 (2003).
Lu, P., Takai, K., Weaver, V. M. & Werb, Z. Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb. Perspect. Biol. 3 10.1101/cshperspect.a005058 (2011).
Discher, D. E., Janmey, P. & Wang, Y. L. Tissue cells feel and respond to the stiffness of their substrate. Science 310, 1139–1143 (2005).
Plotnikov, S. V., Pasapera, A. M., Sabass, B. & Waterman, C. M. Force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151, 1513–1527 (2012).
Schoen, I., Pruitt, B. L. & Vogel, V. The Yin-Yang of rigidity sensing: how forces and mechanical properties regulate the cellular response to materials. Annu. Rev. Mater. Res. 43, 589–618 (2013).
Geiger, B., Spatz, J. P. & Bershadsky, A. D. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 10, 21–33 (2009).
Hoffman, B. D., Grashoff, C. & Schwartz, M. A. Dynamic molecular processes mediate cellular mechanotransduction. Nature 475, 316–323 (2011).
Elosegui-Artola, A. et al. Rigidity sensing and adaptation through regulation of integrin types. Nat. Mater. 13, 631–637 (2014).
Grashoff, C. et al. Measuring mechanical tension across vinculin reveals regulation of focal adhesion dynamics. Nature 466, 263–266 (2010).
Critchley, D. R. Biochemical and structural properties of the integrin-associated cytoskeletal protein talin. Annu. Rev. Biophys. 38, 235–254 (2009).
Kanchanawong, P. et al. Nanoscale architecture of integrin-based cell adhesions. Nature 468, 580–584 (2010).
del Rio, A. et al. Stretching single talin rod molecules activates vinculin binding. Science 323, 638–641 (2009).
Liu, J. et al. Talin determines the nanoscale architecture of focal adhesions. Proc. Natl Acad. Sci. USA 112, E4864–E4873 (2015).
Austen, K., Kluger, C., Freikamp, A., Chrostek-Grashoff, A. & Grashoff, C. Generation and analysis of biosensors to measure mechanical forces within cells. Methods Mol. Biol. 1066, 169–184 (2013).
Cost, A. L., Ringer, P., Chrostek-Grashoff, A. & Grashoff, C. How to measure molecular forces in cells: a guide to evaluating genetically-encoded FRET-based tension sensors. Cell. Mol. Bioeng. 8, 96–105 (2015).
Guo, J., Sachs, F. & Meng, F. Fluorescence-based force/tension sensors: a novel tool to visualize mechanical forces in structural proteins in live cells. Antioxid. Redox. Signal. 20, 986–999 (2014).
Finer, J. T., Simmons, R. M. & Spudich, J. A. Single myosin molecule mechanics: piconewton forces and nanometre steps. Nature 368, 113–119 (1994).
Wang, X. & Ha, T. Defining single molecular forces required to activate integrin and notch signaling. Science 340, 991–994 (2013).
Blakely, B. L. et al. A DNA-based molecular probe for optically reporting cellular traction forces. Nat. Methods 11, 1229–1232 (2014).
Duan, Y. & Kollman, P. A. Pathways to a protein folding intermediate observed in a 1-microsecond simulation in aqueous solution. Science 282, 740–744 (1998).
Zoldak, G., Stigler, J., Pelz, B., Li, H. & Rief, M. Ultrafast folding kinetics and cooperativity of villin headpiece in single-molecule force spectroscopy. Proc. Natl Acad. Sci. USA 110, 18156–18161 (2013).
Zhang, X. et al. Talin depletion reveals independence of initial cell spreading from integrin activation and traction. Nat. Cell Biol. 10, 1062–1068 (2008).
Carisey, A. et al. Vinculin regulates the recruitment and release of core focal adhesion proteins in a force-dependent manner. Curr. Biol. 23, 271–281 (2013).
Dumbauld, D. W. et al. How vinculin regulates force transmission. Proc. Natl Acad. Sci. USA 110, 9788–9793 (2013).
Humphries, J. D. et al. Vinculin controls focal adhesion formation by direct interactions with talin and actin. J. Cell Biol. 179, 1043–1057 (2007).
Hytonen, V. P. & Vogel, V. How force might activate talin’s vinculin binding sites: SMD reveals a structural mechanism. PLoS Comput. Biol. 4, e24 (2008).
Yao, M. et al. Mechanical activation of vinculin binding to talin locks talin in an unfolded conformation. Sci. Rep. 4, 4610 (2014).
Ciobanasu, C., Faivre, B. & Le Clainche, C. Actomyosin-dependent formation of the mechanosensitive talin-vinculin complex reinforces actin anchoring. Nat. Commun. 5, 3095 (2014).
Praekelt, U. et al. New isoform-specific monoclonal antibodies reveal different sub-cellular localisations for talin1 and talin2. Eur. J. Cell Biol. 91, 180–191 (2012).
