Abstract
The term ‘field effect’ (also known as field defect, field cancerization, or field carcinogenesis) has been used to describe a field of cellular and molecular alteration, which predisposes to the development of neoplasms within that territory. We explore an expanded, integrative concept, ‘etiologic field effect’, which asserts that various etiologic factors (the exposome including dietary, lifestyle, environmental, microbial, hormonal, and genetic factors) and their interactions (the interactome) contribute to a tissue microenvironmental milieu that constitutes a ‘field of susceptibility’ to neoplasia initiation, evolution, and progression. Importantly, etiological fields predate the acquisition of molecular aberrations commonly considered to indicate presence of filed effect. Inspired by molecular pathological epidemiology (MPE) research, which examines the influence of etiologic factors on cellular and molecular alterations during disease course, an etiologically focused approach to field effect can: (1) broaden the horizons of our inquiry into cancer susceptibility and progression at molecular, cellular, and environmental levels, during all stages of tumor evolution; (2) embrace host–environment–tumor interactions (including gene-environment interactions) occurring in the tumor microenvironment; and, (3) help explain intriguing observations, such as shared molecular features between bilateral primary breast carcinomas, and between synchronous colorectal cancers, where similar molecular changes are absent from intervening normal colon. MPE research has identified a number of endogenous and environmental exposures which can influence not only molecular signatures in the genome, epigenome, transcriptome, proteome, metabolome and interactome, but also host immunity and tumor behavior. We anticipate that future technological advances will allow the development of in vivo biosensors capable of detecting and quantifying ‘etiologic field effect’ as abnormal network pathology patterns of cellular and microenvironmental responses to endogenous and exogenous exposures. Through an ‘etiologic field effect’ paradigm, and holistic systems pathology (systems biology) approaches to cancer biology, we can improve personalized prevention and treatment strategies for precision medicine.
Similar content being viewed by others
Main
Cancers are fundamentally complex, multifactorial, genomic, and epigenomic diseases,1, 2, 3, 4, 5 which represent a major burden on societies globally. However, many cancers are potentially avoidable, with estimates suggesting that 60% of cancer deaths in the United States are attributable to a limited number of lifestyle factors.6 A better understanding of modifiable contributors to cancer initiation, evolution, and progression is a prerequisite for accurate risk prediction and the development of better strategies for prevention, early detection, treatment, and surveillance.1, 2, 3, 4, 5, 7, 8, 9, 10
The revolutionary theory of field effect (also known as field defect, field cancerization, or field carcinogenesis) has been continually adapted and updated since it was first consolidated by Slaughter et al11 in 1953. The description of ‘field cancerization’11 is regarded as one of the landmarks of the past 100 years of cancer research.12 Several authors have recently reviewed the field effect concept and its evolution,13, 14, 15, 16, 17, 18 and the presence of a recent textbook19 entirely devoted to the topic attests to its continued clinical and scientific importance.
In this article, we offer a reappraisal of field effect, approaching the concept from an etiologic perspective. We term this alternative model ‘etiologic field effect’, where endogenous and exogenous etiologic factors (such as dietary, lifestyle, environmental, microbial, hormonal, and genetic variations), and their interactions, predispose to an abnormal tissue microenvironmental milieu that can influence all stages of tumor evolution. We have taken into account the possible contribution of stromal cells and the microenvironment,20, 21, 22, 23, 24 and developed a paradigm where macroenvironmental and microenvironmental influences, in their totality, contribute to a field of etiologic predisposition to disease. The etiologic field effect concept embraces tumor–host interactions,3, 25, 26, 27, 28, 29, 30, 31, 32 and gene–environment interactions, which have become increasingly important in molecular epidemiology.33 There are several advantages to an etiologically oriented model of field effect, as elaborated upon in the following sections. An etiologic field effect concept can enhance the scope of the traditional field effect model and can effectively explain a variety of phenomena relevant to cancer causation and progression. Effective lifestyle interventions, such as dietary modification and physical activity, can be thought of as mechanisms through which etiologic fields can be attenuated throughout the body, preventing cancer occurrence and progression, and decreasing cancer burden in our society.34, 35
Evolution of the field effect concept
The concept of field effect was proposed by Slaughter et al,11 in their 1953 landmark paper, in an attempt to explain the phenomenon of synchronous or metachronous primary tumors arising within the oral mucosa. In this context, ‘field effect’ implied an inherent predisposition of the non-cancerous mucosa to malignant transformation.
Before the advent of molecular pathology, support for the field effect model relied on pathological observations describing histological abnormalities in grossly normal-appearing tissue adjacent to cancers.11 Subsequently, molecular genetic analyses of cancerous, precancerous, and normal tissues have yielded persuasive evidence that a field of cancer-predisposing molecular alterations can be present even in microscopically normal tissue.13, 14, 15, 16, 17, 18, 19, 36, 37 The existence of field effect has been described in a variety of tissues, spanning almost all organ systems in the human body.13, 14, 15, 16, 17, 18, 19, 38, 39, 40, 41, 42, 43, 44, 45 More recently, studies of epigenetic changes in tumors and normal cells,14, 15, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57 as well as analyses of stromal cells and the tissue microenvironment have contributed to the molecular field effect concept.20, 21, 22, 23, 24
Consequently, the prevailing interpretation of field effect is that a field of somatic molecular alteration in a given organ or tissue predisposes to tumor development within that field. The mechanisms through which geographic fields of molecularly abnormal cells arise are not fully understood, but clonal expansion and intraepithelial migration of genetically altered cells within contiguous epithelial structures has been proposed.36, 58 As a result of supporting evidence that has accrued through histopathological, genetic, and, most recently, epigenetic studies, the field effect concept has become firmly established.13, 14, 15, 16, 17, 18, 19 Importantly, altered molecular field may represent a potential therapeutic target.59, 60
Synchronous primary tumors and field effect: insights and intrigue
Field effects have been implicated in the co-occurrence of tumors in more than one organ. Examples of such multi-organ involvement by field effect include tumors arising within the ductal epithelia of the pancreas, ampulla of Vater, extrahepatic bile ducts, and gallbladder,44 respiratory epithelia of the lung, bronchi, trachea, larynx and nose,59, 61 and urothelium of the bladder, ureters, and renal pelves.45, 62, 63, 64 While shared ontogeny has been hypothesized to contribute to certain multi-organ field effects (eg in epithelia derived from embryonic foregut),44 multi-organ involvement can span tissues arising from more than one embryonic germ layer, as exemplified by urothelial tumors; upper urinary tract structures derive from the mesonephros (mesoderm), while the bladder and urethra arise from the urogenital sinus (endoderm).
The term ‘synchronous tumors’ generally refers to two or more primary tumors arising contemporaneously within a single individual. Synchronous primary neoplasms, particularly those originating within a single contiguous organ system, may develop as a result of field effect phenomena. It should be noted, however, that the absence of synchronous, or even a solitary cancer, does not exclude the existence of field effect; cancer development, per se, is not a requirement for defining field effect. This is an important consideration if we are to exploit field effect to develop screening and preventive strategies.
Synchronous cancers can provide a unique insight into the somatic molecular aberrations that might constitute a field effect.65, 66 Indeed, a number of studies have documented the presence of shared molecular features between synchronous primary tumors,67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77 which would support the influence of a field effect present in the ostensibly ‘normal’ tissue from which they have arisen.13, 14, 15, 16, 17 Alternatively, it can be considered that synchronous primary tumors arise through the interplay between common etiologic contributors, such as genetic predisposition, microbial and environmental exposures, and lifestyle factors, which facilitate progression through certain common carcinogenic pathways.65, 67, 71 Importantly, the latter model does not imply that the molecular features shared by synchronous tumors need to be present in the background ‘normal-appearing’ tissue.
In structurally continuous epithelial tissues, such as the orodigestive and respiratory mucosae, it is perhaps conceptually easy to envisage how multiple primary tumors could arise from a field of molecularly altered cells. It is well documented, however, that field effects appear to influence carcinogenesis in non-contiguous structures, for example, in bilateral primary breast cancer. Right and left breasts are separate organs, with no connections existing between glandular or ductal epithelia of the right and left breasts. Despite this anatomic independence, a number of studies have shown that synchronous and metachronous bilateral breast cancers tend to demonstrate concordance in expression status for the hormone receptors ESR1 (estrogen receptor-alpha) and PGR (progesterone receptor).71 These data imply that, in bilateral breast cancer, independent primary tumors tend to evolve through similar carcinogenic pathways, compared with cancers arising in two different individuals.
