Abstract
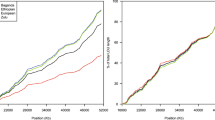
Meiotic recombinations contribute to genetic diversity by yielding new combinations of alleles. Recently, high-resolution recombination maps were inferred from high-density single-nucleotide polymorphism (SNP) data using linkage disequilibrium (LD) patterns that capture historical recombination events1,2. The use of these maps has been demonstrated by the identification of recombination hotspots2 and associated motifs3, and the discovery that the PRDM9 gene affects the proportion of recombinations occurring at hotspots4,5,6. However, these maps provide no information about individual or sex differences. Moreover, locus-specific demographic factors like natural selection7 can bias LD-based estimates of recombination rate. Existing genetic maps based on family data avoid these shortcomings8, but their resolution is limited by relatively few meioses and a low density of markers. Here we used genome-wide SNP data from 15,257 parent–offspring pairs to construct the first recombination maps based on directly observed recombinations with a resolution that is effective down to 10 kilobases (kb). Comparing male and female maps reveals that about 15% of hotspots in one sex are specific to that sex. Although male recombinations result in more shuffling of exons within genes, female recombinations generate more new combinations of nearby genes. We discover novel associations between recombination characteristics of individuals and variants in the PRDM9 gene and we identify new recombination hotspots. Comparisons of our maps with two LD-based maps inferred from data of HapMap populations of Utah residents with ancestry from northern and western Europe (CEU) and Yoruba in Ibadan, Nigeria (YRI) reveal population differences previously masked by noise and map differences at regions previously described as targets of natural selection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
McVean, G. A. et al. The fine-scale structure of recombination rate variation in the human genome. Science 304, 581–584 (2004)
Myers, S., Bottolo, L., Freeman, C., McVean, G. & Donnelly, P. A fine-scale map of recombination rates and hotspots across the human genome. Science 310, 321–324 (2005)
Myers, S., Freeman, C., Auton, A., Donnelly, P. & McVean, G. A common sequence motif associated with recombination hot spots and genome instability in humans. Nature Genet. 40, 1124–1129 (2008)
Baudat, F. et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327, 836–840 (2010)
Myers, S. et al. Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science 327, 876–879 (2010)
Parvanov, E. D., Petkov, P. M. & Paigen, K. Prdm9 controls activation of mammalian recombination hotspots. Science 327, 835 (2010)
O’Reilly, P. F., Birney, E. & Balding, D. J. Confounding between recombination and selection, and the Ped/Pop method for detecting selection. Genome Res. 18, 1304–1313 (2008)
Kong, A. et al. A high-resolution recombination map of the human genome. Nature Genet. 31, 241–247 (2002)
Kong, A. et al. Detection of sharing by descent, long-range phasing and haplotype imputation. Nature Genet. 40, 1068–1075 (2008)
Kong, A. et al. Parental origin of sequence variants associated with complex diseases. Nature 462, 868–874 (2009)
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. B 39, 1–38 (1977)
The International HapMap Consortium A haplotype map of the human genome. Nature 437, 1299–1320 (2005)
Lang, M. R., Patterson, L. B., Gordon, T. N., Johnson, S. L. & Parichy, D. M. Basonuclin-2 requirements for zebrafish adult pigment pattern development and female fertility. PLoS Genet. 5, e1000744 (2009)
Broman, K. W., Murray, J. C., Sheffield, V. C., White, R. L. & Weber, J. L. Comprehensive human genetic maps: individual and sex-specific variation in recombination. Am. J. Hum. Genet. 63, 861–869 (1998)
Kong, A. et al. Sequence variants in the RNF212 gene associate with genome-wide recombination rate. Science 319, 1398–1401 (2008)
Akey, J. M. Constructing genomic maps of positive selection in humans: where do we go from here? Genome Res. 19, 711–722 (2009)
Stefansson, H. et al. A common inversion under selection in Europeans. Nature Genet. 37, 129–137 (2005)
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999)
Acknowledgements
We thank D. Reich for discussion and suggestions.
Author information
Authors and Affiliations
Contributions
A.K. and K.S. planned and directed the research. A.K. wrote the first draft of the paper and, with K.S., U.T. and A.H., wrote most of the final version. D.F.G. improved previous phasing procedures and, with G.M., made the recombination calls. G.T. created the maps and, with M.L.F., assisted A.K. in the analyses. U.T., Aslaug J., A.S., Adalbjorg J., K.T.K. and G.B.W. performed experiments providing information on sequences at the PRDM9 gene. A.G. did the variant imputations. G.M. and S.A.G. determined the locations and intensities of genomic features. A.H. assisted in the study on selection.
Corresponding authors
Ethics declarations
Competing interests
The authors are all employees of deCode Genetics, a biotechnology company that provides genetic testing services, and own stocks or stock options in the company.
Additional information
The maps constructed in this study are available at http://www.decode.com/addendum.
Supplementary information
Supplementary Information
The file contains Supplementary Notes 1-10, Supplementary Tables 1-6, Supplementary Figures 1-4 with legends and additional references. (PDF 656 kb)
Rights and permissions
About this article
Cite this article
Kong, A., Thorleifsson, G., Gudbjartsson, D. et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 467, 1099–1103 (2010). https://doi.org/10.1038/nature09525
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09525
This article is cited by
-
Variant in the synaptonemal complex protein SYCE2 associates with pregnancy loss through effect on recombination
Nature Structural & Molecular Biology (2024)
-
Inversion polymorphism in a complete human genome assembly
Genome Biology (2023)
-
Emergence and influence of sequence bias in evolutionarily malleable, mammalian tandem arrays
BMC Biology (2023)
-
Analysis of archaic human haplotypes suggests that 5hmC acts as an epigenetic guide for NCO recombination
BMC Biology (2022)
-
Genome-wide recombination map construction from single sperm sequencing in cattle
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.