Borghi, N. et al. E-cadherin is under constitutive actomyosin-generated tension that is increased at cell–cell contacts upon externally applied stretch. Proc. Natl Acad. Sci. USA 109, 12568–12573 (2012).
Krieg, M., Dunn, A. R. & Goodman, M. B. Mechanical control of the sense of touch by β-spectrin. Nat. Cell Biol. 16, 224–233 (2014).
Paszek, M. J. et al. The cancer glycocalyx mechanically primes integrin-mediated growth and survival. Nature 511, 319–325 (2014).
Cai, D. et al. Mechanical feedback through E-cadherin promotes direction sensing during collective cell migration. Cell 157, 1146–1159 (2014).
Conway, D. E. et al. Fluid shear stress on endothelial cells modulates mechanical tension across VE-cadherin and PECAM-1. Curr. Biol. 23, 1024–1030 (2013).
Moser, M., Legate, K. R., Zent, R. & Fassler, R. The tail of integrins, talin, and kindlins. Science 324, 895–899 (2009).
Miller, K., Chinzei, K., Orssengo, G. & Bednarz, P. Mechanical properties of brain tissue in-vivo: experiment and computer simulation. J. Biomech. 33, 1369–1376 (2000).
Anthis, N. J., Wegener, K. L., Critchley, D. R. & Campbell, I. D. Structural diversity in integrin/talin interactions. Structure 18, 1654–1666 (2010).
Cecconi, C., Shank, E. A., Dahlquist, F. W., Marqusee, S. & Bustamante, C. Protein-DNA chimeras for single molecule mechanical folding studies with the optical tweezers. Eur. Biophys. J. 37, 729–738 (2008).
Stigler, J., Ziegler, F., Gieseke, A., Gebhardt, J. C. & Rief, M. The complex folding network of single calmodulin molecules. Science 334, 512–516 (2011).
von Hansen, Y., Mehlich, A., Pelz, B., Rief, M. & Netz, R. R. Auto- and cross-power spectral analysis of dual trap optical tweezer experiments using Bayesian inference. Rev. Sci. Instrum. 83, 095116 (2012).
Mehlich, A., Austen, K., Ringer, P., Rief, M. & Grashoff, C. Evaluation of molecular tension sensors using single-molecule force spectroscopy and live cell FRET imaging. Protoc. Exch. http://dx.doi.org/10.1038/protex.2015.095 (2015).
Subauste, M. C. et al. Rho family proteins modulate rapid apoptosis induced by cytotoxic T lymphocytes and Fas. J. Biol. Chem. 275, 9725–9733 (2000).
Conti, F. J., Monkley, S. J., Wood, M. R., Critchley, D. R. & Muller, U. Talin 1 and 2 are required for myoblast fusion, sarcomere assembly and the maintenance of myotendinous junctions. Development 136, 3597–3606 (2009).
Monkley, S. J. et al. Disruption of the talin gene arrests mouse development at the gastrulation stage. Dev. Dyn. 219, 560–574 (2000).
Schiller, H. B. et al. β1- and αv-class integrins cooperate to regulate myosin II during rigidity sensing of fibronectin-based microenvironments. Nat. Cell Biol. 15, 625–636 (2013).
Sprague, B. L. & McNally, J. G. FRAP analysis of binding: proper and fitting. Trends Cell Biol. 15, 84–91 (2005).
Carisey, A., Stroud, M., Tsang, R. & Ballestrem, C. Fluorescence recovery after photobleaching. Methods Mol. Biol. 769, 387–402 (2011).
Goult, B. T. et al. The structure of an interdomain complex that regulates talin activity. J. Biol. Chem. 284, 15097–15106 (2009).
Goksoy, E. et al. Structural basis for the autoinhibition of talin in regulating integrin activation. Mol. Cell 31, 124–133 (2008).
Goult, B. T. et al. Structural studies on full-length talin1 reveal a compact auto-inhibited dimer: implications for talin activation. J. Struct. Biol. 184, 21–32 (2013).
Wurflinger, T., Gamper, I., Aach, T. & Sechi, A. S. Automated segmentation and tracking for large-scale analysis of focal adhesion dynamics. J. Microsc. 241, 37–53 (2011).
Yeung, T. et al. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil. Cytoskeleton 60, 24–34 (2005).
Boudou, T., Ohayon, J., Picart, C., Pettigrew, R. I. & Tracqui, P. Nonlinear elastic properties of polyacrylamide gels: implications for quantification of cellular forces. Biorheology 46, 191–205 (2009).
Lembong, J., Sabass, B., Sun, B., Rogers, M. E. & Stone, H. A. Mechanics regulates ATP-stimulated collective calcium response in fibroblast cells. J. R. Soc. Interface 12, 20150140 (2015).