Similarly, spatially distinct primary tumors can arise within a single organ system despite the complete absence of detectable classical field change in the background normal mucosa. We have previously demonstrated that synchronous colorectal cancers tend to show concordant molecular features (including DNA hypomethylation and CpG island hypermethylation) without similar molecular changes in intervening normal colonic mucosa.67 While these observations are not necessarily at odds with the conventional field effect model, they underscore the need for emphasis to be placed on the putative factors that predispose to tumor initiation, or facilitate tumor evolution through specific, common carcinogenic pathways, where the end result is tumors with shared molecular features. It would also seem advantageous to develop a model where detectable somatic alterations, similar to those found in established cancers, are not prerequisite for defining a field.
Predisposition to neoplasia: exploring the geographic limits of field effect
It is incontrovertible that multiple primary tumors can arise within a background population of genetically predisposed cells. One concrete example of this mechanism is the presence of highly penetrant cancer syndromes,78 such as Lynch syndrome, where genetic predisposition to multiple primary tumors in one or more organ systems has been described.79, 80 In genetic predisposition syndromes, virtually every cell in the body carries a copy of the mutated gene and, as such, these syndromes may be considered whole-body mutational field effects. Germline inheritance of cancer-predisposing variants is perhaps beyond the intended scope of the conventional field effect concept. Furthermore, familial cancer predisposition syndromes contribute only a relatively small proportion of cases to the overall incidence figures for common cancers. Genetic influences, however, remain important in ‘sporadic’ cancers, which are considered to result from the interplay of genetic and environmental influences.81, 82
In non-syndromic cancers, evidence suggests that high-prevalence low-penetrance genetic variants (including those identified by genome-wide association study, GWAS) predispose to the acquisition of specific somatic molecular alterations.83, 84 For example, studies have shown a consistent association between a common single nucleotide polymorphism in the MGMT promoter (rs16906252), and MGMT promoter CpG island hypermethylation in several cell types, including colorectal cancer,85 normal colonic cells,86 normal peripheral blood cells,87 lung adenocarcinoma and premalignant lesions,88 and malignant pleural mesothelioma.89 In this example, a whole-body field effect, conferred by the common MGMT promoter single nucleotide polymorphism, appears to predispose many distinct cell types to an acquired epigenetic event, ie, somatic MGMT promoter hypermethylation.
In a manner analogous to inherited genetic variants, environmental, and other exogenous exposures, such as dietary and lifestyle factors, may also actuate or promote the accrual of specific somatic genetic or epigenetic alterations.90, 91 Indeed, epigenetic mechanisms are recognized to serve as a link between environmental influences and gene regulation.92
Beyond cancer syndromes, it is postulated that genetic determinants, endogenous, and exogenous environmental exposures can influence neoplastic transformation at multiple body sites. One would therefore expect there to be evidence of a wider, multi-organ, or even whole-body, field effect in cancer-predisposed individuals. It has been proposed that biomarkers at ‘surrogate anatomic/functional sites’ can be evaluated for presence of ‘an extended field effect’, indicative of elevated cancer risk.54, 61, 62, 93, 94 Likewise, epigenetic aberrations detectable in peripheral blood cells have been speculated to reflect constitutional cancer susceptibility.95, 96 Importantly, epidemiological studies have demonstrated clear pleiotropic effects for certain etiologic exposures, most notably smoking. Moreover, data from a large population-based study suggest increased familial clustering of cancers at different sites, which may be due to common genetic susceptibility as well as shared environmental exposures.97 Thus, evidence does exist to support a more systemic etiologic contribution to field effect, involving the interplay between germline genetic variants and environmental exposures.
The genesis of the ‘etiologic field effect’
The evolving field of molecular pathological epidemiology (MPE) has resulted in the discovery of a number of robust relationships between etiologic factors and somatic molecular alterations in human cancers and normal tissues.90, 91 These relationships include the associations between the following: reproductive and hormonal influences (such as age at menarche, parity, age at first full term-pregnancy, lactation, and hormone therapy) and risk of molecularly defined subtypes of breast cancer (by ESR1, PGR and ERBB2 expression);98, 99, 100, 101 the MLH1 rs1800734 single nucleotide polymorphism and MLH1 promoter hypermethylation (or microsatellite instability) in endometrial and colorectal cancers;102, 103, 104, 105, 106 genetic modifiers of one-carbon metabolism, micronutrient intake, and DNA methylation in colorectal cancer;107, 108, 109, 110, 111, 112, 113, 114 body mass index (or obesity), microsatellite instability, and fatty acid synthase (FASN) expression level in colorectal cancer;115, 116, 117, 118, 119, 120, 121 cigarette smoking and microsatellite instability, CpG island methylator phenotype, and BRAF mutation in colorectal cancer;122, 123, 124, 125, 126, 127 cigarette smoking and KRAS mutation in lung tumors;128, 129, 130 young age of onset and family history of colorectal cancer, and LINE-1-hypomethylated colorectal cancer;131, 132, 133, 134, 135 interactions between aspirin use and molecular features of colorectal cancer;136, 137, 138, 139 Epstein–Barr virus and CpG island hypermethylation in gastric cancer;140, 141, 142 H. pylori infection and CpG island methylation in gastric epithelial cells;143, 144 viral hepatitis and CpG island hypermethylation in hepatocellular carcinoma;145, 146, 147 and the MGMT rs16906252 promoter single nucleotide polymorphism and MGMT promoter hypermethylation in normal cells and various cancers.85, 86, 87, 88, 89 The MPE paradigm, which can encompass all human diseases,148 is firmly established,149, 150, 151, 152 and has gained widespread recognition.121, 153, 154, 155, 156, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 167, 168, 169, 170, 171, 172, 173, 174, 175, 176
As a result of MPE research, considerable evidence has accumulated to support the concept that cells in one or more organs, or organ systems, can become predisposed, in a tissue-specific or less specific fashion, to the acquisition of well-defined somatic molecular alterations. Thus, internal susceptibility, in the form of heritable genetic and epigenetic variants, and exposure to exogenous influences, such as microorganisms, environmental toxins, dietary components, and lifestyle factors, converge at the level of the tissue microenvironment, and mediate the propensity to neoplastic transformation and progression through cell–cell and extracellular matrix–cell interactions.4, 177, 178 In other words, every cancer (or, indeed, every disease process) results from changes in interactomes, with interactomes ultimately dictating how tumors behave. Interactomes179 encompass the entirety of complex molecular interactions within a cell, between cells and stromal components in the tissue microenvironment, within tissues and organs, and even at the level of a whole organism. The interactome is the ultimate interface through which external stimuli interact with host biological systems. The interactome therefore includes gene–environment interactions, which have become increasingly important as the basis for molecular epidemiology studies. Early changes in interactomes can be regarded as an expanded notion of ‘field effect’, and could be utilized as a marker of increased tumor or disease risk.
There is no real provision for the interaction of heritable and environmental risk modifiers in the conventional interpretation of field effect. Taking into account the aforementioned MPE research findings, and cognizant of the importance of the potential for interplay between diverse etiologic exposures in carcinogenesis, we developed an alternative field effect model, which we term ‘etiologic field effect’. In contrast to the conventional field effect model, where the ‘cancer-susceptible field’ comprises a distinct molecular or cellular change in an anatomically defined area, the ‘etiologic field effect’ focuses principally on the dynamic interplay between fields of exposure to etiologic factors, which may alter a tissue’s microenvironmental milieu. An etiologic field can be defined as ‘a functional field of altered tissue microenvironment that predisposes to the acquisition of specific somatic molecular changes through alterations in cellular and extracellular interactomes’. Etiologic fields are characterized by the presence of common etiologic exposures, rather than by cellular molecular aberrations. Since exposures frequently transcend anatomic boundaries, etiologic fields are not restricted to contiguous epithelial structures. As etiologic exposures predate the establishment of pathological cellular and molecular aberrations that lead to neoplastic initiation and progression, ‘etiologic field effects’ are potentially reversible and represent modifiable targets for intervention. Table 1 contrasts key features of the conventional field effect with those of the ‘etiologic field effect’ concept.
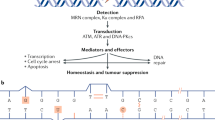
There are tangible examples that illustrate how an etiologically based model is better positioned to explain certain field effect phenomena. Smoking is well documented as a risk factor for colorectal cancers that display specific molecular features, namely, CpG island methylator phenotype, microsatellite instability, and BRAF mutation.122, 123, 124, 125, 126, 127 Smoking has been associated with genome-wide DNA methylation changes in blood leukocytes, which may imply its systemic effect on cellular epigenetic status.180, 181 Smoking has also been shown to be a strong risk factor for synchronous primary colorectal cancers (R Nishihara et al, unpublished data) and synchronous multiple polyps,182 especially serrated polyps,183 which are recognized as precursors for colorectal cancers with CpG island methylator phenotype, microsatellite instability and/or BRAF mutation.159, 184, 185, 186 CpG island methylator phenotype-high, microsatellite instability-high and BRAF mutation can co-occur in colorectal cancer,9, 30, 113, 184, 187, 188, 189, 190, 191, 192, 193, 194, 195 and are common characteristics of synchronous colorectal cancers.67, 68, 69, 70 Furthermore, synchronous primary colorectal cancers are considered to arise due to some form of predisposition, likely involving both genetic and environmental factors.196 However, there has been little evidence for conventional field effect in individuals with synchronous colorectal cancers, ie, CpG island methylator phenotype, microsatellite instability, and BRAF mutation are not found in normal colonic cells adjacent to synchronous colorectal cancers that demonstrate these somatic molecular aberrations.67 Furthermore, smoking has been consistently associated with BRAF-mutated colorectal cancer,122, 123, 124, 125, 126, 127 but not with KRAS-mutated colorectal cancer, where data are conflicting and complicated by publication bias.197, 198, 199, 200, 201 It is difficult to explain the gene specificity of mutations if one assumes the role of tobacco smoke as only a direct mutagen. Considering these pieces of evidence together, it seems plausible that smoking generates a field of tissue microenvironmental changes (as opposed to directly causing CpG island methylator phenotype, microsatellite instability or BRAF mutation), which may be advantageous for the growth of specific neoplastic/preneoplastic cells harboring BRAF mutation, but not so conducive to the grown of KRAS-mutated cells. The etiologic field of microenvironmental changes induced by smoking may predispose to the development of multiple cancers through similar carcinogenic pathways (Figure 1).
An example of evidence for an ‘etiologic field effect’ phenomenon. Smoking has been shown to increase the incidence of colorectal cancer displaying CpG island methylator phenotype, microsatellite instability, and BRAF mutation, as well as the incidence of synchronous colorectal cancers. Smoking, as an etiologic exposure, creates a field effect manifest as altered colonic tissue microenvironment. Microenvironmental change induced by smoking promotes carcinogenesis via specific pathways resulting in synchronous tumors with shared molecular features. As one might expect of an etiologic field, somatic alterations (such as CpG island methylator phenotype, microsatellite instability, and BRAF mutation) are absent from non-neoplastic tissue within the field.
Another example that profits from the adoption of an ‘etiologic field effect’ model is the association between early breast neoplasia (including lobular carcinoma in situ and atypical ductal hyperplasia) in one breast, and increased risk of subsequent invasive breast cancer in the same breast, and also the contralateral breast.75, 76 This phenomenon is likely related to tissue-specific gene-environmental interactions, which can be considered to constitute a type of field effect. The presence of an etiological field therefore adequately explains this observation; certain shared etiologic exposures lead to changes in the breast tissue microenvironment resulting in cancer susceptibility in both breasts.
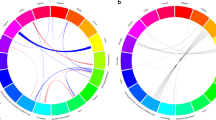
In Figure 2, using the colon as an example, we illustrate the temporal extent of ‘etiologic field effect’ concept and compare it with the conventional model of field effect. In the conventional model, cancers arise in a field of cells harboring acquired somatic molecular alterations. The ‘etiologic field effect’ model takes into account both host and exogenous factors, which, together, constitute a field of microenvironmental alterations and susceptibility to cancer development and progression.3, 90, 91
Comparison of conventional and etiologic field effect models, using colon cancer as an example. (a) In the conventional field effect model, a field of alteration ‘X’ in normal tissue (eg, colon) makes an individual prone to cancer development within that field. Normal colon in the field, and resulting synchronous tumors, show the same molecular alteration ‘X’. (b) In an ‘etiologic field effect’ model, etiologic factors (which can be multifactorial) generate a field of tissue microenvironmental changes, favoring the development of cancers through a common carcinogenic pathway characterized by molecular alteration ‘X’. In this setting, resulting synchronous tumors demonstrate the same alteration ‘X’, but normal tissue (eg, colon) need not necessarily display the same molecular feature.
Tumor initiation, evolution and progression: exploring the temporal limits of field effect
In contrast to the conventional field effect concept, which essentially pertains to molecular events associated with the initiating phases of neoplasia, ‘etiologic field effect’ extends temporally to incorporate biologic and physical etiologic factors that promote microenvironmental changes leading to cellular transformation, invasion, and metastasis (Figure 3). For example, etiologic factors that provoke or modulate inflammation (eg microbes, genetic polymorphisms, drugs, and dietary exposures) could contribute to an ‘etiologic field effect’ that remains influential at all stages of tumor evolution. Indeed, accumulating evidence on the anti-neoplastic effects of aspirin, and other inhibitors of PTGS2 (cyclooxygenase-2), supports exactly such a model; inflammatory processes, susceptible to the effects of these drugs, appear to be important in the early phases of neoplasia (eg colonic adenomagenesis),202, 203 during cancer evolution,204, 205, 206 and after cancer diagnosis,137, 138 possibly including the development of distant metastases.205 Tobacco smoke is a further example of an exposure that influences multiple phases of tumor evolution. In addition to a role in the initiating phases of bronchial carcinogenesis, components of tobacco smoke are implicated in promoting lung cancer growth and metastasis.207, 208 Similarly, cigarette smoke, a risk factor for breast cancer, may promote epithelial-mesenchymal transition and increase the metastatic potential of breast cancer cells.209
One of the principal differences between the ‘etiologic field effect’ and conventional field effect concepts is their temporal associations. The conventional field effect typically spans a relatively narrow part of neoplastic evolution spectrum, from the acquisition of somatic aberrations to histologically dysplastic pre-malignancy. The ‘etiologic field effect’, by comparison, is relevant at all stages from neoplasia initiation to patient death. The presence of ‘etiologic field effect’ precedes the acquisition of pathologic somatic changes, and extends to be influential in tumor evolution, invasion, and metastatic growth.
Tumor establishment at metastatic sites is dependent on physical cellular interactions and cross-talk between genetic, epigenetic, metabolomic and environmental factors occurring in the local tissue microenvironment.210 Thus, both tumor cell migration and the presence of a pro-metastatic microenvironmental niche, conducive to tumor seeding, could be ascribed to the presence of etiologic field effects.
The ‘etiologic field effect’ is not limited to epithelial cells, and embraces tumor–stromal interactions in the microenvironment, as well as macroenvironmental exposures and gene–environment interactions that effect microenvironmental change.33 Interestingly, tumor stroma and microenvironment may determine cancer molecular phenotype,211 and even response to molecularly targeted therapies.27, 212, 213 A small number of studies have highlighted the contribution of tumor stroma to field effect phenomena;20, 21, 22, 23, 24 however, the discussion in these studies20, 21, 22, 23, 24 has tended to be limited by the conventional notion of field effect.
Of note, ‘etiologic field effect’ phenomena do not need to be discrete from phenomena that constitute conventional field effect; rather, the two overlap within the spectrum of cancer predisposition. As with many features of biological systems, a ‘continuum’ model may afford a better representation of reality.78, 214, 215 Etiologic fields may seamlessly span multiple phases of neoplastic evolution, while their anatomic boundaries are likely to be gradients of tissue microenvironmental change, determined by variation in the magnitude of and sensitivity to a particular exposure.
Advantages and implications of the ‘etiologic field effect’ concept
The ‘etiologic field effect’ concept, as we perceive it, is attractive. First, it does not conflict or diminish the importance of the conventional notion of field effect; rather, the ‘etiologic field effect’ concept extends the temporal boundaries of the existing paradigm, to encompass the entire process of tumor evolution; exposure to etiologic factors generates a field of altered tissue interactome that remains influential throughout carcinogenesis, from initiation to progression and metastasis, and, ultimately, the demise of the patient. Second, the ‘etiologic field effect’ concept shifts the focus of attention from somatic genetic and epigenetic alterations to the influence of etiologic factors that might predispose to the acquisition of pathological molecular alterations in the first place. The ‘etiologic field effect’ therefore broadens the horizons of our inquiry into cancer susceptibility at molecular, cellular, and environmental levels.
Most cancer and pre-cancer surveillance protocols are based on the assumption of a persisting etiologic susceptibility. In clinical practice, modifiable components of etiological risk can be easily overlooked; in scheduling a polyp surveillance colonoscopy, we might miss opportunities for lifestyle interventions targeted at adenoma risk factors.216, 217 The term ‘etiologic field effect’ can successfully conceptualize the rather vague assumption of etiologic predisposition into a more concrete biomedical paradigm with a focus on risk modification and disease prevention. This concept can successfully act as model for dietary and lifestyle interventions, which have the potential to attenuate etiologic fields.