Sabass, B., Gardel, M. L., Waterman, C. M. & Schwarz, U. S. High resolution traction force microscopy based on experimental and computational advances. Biophys. J. 94, 207–220 (2008).
Jares-Erijman, E. A. & Jovin, T. M. FRET imaging. Nat. Biotechnol. 21, 1387–1395 (2003).
Acknowledgements
C.G. is supported by the German Research Council (DFG, GR3399/2-1 and GR3399/5-1), the Collaborative Research Centre SFB863 (B9) and a Paul Gerson Unna Research Group of the Max Planck Förderstiftung. A.M. is supported by the Nanosystems Initiative Munich (NIM); M.R. is supported by the German Research Council through the Collaborative Research Centre SFB863 (A2). R.Z. is supported by R01-DK083187, R01-DK075594, R01-DK069221 and VA Merit Award 1I01BX002196. B.S. acknowledges financial support from the DAAD. The authors thank R. Fässler for continuous support and helpful discussions; F. Schnorrer for critically reading the manuscript; A. Lambacher and the MPIB core facility for technical support; G. Zoldak for helpful discussion, A. Sechi for sharing data analysis software and D. Critchley (University of Leicester, UK) for providing talin-2 cDNA.
Author information
Authors and Affiliations
Contributions
C.G. and K.A. initiated the project, generated cell lines, performed cellular and biochemical experiments and analysed data; P.R. generated cell lines, performed cellular experiments, wrote data analysis software and analysed data. A.M. and M.R. performed the single-molecule calibration and theoretical modelling. A.C.-G. created talin expression constructs, C.Kluger generated vinculin expression constructs and cell lines, C.Kluger and C.Klingner wrote data analysis software. K.A., C.Kluger and B.S. performed traction force microscopy experiments and analyses. R.Z. provided genetically modified talin cells. C.G. wrote the manuscript with the input from all authors. K.A., P.R. and A.M. contributed equally.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Single-molecule calibration of HP35-TS and HP35st-TS.
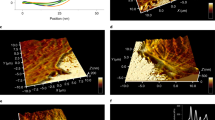
(a) Filtered force-extension traces (FE) of consecutive stretch (blue) and relax (grey) cycles of HP35-TS; black solid lines are WLC model fits to the data marking folded and unfolded protein states. (b) FE of consecutive stretch (red) and relax (grey) cycles of HP35st-TS; black solid lines are WLC model fits to the data. (c) Superimposed HP35-TS (blue) and HP35st-TS (red) stretch and relax cycles; solid lines are entire data fits, dashed lines show WLC model fits. (d) Standard deviation of bead deflection fluctuations as a function of mean force for DNA-HP35-TS (blue), DNA-HP35st-TS (red) and only DNA (black). (e) Folding free energy distributions of HP35-TS (blue) and HP35st-TS (red); solid lines are Gaussian fits to the data. (f) Modelled force-extension curve using a two-state model with mechanical parameters of HP35st-TS. The dashed black line represents the folded state with zero contour length; the black solid line represents the completely unfolded state with the contour length Ltot; the red line indicates the average protein extension 〈xp〉. The corresponding probability plot for the folded and unfolded state is shown next to the force-extension plot. (g) Force-extension model fit for HP35st-TS using a three-state model (also shown in Fig. 1g, h). The dashed black line represents the folded state, the grey dashed line the half folded/unfolded state with contour length Ltot/2 and the black solid line the completely unfolded state with contour length Ltot; the red line indicates the average protein extension 〈xp〉. The corresponding probability plot for the folded, half folded/unfolded and unfolded state is shown next to the force-extension plot. (h) Comparison of a two-state model fit with a three-state model fit to HP35-TS force-extension data (same data as in Fig. 1d). The residual plot in the upper graph indicates a better fit when a three-state model is used. (i) Protein stability curve for HP35 (black line) resulting from a combined fit to published experimental data. Data covering 0–40 °C (indicated as black empty squares) were taken from1, data for 55–80 °C (indicated by the thick grey line) were derived from2. Red circles mark the free energy values at 30 °C and 37 °C.
Supplementary Figure 2 The talin tension sensor rescues the talin-1/2 knockout phenotype.