Epidemiologic and translational studies tend to use cancer occurrences or premalignant intermediaries (eg colorectal adenomas) as end points. For effective preventive strategies, we must strive to identify earlier markers of disease risk. Epidemiologic studies have linked low folate intake to increased incidence of colorectal adenomas and cancer.218, 219 It is also known that tissue folate status can modify molecular events in normal colonic mucosa.220 As not all individuals with low tissue folate will develop colorectal adenomas or cancer, the ability to interrogate the normal mucosal interactome for evidence of a pathological response to folate depletion could aid risk stratification, allowing for dietary or pharmacologic therapy to be instituted where appropriate. Biosensors of the presence and magnitude of ‘etiologic field effect’ could therefore facilitate the development of personalized prevention and treatment strategies.
An ‘etiologic field effect’ model may be informative for clinical and translational research. Clearly, the potential for etiologic field effect to act as a marker of the risk of cancer development, progression and/or metastasis constitutes an important area for future investigation. If etiologic field can act as a surrogate for disease risk, then evidence of etiological field could be used as an outcome measure in interventional studies of lifestyle modification or pharmacologic therapies. The possibility that reversal or modification of the etiologic field to a more ‘normal’ state can serve as a preventive strategy also demands scrutiny in future studies.
As described above, etiologic field effect can manifest as altered microenvironmental properties or abnormal patterns of cellular and tissue response to various endogenous and exogenous factors. We speculate that, in the future, the integrated efforts of ‘omics’ research, physical sciences, systems biology,2, 221 and nanotechnologies222 will help characterize complex molecular patterns (epigenomic, proteomic, metabolomic, etc) of host interactions with exogenous factors, giving rise to markers capable of indicating exposure to etiologic fields (Table 2). Markers of etiologic fields are likely to reflect altered exposome, with the exposome encompassing all exposures to which an individual is subjected.223 We would suggest that future biomarker discovery and validation efforts should focus on the identification of biosensors that signal the presence of disease susceptibility, rather than indicate established, albeit early pathologic changes. By means of illustration, recent data provide evidence for an epigenetic field of promoter methylation in normal colonic mucosa, involving genes distinct from those methylated in colorectal cancers.224 Furthermore, pathway analysis demonstrates that many of the differentially methylated genes are involved in carbohydrate metabolism, suggesting complex interactions between luminal contents and the gut microbiome and metabolome.
Several biosensors and nanotechnologies have already demonstrated the capacity for in vivo assessment of tissue ultrastructural and microvascular correlates of the genetic and epigenetic aberrations that define conventional field effects.222, 225, 226 Indeed, nano-cytology and nano-cytoarchitecture have been proposed as screening targets for field carcinogenesis.226, 227 It is therefore conceivable that the sophistication of future biosensors will enable accurate assessment of in vivo real-time changes in the microenvironmental interactome with high resolution (ie, in vivo molecular pathology). By detecting the very earliest alterations in the biologic or physical properties of tissues, exposure to detrimental etiological fields may be revealed long before the acquisition of irreversible molecular aberrations.
Conclusions
The existence of the conventional field effect is irrefutable, and is supported by an abundance of published research. Historically, advances in science and technology have allowed us to define field effects at increasingly earlier stages in carcinogenesis. The new paradigm of ‘etiologic field effect’ represents a further advancement of the field effect concept. This overarching interpretation of field effect has not previously been conceptualized or consolidated, as far as we are aware. Importantly, the ‘etiologic field effect’ is better positioned to explain several intriguing observations that are inadequately accounted for by the conventional interpretation of field effect. An important agenda for future research in complex diseases, such as cancer, is to decipher the complex interaction of etiologic exposures in an attempt to understand how they predispose to the acquisition of specific molecular aberrations and facilitate malignant transformation, tumor growth, migration, and metastasis. We believe that an etiologically focused, holistic, approach to the field effect paradigm can lead to a better understanding of cancer predisposition and progression, which, in turn, can facilitate the design of improved personalized cancer prevention and treatment strategies.
References
Marusyk A, Almendro V, Polyak K . Intra-tumor heterogeneity: a looking glass for cancer. Nat Rev Cancer 2012;12:323–334.
Gonzalez-Angulo AM, Hennessy BT, Mills GB . Future of personalized medicine in oncology: a systems biology approach. J Clin Oncol 2010;28:2777–2783.
Ogino S, Galon J, Fuchs CS et al. Cancer immunology-analysis of host and tumor factors for personalized medicine. Nat Rev Clin Oncol 2011;8:711–719.
Tjalsma H, Boleij A, Marchesi JR et al. A bacterial driver-passenger model for colorectal cancer: beyond the usual suspects. Nat Rev Microbiol 2012;10:575–582.
Baylin SB, Jones PA . A decade of exploring the cancer epigenome—biological and translational implications. Nat Rev Cancer 2011;11:726–734.
Schottenfeld D, Beebe-Dimmer JL, Buffler PA et al. Current perspective on the global and United States cancer burden attributable to lifestyle and environmental risk factors. Annu Rev Public Health 2013;34:97–117.
Topalian SL, Weiner GJ, Pardoll DM . Cancer immunotherapy comes of age. J Clin Oncol 2011;29:4828–4836.
Weitzel JN, Blazer KR, Macdonald DJ et al. Genetics, genomics, and cancer risk assessment: state of the art and future directions in the era of personalized medicine. CA Cancer J Clin 2011;61:327–359.
Myers MB, Wang Y, McKim KL et al. Hotspot oncomutations: implications for personalized cancer treatment. Expert Rev Mol Diagn 2012;12:603–620.
Sethi S, Ali S, Phillip PA et al. Clinical advances in molecular biomarkers for cancer diagnosis and therapy. Int J Mol Sci 2013;14:14771–14784.
Slaughter DP, Southwick HW, Smejkal W . Field cancerization in oral stratified squamous epithelium; clinical implications of multicentric origin. Cancer 1953;6:963–968.
Lippman SM, Hawk ET . Cancer prevention: from 1727 to milestones of the past 100 years. Cancer Res 2009;69:5269–5284.
Gazdar AF, Minna JD . Multifocal lung cancers—clonality vs field cancerization and does it matter? J Natl Cancer Inst 2009;101:541–543.
Giovannucci E, Ogino S . DNA methylation, field effects, and colorectal cancer. J Natl Cancer Inst 2005;97:1317–1319.
Ushijima T . Epigenetic field for cancerization. J Biochem Mol Biol 2007;40:142–150.
Dakubo GD, Jakupciak JP, Birch-Machin MA et al. Clinical implications and utility of field cancerization. Cancer Cell Int 2007;7:2.
Chai H, Brown RE . Field effect in cancer-an update. Ann Clin Lab Sci 2009;39:331–337.
Ramachandran K, Singal R . DNA methylation and field cancerization. Epigenomics 2012;4:243–245.
Dakubo GD, ed Field Cancerization: Basic Science and Clinical Applications. Nova Science Publishers: Hauppauge, NY, 2011.
Stearman RS, Dwyer-Nield L, Grady MC et al. A macrophage gene expression signature defines a field effect in the lung tumor microenvironment. Cancer Res 2008;68:34–43.
Martinez-Outschoorn UE, Balliet RM, Rivadeneira DB et al. Oxidative stress in cancer associated fibroblasts drives tumor-stroma co-evolution: A new paradigm for understanding tumor metabolism, the field effect and genomic instability in cancer cells. Cell Cycle 2010;9:3256–3276.
Ge L, Meng W, Zhou H et al. Could stroma contribute to field cancerization? Med Hypotheses 2010;75:26–31.
Hu B, Castillo E, Harewood L et al. Multifocal epithelial tumors and field cancerization from loss of mesenchymal CSL signaling. Cell 2012;149:1207–1220.
Vanharanta S, Massague J . Field cancerization: something new under the sun. Cell 2012;149:1179–1181.
Allen M, Louise Jones J . Jekyll and Hyde: the role of the microenvironment on the progression of cancer. J Pathol 2011;223:162–176.
Palmqvist R, Wikberg ML, Ling A et al. The association of immune cell infiltration and prognosis in colorectal cancer. Curr Colorectal Cancer Rep 2013;9:372–379.
Galon J, Mlecnik B, Bindea G et al. Towards the introduction of the Immunoscore in the classification of malignant tumors. J Pathol 2014;232:199–209.
Ogino S, Nosho K, Irahara N et al. Lymphocytic reaction to colorectal cancer is associated with longer survival, independent of lymph node count, microsatellite instability, and CpG island methylator phenotype. Clin Cancer Res 2009;15:6412–6420.
Nosho K, Baba Y, Tanaka N et al. Tumour-infiltrating T-cell subsets, molecular changes in colorectal cancer and prognosis: cohort study and literature review. J Pathol 2010;222:350–366.
Dahlin AM, Henriksson ML, Van Guelpen B et al. Colorectal cancer prognosis depends on T-cell infiltration and molecular characteristics of the tumor. Mod Pathol 2011;24:671–682.
Fridman WH, Pages F, Sautes-Fridman C et al. The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer 2012;12:298–306.
Edin S, Wikberg ML, Dahlin AM et al. The distribution of macrophages with a m1 or m2 phenotype in relation to prognosis and the molecular characteristics of colorectal cancer. PloS One 2012;7:e47045.
Thomas D . Gene–environment-wide association studies: emerging approaches. Nat Rev Genet 2010;11:259–272.
Ballard-Barbash R, Friedenreich CM, Courneya KS et al. Physical activity, biomarkers, and disease outcomes in cancer survivors: a systematic review. J Natl Cancer Inst 2012;104:815–840.
Vrieling A, Kampman E . The role of body mass index, physical activity, and diet in colorectal cancer recurrence and survival: a review of the literature. Am J Clin Nutr 2010;92:471–490.
Braakhuis BJM, Tabor MP, Kummer JA et al. A genetic explanation of Slaughter's concept of field cancerization: evidence and clinical implications. Cancer Res 2003;63:1727–1730.
Heaphy CM, Griffith JK, Bisoffi M . Mammary field cancerization: molecular evidence and clinical importance. Breast Cancer Res Treat 2009;118:229–239.
Franklin WA, Gazdar AF, Haney J et al. Widely dispersed p53 mutation in respiratory epithelium. A novel mechanism for field carcinogenesis. J Clin Invest 1997;100:2133–2137.
Prevo LJ, Sanchez CA, Galipeau PC et al. p53-mutant clones and field effects in Barrett's esophagus. Cancer Res 1999;59:4784–4787.
Rosenthal AN, Ryan A, Hopster D et al. Molecular evidence of a common clonal origin and subsequent divergent clonal evolution in vulval intraepithelial neoplasia, vulval squamous cell carcinoma and lymph node metastases. Int J Cancer 2002;99:549–554.
Chu TY, Shen CY, Lee HS et al. Monoclonality and surface lesion-specific microsatellite alterations in premalignant and malignant neoplasia of uterine cervix: a local field effect of genomic instability and clonal evolution. Genes Chromosomes Cancer 1999;24:127–134.
Heaphy CM, Bisoffi M, Fordyce CA et al. Telomere DNA content and allelic imbalance demonstrate field cancerization in histologically normal tissue adjacent to breast tumors. Int J Cancer 2006;119:108–116.
Takahashi T, Habuchi T, Kakehi Y et al. Clonal and chronological genetic analysis of multifocal cancers of the bladder and upper urinary tract. Cancer Res 1998;58:5835–5841.
Henson DE, Schwartz AM, Nsouli H et al. Carcinomas of the pancreas, gallbladder, extrahepatic bile ducts, and ampulla of vater share a field for carcinogenesis: a population-based study. Arch Pathol Lab Med 2009;133:67–71.
Davidson DD, Cheng L . ‘Field cancerization’ in the urothelium of the bladder. Anal Quant Cytol Histol 2006;28:337–338.
Shen L, Kondo Y, Rosner GL et al. MGMT promoter methylation and field defect in sporadic colorectal cancer. J Natl Cancer Inst 2005;97:1330–1338.
Issa JP, Ahuja N, Toyota M et al. Accelerated age-related CpG island methylation in ulcerative colitis. Cancer Res 2001;61:3573–3577.
Eads CA, Lord RV, Kurumboor SK et al. Fields of aberrant CpG island hypermethylation in Barrett's esophagus and associated adenocarcinoma. Cancer Res 2000;60:5021–5026.
Habuchi T, Takahashi T, Kakinuma H et al. Hypermethylation at 9q32-33 tumour suppressor region is age-related in normal urothelium and an early and frequent alteration in bladder cancer. Oncogene 2001;20:531–537.
Guo M, House MG, Hooker C et al. Promoter hypermethylation of resected bronchial margins: a field defect of changes? Clin Cancer Res 2004;10:5131–5136.
Yan PS, Venkataramu C, Ibrahim A et al. Mapping geographic zones of cancer risk with epigenetic biomarkers in normal breast tissue. Clin Cancer Res 2006;12:6626–6636.
Kamiyama H, Suzuki K, Maeda T et al. DNA demethylation in normal colon tissue predicts predisposition to multiple cancers. Oncogene 2012;31:5029–5037.
Asada K, Ando T, Niwa T et al. FHL1 on chromosome X is a single-hit gastrointestinal tumor-suppressor gene and contributes to the formation of an epigenetic field defect. Oncogene 2013;32:2140–2149.
Shigaki H, Baba Y, Watanabe M et al. LINE-1 hypomethylation in noncancerous esophageal mucosae is associated with smoking history. Ann Surg Oncol 2012;19:4238–4243.
Subramaniam MM, Loh M, Chan JY et al. The topography of DNA methylation in the non-neoplastic colonic mucosa surrounding colorectal cancers. Mol Carcinog 2013;53:98–108.
Kunte DP, Delacruz M, Wali RK et al. Dysregulation of MicroRNAs in colonic field carcinogenesis: implications for screening. PloS One 2012;7:e45591.
Goel A, Boland CR . Epigenetics of colorectal cancer. Gastroenterology 2012;143:1442–60 e1.
Hockel A, Dornhofer N . The hydra phenomenon of cancer: why tumors recur locally after microscopically complete resection. Cancer Res. 2005;65:2997–3002.
Gustafson AM, Soldi R, Anderlind C et al. Airway PI3K pathway activation is an early and reversible event in lung cancer development. Sci Transl Med 2010;2:26ra5.
Cheng L, Zhang S, Davidson DD et al. Molecular determinants of tumor recurrence in the urinary bladder. Future Oncol 2009;5:843–857.
Steiling K, Ryan J, Brody JS et al. The field of tissue injury in the lung and airway. Cancer Prev Res (Phila) 2008;1:396–403.
Kopelovich L, Henson DE, Gazdar AF et al. Surrogate anatomic/functional sites for evaluating cancer risk: an extension of the field effect. Clin Cancer Res 1999;5:3899–3905.
Jones TD, Wang M, Eble JN et al. Molecular evidence supporting field effect in urothelial carcinogenesis. Clin Cancer Res 2005;11:6512–6519.
Cheng L, Davidson DD, Maclennan GT et al. The origins of urothelial carcinoma. Expert Rev Anticancer Ther 2010;10:865–880.
Ogino S, Brahmandam M, kawasaki T et al. Epigenetic profiling of synchronous colorectal neoplasias by quantitative DNA methylation analysis. Mod Pathol 2006;19:1083–1090.
Talbott VA, Yeo CJ, Brody JR et al. Molecular profiling of synchronous and metachronous cancers of the pancreas reveal molecular mimicry between samples from the same patient. J Surg Res 2012;176:154–158.
Nosho K, Kure S, Irahara N et al. A prospective cohort study shows unique epigenetic, genetic, and prognostic features of synchronous colorectal cancers. Gastroenterology 2009;137:1609–1620 e3.
Konishi K, Shen L, Jelinek J et al. Concordant DNA methylation in synchronous colorectal carcinomas. Cancer Prev Res (Phila Pa) 2009;2:814–822.
Gonzalo V, Lozano JJ, Munoz J et al. Aberrant gene promoter methylation associated with sporadic multiple colorectal cancer. PloS One 2010;5:e8777.
Bae JM, Cho NY, Kim TY et al. Clinicopathologic and molecular characteristics of synchronous colorectal cancers: heterogeneity of clinical outcome depending on microsatellite instability status of individual tumors. Dis Colon Rectum 2012;55:181–190.
Begg CB . A strategy for distinguishing optimal cancer subtypes. Int J Cancer 2011;129:931–937.
Arpino G, Weiss HL, Clark GM et al. Hormone receptor status of a contralateral breast cancer is independent of the receptor status of the first primary in patients not receiving adjuvant tamoxifen. J Clin Oncol 2005;23:4687–4694.
Kollias J, Pinder SE, Denley HE et al. Phenotypic similarities in bilateral breast cancer. Breast Cancer Res Treat 2004;85:255–261.
Swain SM, Wilson JW, Mamounas EP et al. Estrogen receptor status of primary breast cancer is predictive of estrogen receptor status of contralateral breast cancer. J Natl Cancer Inst 2004;96:516–523.
Weitzel JN, Robson M, Pasini B et al. A comparison of bilateral breast cancers in BRCA carriers. Cancer Epidemiol Biomarkers Prev 2005;14:1534–1538.
Brown M, Bauer K, Pare M . Tumor marker phenotype concordance in second primary breast cancer, California, 1999-2004. Breast Cancer Res Treat 2010;120:217–227.
Zauber P, Huang J, Sabbath-Solitare M et al. Similarities of molecular genetic changes in synchronous and metachronous colorectal cancers are limited and related to the cancers' proximities to each other. J Mol Diagn 2013;15:652–660.
Berger AH, Knudson AG, Pandolfi PP . A continuum model for tumour suppression. Nature 2011;476:163–169.