(a) Representative images from 3 independent experiments showing paxillin-stained (red) Tln1f/fTln2−/− and Tln1−/−Tln2−/− cells; note the lack of FA formation and cell spreading; scale bars, 20 μm. (b) Representative western blots from 3 independent experiments demonstrating comparable expression levels and rescue of FAK phosphorylation in Tln1−/−Tln2−/− cells by expression of Tln1Y (Y), Tln1TS (TS) and Tln1Con (Con); phospho-FAK blots indicate FAK pY-397. Unprocessed original scans of western blots are shown in Supplementary Fig. 5. (c) Live cell FRET efficiencies as a function of mean fluorescence intensities for Tln1TS (blue) and Tln1Con (red) (n = 129 and 137 cells respectively; pooled from 5 independent experiments) indicating FRET efficiencies are independent of the used fluorescence intensities. (d) Ratiometric FRET analysis of Tln1Con and Tln1TS cells imaged at 30 °C and 37 °C (n = 29, 22, 27 and 24 cells respectively from left to right; 3 independent experiments). (e) FRET increase in Tln1TS cells in the absence of vinculin indicates reduced tension across talin-1; re-expression of full length vinculin (V-wt), but not a mutant vinculin unable to engage the f-actin cytoskeleton (V-mut), rescues talin tension (n = 23, 16, 21 and 26 cells respectively from left to right; 3 independent experiments). (d,e: Kolmogorov–Smirnov test, ∗∗∗:p < 0.001; ∗∗:p < 0.01; ∗:p < 0.05; not significant (n.s.): p > 0.05). Boxplots indicate the median (red line) as well as 25th and 75th percentiles; whiskers reach out to 2.7 standard deviations (σ). Statistic source data are available in Supplementary Table 1.
Supplementary Figure 3 Effects of talin deletion mutants on the organization of the actin cytoskeleton.
(a) Actin networks of the representative Tln1Y, Tln1-2300Y and Tln1-950Y cells shown in Fig. 4b; note the lack of actin stress fibers in Tln1-950Y cells; scale bars, 20 μm. (b,c) Representative western blots from 3 independent experiments demonstrating comparable talin (b) and vinculin (c) expression levels in Tln1Y and Tln1-950Y cells. Unprocessed original scans of western blots are shown in Supplementary Fig. 5.
Supplementary Figure 4 Talin isoform studies.
(a,b) Western blot analysis (3 independent experiments) of protein lysates from Tln1Y and Tln2Y cells demonstrating comparable expression levels and equal rescue of FAK phosphorylation. Unprocessed original scans of western blots are shown in Supplementary Fig. 5. (c) Representative FACS histogram of cells expressing Tln1Y (black) or Tln2Y (red) labelled for beta1 integrin; the negative control is shown in grey (4 independent experiments). (d) Reduced mobile fraction in Tln2Y expressing cells (n = 18 (Tln1Y) and 17 (Tln2Y) cells; pooled from 3 independent experiments). (e) Intermolecular FRET analysis in cells co-expressing Tln1C-i/Tln1Y-i or Tln2C-i/Tln2Y-i seeded on FN-coated glass coverslips; note equally low intermolecular FRET levels (n = 31 (Tln1C-i/Tln1Y-i) and 18 (Tln2C-i/Tln2Y-i) cells; 3 independent experiments). (f) No FRET efficiency differences between Tln1Con and Tln2Con as well as Tln1TS and Tln2TS cells when seeded on pL-coated glass slides indicating integrin-specificity of isoform-specific differences (n = 22, 29, 35 and 11 cells respectively from left to right; 3 independent experiments). (g) Elevated FRET efficiencies for HP35st (10 pN) probes in Tln1TS and Tln2TS as compared to HP35 (7 pN) when cells were seeded on FN-coated glass coverslips (n = 18, 27, 19 and 27 cells respectively from left to right; 6 independent experiments). These data indicate that talin-2, similar to talin-1, is exposed to a range of forces. (d–g Kolmogorov–Smirnov test. ∗∗∗:p < 0.001; ∗∗:p < 0.01, not significant (n.s.): p > 0.05). Boxplots indicate the median (red line) as well as 25th and 75th percentiles; whiskers reach out to 2.7 standard deviations (σ). Statistic source data are available in Supplementary Table 1.
Supplementary Figure 5 Unprocessed representative western blots.
Black boxes with dashed lines indicate how blots were cropped for Supplementary Figs 2–4.
Supplementary information
Supplementary Information
Supplementary Information (PDF 5913 kb)
Supplementary Table 1
Supplementary Information (XLSX 206 kb)
Rights and permissions
About this article
Cite this article
Austen, K., Ringer, P., Mehlich, A. et al. Extracellular rigidity sensing by talin isoform-specific mechanical linkages. Nat Cell Biol 17, 1597–1606 (2015). https://doi.org/10.1038/ncb3268
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3268
This article is cited by
-
Curved adhesions mediate cell attachment to soft matrix fibres in three dimensions
Nature Cell Biology (2023)
-
Nuclear lamina strain states revealed by intermolecular force biosensor
Nature Communications (2023)
-
Molecular mechanocytometry using tension-activated cell tagging
Nature Methods (2023)
-
Talin and kindlin use integrin tail allostery and direct binding to activate integrins
Nature Structural & Molecular Biology (2023)
-
Tension-tuned receptors for synthetic mechanotransduction and intercellular force detection
Nature Biotechnology (2023)