Win AK, Young JP, Lindor NM et al. Colorectal and other cancer risks for carriers and noncarriers from families with a DNA mismatch repair gene mutation: a prospective cohort study. J Clin Oncol 2012;30:958–964.
Win AK, Lindor NM, Young JP et al. Risks of primary extracolonic cancers following colorectal cancer in Lynch syndrome. J Natl Cancer Inst 2012;104:1363–1372.
Lichtenstein P, Holm NV, Verkasalo PK et al. Environmental and heritable factors in the causation of cancer—analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med 2000;343:78–85.
Galvan A, Ioannidis JP, Dragani TA . Beyond genome-wide association studies: genetic heterogeneity and individual predisposition to cancer. Trends Genet 2010;26:132–141.
Slattery ML, Herrick J, Curtin K et al. Increased risk of colon cancer associated with a genetic polymorphism of SMAD7. Cancer Res 2010;70:1479–1485.
Garcia-Albeniz X, Nan H, Valeri L et al. Phenotypic and tumor molecular characterization of colorectal cancer in relation to a susceptibility SMAD7 variant associated with survival. Carcinogenesis 2013;34:292–298.
Ogino S, Hazra A, Tranah GJ et al. MGMT germline polymorphism is associated with somatic MGMT promoter methylation and gene silencing in colorectal cancer. Carcinogenesis 2007;28:1985–1990.
Hawkins NJ, Lee JH, Wong JJ et al. MGMT methylation is associated primarily with the germline C>T SNP (rs16906252) in colorectal cancer and normal colonic mucosa. Mod Pathol 2009;22:1588–1599.
Candiloro IL, Dobrovic A . Detection of MGMT promoter methylation in normal individuals is strongly associated with the T allele of the rs16906252 MGMT promoter single nucleotide polymorphism. Cancer Prev Res (Phila Pa) 2009;2:862–867.
Leng S, Bernauer AM, Hong C et al. The A/G allele of Rs16906252 predicts for MGMT methylation and is selectively silenced in premalignant lesions from smokers and in lung adenocarcinomas. Clin Cancer Res 2011;17:2014–2023.
Kristensen LS, Nielsen HM, Hager H et al. Methylation of MGMT in malignant pleural mesothelioma occurs in a subset of patients and is associated with the T allele of the rs16906252 MGMT promoter SNP. Lung Cancer 2011;71:130–136.
Ogino S, Stampfer M . Lifestyle factors and microsatellite instability in colorectal cancer: the evolving field of molecular pathological epidemiology. J Natl Cancer Inst 2010;102:365–367.
Ogino S, Chan AT, Fuchs CS et al. Molecular pathological epidemiology of colorectal neoplasia: an emerging transdisciplinary and interdisciplinary field. Gut 2011;60:397–411.
Herceg Z, Vaissiere T . Epigenetic mechanisms and cancer: an interface between the environment and the genome. Epigenetics 2011;6:804–819.
Kitkumthorn N, Mutirangura A . Long interspersed nuclear element-1 hypomethylation in cancer: biology and clinical applications. Clin Epigenet 2012;2:315–330.
Iwagami S, Baba Y, Watanabe M et al. Pyrosequencing assay to measure LINE-1 methylation level in esophageal squamous cell carcinoma. Ann Surg Oncol 2012;19:2726–2732.
Flanagan JM, Munoz-Alegre M, Henderson S et al. Gene-body hypermethylation of ATM in peripheral blood DNA of bilateral breast cancer patients. Hum Mol Genet 2009;18:1332–1342.
Brennan K, Flanagan JM . Is there a link between genome-wide hypomethylation in blood and cancer risk? Cancer Prev Res (Phila) 2012;5:1345–1357.
Amundadottir LT, Thorvaldsson S, Gudbjartsson DF et al. Cancer as a complex phenotype: pattern of cancer distribution within and beyond the nuclear family. PLoS Med 2004;1:e65.
Millikan RC, Newman B, Tse CK et al. Epidemiology of basal-like breast cancer. Breast Cancer Res Treat 2008;109:123–139.
Yang XR, Sherman ME, Rimm DL et al. Differences in risk factors for breast cancer molecular subtypes in a population-based study. Cancer Epidemiol Biomarkers Prev 2007;16:439–443.
Phipps AI, Malone KE, Porter PL, Daling JR, Li CI . Reproductive and hormonal risk factors for postmenopausal luminal, HER-2-overexpressing, and triple-negative breast cancer. Cancer 2008;113:1521–1526.
Dolle JM, Daling JR, White E et al. Risk factors for triple-negative breast cancer in women under the age of 45 years. Cancer Epidemiol Biomarkers Prev 2009;18:1157–1166.
Chen H, Taylor NP, Sotamaa KM et al. Evidence for heritable predisposition to epigenetic silencing of MLH1. Int J Cancer 2007;120:1684–1688.
Raptis S, Mrkonjic M, Green RC et al. MLH1 -93G>A promoter polymorphism and the risk of microsatellite-unstable colorectal cancer. J Natl Cancer Inst 2007;99:463–474.
Samowitz WS, Curtin K, Wolff RK et al. The MLH1 -93 G>A promoter polymorphism and genetic and epigenetic alterations in colon cancer. Genes Chromosomes Cancer 2008;47:835–844.
Allan JM, Shorto J, Adlard J et al. MLH1 -93G>A promoter polymorphism and risk of mismatch repair deficient colorectal cancer. Int J Cancer 2008;123:2456–2459.
Campbell PT, Curtin K, Ulrich CM et al. Mismatch repair polymorphisms and risk of colon cancer, tumour microsatellite instability and interactions with lifestyle factors. Gut 2009;58:661–667.
Oyama K, Kawakami K, Maeda K, Ishiguro K, Watanabe G . The association between methylenetetrahydrofolate reductase polymorphism and promoter methylation in proximal colon cancer. Anticancer Res 2004;24:649–654.
Curtin K, Slattery ML, Ulrich CM et al. Genetic polymorphisms in one-carbon metabolism: associations with CpG island methylator phenotype (CIMP) in colon cancer and the modifying effects of diet. Carcinogenesis 2007;28:1672–1679.
de Vogel S, Wouters KA, Gottschalk RW et al. Genetic variants of methyl metabolizing enzymes and epigenetic regulators: associations with promoter CpG island hypermethylation in colorectal cancer. Cancer Epidemiol Biomarkers Prev 2009;18:3086–3096.
Schernhammer ES, Giovannucci E, Kawasaki T et al. Dietary folate, alcohol, and B vitamins in relation to LINE-1 hypomethylation in colon cancer. Gut 2010;59:794–799.
Hazra A, Fuchs CS, Kawasaki T et al. Germline polymorphisms in the one-carbon metabolism pathway and DNA methylation in colorectal cancer. Cancer Causes Control 2010;21:331–345.
Van Guelpen B, Dahlin AM, Hultdin J et al. One-carbon metabolism and CpG island methylator phenotype status in incident colorectal cancer: a nested case-referent study. Cancer Causes Control 2010;21:557–566.
Curtin K, Samowitz WS, Ulrich CM et al. Nutrients in folate-mediated, one-carbon metabolism and the risk of rectal tumors in men and women. Nutr Cancer 2011;63:357–366.
Schernhammer ES, Giovannucci E, Baba Y, Fuchs CS, Ogino S . B vitamins, methionine and alcohol intake and risk of colon cancer in relation to BRAF mutation and CpG island methylator phenotype (CIMP). PloS One 2011;6:e21102.
Slattery ML, Curtin K, Anderson K et al. Associations between cigarette smoking, lifestyle factors, and microsatellite instability in colon tumors. J Natl Cancer Inst 2000;92:1831–1836.
Satia JA, Keku T, Galanko JA et al. Diet, lifestyle, and genomic instability in the north Carolina colon cancer study. Cancer Epidemiol Biomarkers Prev 2005;14:429–436.
Ogino S, Nosho K, Meyerhardt JA et al. Cohort study of fatty acid synthase expression and patient survival in colon cancer. J Clin Oncol 2008;26:5713–5720.
Campbell PT, Jacobs ET, Ulrich CM et al. Case-control study of overweight, obesity, and colorectal cancer risk, overall and by tumor microsatellite instability status. J Natl Cancer Inst 2010;102:391–400.
Kuchiba A, Morikawa T, Yamauchi M et al. Body mass index and risk of colorectal cancer according to fatty acid synthase expression in the nurses' health study. J Natl Cancer Inst 2012;104:415–420.
Sinicrope FA, Foster NR, Yoon HH et al. Association of obesity with DNA mismatch repair status and clinical outcome in patients with stage II or III colon carcinoma participating in NCCTG and NSABP adjuvant chemotherapy trials. J Clin Oncol 2012;30:406–412.
Hughes LA, Williamson EJ, van Engeland M et al. Body size and risk for colorectal cancers showing BRAF mutation or microsatellite instability: a pooled analysis. Int J Epidemiol 2012;41:1060–1072.
Samowitz WS, Albertsen H, Sweeney C et al. Association of smoking, CpG island methylator phenotype, and V600E BRAF mutations in colon cancer. J Natl Cancer Inst 2006;98:1731–1738.
Poynter JN, Haile RW, Siegmund KD et al. Associations between smoking, alcohol consumption, and colorectal cancer, overall and by tumor microsatellite instability status. Cancer Epidemiol Biomarkers Prev 2009;18:2745–2750.
Rozek LS, Herron CM, Greenson JK et al. Smoking, gender, and ethnicity predict somatic BRAF mutations in colorectal cancer. Cancer Epidemiol Biomarkers Prev 2010;19:838–843.
Limsui D, Vierkant RA, Tillmans LS et al. Cigarette smoking and colorectal cancer risk by molecularly defined subtypes. J Natl Cancer Inst 2010;102:1012–1022.
Curtin K, Samowitz WS, Wolff RK et al. Somatic alterations, metabolizing genes and smoking in rectal cancer. Int J Cancer 2009;125:158–164.
Nishihara R, Morikawa T, Kuchiba A et al. A prospective study of duration of smoking cessation and colorectal cancer risk by epigenetics-related tumor classification. Am J Epidemiol 2013;178:84–100.
Riely GJ, Kris MG, Rosenbaum D et al. Frequency and distinctive spectrum of KRAS mutations in never smokers with lung adenocarcinoma. Clin Cancer Res 2008;14:5731–5734.
Kawaguchi T, Ando M, Kubo A et al. Long exposure of environmental tobacco smoke associated with activating EGFR mutations in never-smokers with non-small cell lung cancer. Clin Cancer Res 2011;17:39–45.
Martinez ME, Maltzman T, Marshall JR et al. Risk factors for Ki-ras protooncogene mutation in sporadic colorectal adenomas. Cancer Res 1999;59:5181–5185.
Baba Y, Huttenhower C, Nosho K et al. Epigenomic diversity of colorectal cancer indicated by LINE-1 methylation in a database of 869 tumors. Mol Cancer 2010;9:125.
Goel A, Xicola RM, Nguyen TP et al. Aberrant DNA methylation in hereditary nonpolyposis colorectal cancer without mismatch repair deficiency. Gastroenterology 2010;138:1854–1862.
Antelo M, Balaguer F, Shia J et al. A high degree of LINE-1 hypomethylation is a unique feature of early-onset colorectal cancer. PloS One 2012;7:e45357.
Ogino S, Nishihara R, Lochhead P et al. Prospective study of family history and colorectal cancer risk by tumor LINE-1 methylation level. J Natl Cancer Inst 2013;105:130–140.
Ogino S, Nishihara R, Lochhead P et al. Prospective study of family history and colorectal cancer risk by tumor LINE-1 methylation level. J Natl Cancer Inst 2013;105:130–140.
Chan AT, Ogino S, Fuchs CS . Aspirin and the risk of colorectal cancer in relation to the expression of COX-2. N Engl J Med 2007;356:2131–2142.
Chan AT, Ogino S, Fuchs CS . Aspirin use and survival after diagnosis of colorectal cancer. JAMA 2009;302:649–658.
Liao X, Lochhead P, Nishihara R et al. Aspirin use, tumor PIK3CA mutation status, and colorectal cancer survival. N Engl J Med 2012;367:1596–1606.
Pasche B . Aspirin—from prevention to targeted therapy. N Engl J Med 2012;367:1650–1651.
Kang GH, Lee S, Kim WH et al. Epstein-barr virus-positive gastric carcinoma demonstrates frequent aberrant methylation of multiple genes and constitutes CpG island methylator phenotype-positive gastric carcinoma. Am J Pathol 2002;160:787–794.
Chang MS, Uozaki H, Chong JM et al. CpG island methylation status in gastric carcinoma with and without infection of Epstein-Barr virus. Clin Cancer Res 2006;12:2995–3002.
Ryan JL, Jones RJ, Kenney SC et al. Epstein-Barr virus-specific methylation of human genes in gastric cancer cells. Infect Agent Cancer 2010;5:27.
Ding SZ, Goldberg JB, Hatakeyama M . Helicobacter pylori infection, oncogenic pathways and epigenetic mechanisms in gastric carcinogenesis. Future Oncol 2010;6:851–862.
Peterson AJ, Menheniott TR, O'Connor L et al. Helicobacter pylori infection promotes methylation and silencing of trefoil factor 2, leading to gastric tumor development in mice and humans. Gastroenterology 2010;139:2005–2017.
Hernandez-Vargas H, Lambert MP, Le Calvez-Kelm F et al. Hepatocellular carcinoma displays distinct DNA methylation signatures with potential as clinical predictors. PloS One 2010;5:e9749.
Gao W, Kondo Y, Shen L et al. Variable DNA methylation patterns associated with progression of disease in hepatocellular carcinomas. Carcinogenesis 2008;29:1901–1910.
Vivekanandan P, Daniel HD, Kannangai R, Martinez-Murillo F, Torbenson M . Hepatitis B virus replication induces methylation of both host and viral DNA. J Virol 2010;84:4321–4329.
Ogino S, Lochhead P, Chan AT et al. Molecular pathological epidemiology of epigenetics: Emerging integrative science to analyze environment, host, and disease. Mod Pathol 2013;26:465–484.
Ogino S, Giovannucci E . Commentary: lifestyle factors and colorectal cancer microsatellite instability—molecular pathological epidemiology science, based on unique tumour principle. In J Epidemiol 2012;41:1072–1074.
Ogino S, King EE, Beck AH et al. Interdisciplinary education to integrate pathology and epidemiology: towards molecular and population-level health science. Am J Epidemiol 2012;176:659–667.
Ogino S, Beck AH, King EE et al. Ogino et al. respond to ‘The 21st century epidemiologist’. Am J Epidemiol 2012;176:672–674.
Ogino S, Fuchs CS, Giovannucci E . How many molecular subtypes? Implications of the unique tumor principle in personalized medicine. Expert Rev Mol Diagn 2012;12:621–628.
Curtin K, Slattery ML, Samowitz WS . CpG island methylation in colorectal cancer: past, present and future. Pathol Res Int 2011;2011:902674.
Hughes LA, Simons CC, van den Brandt PA et al. Body size, physical activity and risk of colorectal cancer with or without the CpG island methylator phenotype (CIMP). PloS One 2011;6:e18571.
Gehoff A, Basten O, Sprenger T et al. Optimal lymph node harvest in rectal cancer (UICC Stages II and III) after preoperative 5-FU-based radiochemotherapy. acetone compression is a new and highly efficient method. Am J Surg Pathol 2012;36:202–213.
Hughes LA, Khalid-de Bakker CA, Smits KM et al. The CpG island methylator phenotype in colorectal cancer: progress and problems. Biochim Biophys Acta 2012;1825:77–85.
Ku CS, Cooper DN, Wu M et al. Gene discovery in familial cancer syndromes by exome sequencing: prospects for the elucidation of familial colorectal cancer type X. Mod Pathol 2012;25:1055–1068.
Kanthan R, Senger JL, Kanthan SC . Molecular events in primary and metastatic colorectal carcinoma: a review. Pathol Res Int 2012;2012:597497.
Rex DK, Ahnen DJ, Baron JA et al. Serrated lesions of the colorectum: review and recommendations from an expert panel. Am J Gastroenterol 2012;107:1315–1329.
Dogan S, Shen R, Ang DC et al. Molecular epidemiology of EGFR and KRAS mutations in 3026 lung adenocarcinomas: higher susceptibility of women to smoking-related KRAS-mutant cancers. Clin Cancer Res 2012;18:6169–6177.
Galon J, Franck P, Marincola FM et al. Cancer classification using the Immunoscore: a worldwide task force. J Transl Med 2012;10:205.
Spitz MR, Caporaso NE, Sellers TA . Integrative cancer epidemiology—the next generation. Cancer Discov 2012;2:1087–1090.
Rosty C, Young JP, Walsh MD et al. Colorectal carcinomas with KRAS mutation are associated with distinctive morphological and molecular features. Mod Pathol 2013;26:825–834.
Campbell PT, Patel AV, Newton CC, Jacobs EJ, Gapstur SM . Associations of recreational physical activity and leisure time spent sitting with colorectal cancer survival. J Clin Oncol 2013;31:876–885.
Weijenberg MP, Hughes LA, Bours MJ et al. The mTOR pathway and the role of energy balance throughout life in colorectal cancer etiology and prognosis: unravelling mechanisms through a multidimensional molecular epidemiologic approach. Curr Nutr Rep 2013;2:19–26.
Alvarez MC, Santos JC, Maniezzo N et al. MGMT and MLH1 methylation in Helicobacter pylori-infected children and adults. World J Gastroenterol 2013;19:3043–3051.
Hagland HR, Berg M, Jolma IW, Carlsen A, Soreide K . Molecular pathways and cellular metabolism in colorectal cancer. Digestive Surg 2013;30:12–25.
Burnett-Hartman AN, Newcomb PA, Potter JD et al. Genomic aberrations occuring in subsets of serrated colorectal lesions but not conventional adenomas. Cancer Res 2013;73:2863–2872.
Zaidi N, Lupien L, Kuemmerle NB et al. Lipogenesis and lipolysis: the pathways exploited by the cancer cells to acquire fatty acids. Prog Lipid Res 2013;52:585–589.
Abbenhardt C, Poole EM, Kulmacz RJ et al. Phospholipase A2G1B polymorphisms and risk of colorectal neoplasia. Int J Mol Epidemiol Genet 2013;4:140–149.
Hughes LA, Melotte V, de Schrijver J et al. The CpG island methylator phenotype: what's in a name? Cancer Res 2013;73:5858–5868.
Amirian ES, Petrosino JF, Ajami NJ et al. Potential role of gastrointestinal microbiota composition in prostate cancer risk. Infect Agent Cancer 2013;8:42.
Bae JM, Kim JH, Cho NY, Kim TY, Kang GH . Prognostic implication of the CpG island methylator phenotype in colorectal cancers depends on tumour location. Br J Cancer 2013;109:1004–1012.
Ikramuddin S, Livingston EH . New insights on bariatric surgery outcomes. JAMA 2013;310:2401–2402.
Hoffmeister M, Blaker H, Kloor M et al. Body mass index and microsatellite instability in colorectal cancer: a population-based study. Cancer Epidemiol Biomarkers Prev 2013;22:2303–2311.
Zhu Y, Yang SR, Wang PP et al. Influence of pre-diagnostic cigarette smoking on colorectal cancer survival: overall and by tumour molecular phenotype. Br J Cancer 2014;110:1359–1366.
Larsson E, Tremaroli V, Lee YS et al. Analysis of gut microbial regulation of host gene expression along the length of the gut and regulation of gut microbial ecology through MyD88. Gut 2012;61:1124–1131.
Yamauchi M, Morikawa T, Kuchiba A et al. Assessment of colorectal cancer molecular features along bowel subsites challenges the conception of distinct dichotomy of proximal versus distal colorectum. Gut 2012;61:847–854.
Vidal M, Cusick ME, Barabasi AL . Interactome networks and human disease. Cell 2011;144:986–998.
Wan ES, Qiu W, Baccarelli A et al. Cigarette smoking behaviors and time since quitting are associated with differential DNA methylation across the human genome. Hum Mol Genet 2012;21:3073–3082.
Shenker NS, Polidoro S, van Veldhoven K et al. Epigenome-wide association study in the European Prospective Investigation into Cancer and Nutrition (EPIC-Turin) identifies novel genetic loci associated with smoking. Hum Mol Genet 2013;22:843–851.
Toyomura K, Yamaguchi K, Kawamoto H et al. Relation of cigarette smoking and alcohol use to colorectal adenomas by subsite: the self-defense forces health study. Cancer Sci 2004;95:72–76.
Buda A, De Bona M, Dotti I et al. Prevalence of different subtypes of serrated polyps and risk of synchronous advanced colorectal neoplasia in average-risk population undergoing first-time colonoscopy. Clin Transl Gasntroenterol 2012;3:e6.
Yamamoto E, Suzuki H, Yamano HO et al. Molecular dissection of premalignant colorectal lesions reveals early onset of the CpG Island methylator phenotype. Am J Pathol 2012;181:1847–1861.
Bardhan K, Liu K . Epigenetics and colorectal cancer pathogenesis. Cancers 2013;5:676–713.
Colussi D, Brandi G, Bazzoli F, Ricciardiello L . Molecular pathways involved in colorectal cancer: implications for disease behavior and prevention. Int J Mol Sci 2013;14:16365–16385.
Phipps AI, Buchanan DD, Makar KW et al. BRAF mutation status and survival after colorectal cancer diagnosis according to patient and tumor characteristics. Cancer Epidemiol Biomarkers Prev 2012;21:1792–1798.
Gavin P, Colangelo LH, Fumagalli D et al. Mutation profiling and microsatellite instability in stage II and III colon cancer: an assessment of their prognostic and oxaliplatin predictive value. Clin Cancer Res 2012;18:6531–6541.
Lao VV, Grady WM . Epigenetics and colorectal cancer. Nat Rev Gastroenterol Hepatol 2011;8:686–700.
Dahlin AM, Palmqvist R, Henriksson ML et al. The role of the CpG island methylator phenotype in colorectal cancer prognosis depends on microsatellite instability screening status. Clin Cancer Res 2010;16:1845–1855.
Zlobec I, Bihl M, Foerster A, Rufle A, Lugli A . Comprehensive analysis of CpG island methylator phenotype (CIMP)-high, -low, and -negative colorectal cancers based on protein marker expression and molecular features. J Pathol 2011;225:336–343.
Ang PW, Loh M, Liem N et al. Comprehensive profiling of DNA methylation in colorectal cancer reveals subgroups with distinct clinicopathological and molecular features. BMC Cancer 2010;10:227.
Wu C, Bekaii-Saab T . CpG island methylation, microsatellite instability, and BRAF mutations and their clinical application in the treatment of colon cancer. Chemother Res Prac 2012;2012:359041.
Beggs AD, Jones A, El-Bahwary M et al. Whole-genome methylation analysis of benign and malignant colorectal tumours. J Pathol 2013;229:697–704.
Zlobec I, Bihl MP, Foerster A, Rufle A, Lugli A . The impact of CpG island methylator phenotype and microsatellite instability on tumour budding in colorectal cancer. Histopathology 2012;61:777–787.
Leggett BA, Worthley DL . Synchronous colorectal cancer: not just bad luck? Gastroenterology 2009;137:1559–1562.
Naguib A, Mitrou PN, Gay LJ et al. Dietary, lifestyle and clinicopathological factors associated with BRAF and K-ras mutations arising in distinct subsets of colorectal cancers in the EPIC Norfolk study. BMC Cancer 2010;10:99.
Weijenberg MP, Aardening PW, de Kok TM, de Goeij AF, van den Brandt PA . Cigarette smoking and KRAS oncogene mutations in sporadic colorectal cancer: results from the Netherlands Cohort Study. Mutat Res 2008;652:54–64.
Diergaarde B, Vrieling A, van Kraats AA et al. Cigarette smoking and genetic alterations in sporadic colon carcinomas. Carcinogenesis 2003;24:565–571.
Slattery ML, Anderson K, Curtin K et al. Lifestyle factors and Ki-ras mutations in colon cancer tumors. Mutat Res 2001;483:73–81.
Samadder NJ, Vierkant RA, Tillmans LS et al. Cigarette smoking and colorectal cancer risk by KRAS mutation status among older women. Am J Gastroenterol 2012;107:782–789.
Bertagnolli MM, Eagle CJ, Zauber AG et al. Five-year efficacy and safety analysis of the Adenoma Prevention with Celecoxib Trial. Cancer Prev Res (Phila) 2009;2:310–321.
Benamouzig R, Uzzan B, Deyra J et al. Prevention by daily soluble aspirin of colorectal adenoma recurrence: 4-year results of the APACC randomised trial. Gut 2012;61:255–261.
Nishihara R, Lochhead P, Kuchiba A et al. Aspirin use and risk of colorectal cancer according to BRAF mutation status. JAMA 2013;309:2563–2571.
Rothwell PM, Wilson M, Price JF et al. Effect of daily aspirin on risk of cancer metastasis: a study of incident cancers during randomised controlled trials. Lancet 2012;379:1591–1601.
Algra AM, Rothwell PM . Effects of regular aspirin on long-term cancer incidence and metastasis: a systematic comparison of evidence from observational studies versus randomised trials. Lancet Oncol 2012;13:518–527.
Schaal C, Chellappan SP . Nicotine-mediated cell proliferation and tumor progression in smoking-related cancers. Mol Cancer Res 2014;12:14–23.
Yoshino I, Kometani T, Shoji F et al. Induction of epithelial-mesenchymal transition-related genes by Benzo[a]Pyrene in lung cancer cells. Cancer 2007;110:369–374.
Di Cello F, Flowers VL, Li HL et al. Cigarette smoke induces epithelial to mesenchymal transition and increases the metastatic ability of breast cancer cells. Mol Cancer 12:90 2013.
Wirtz D, Konstantopoulos K, Searson PC . The physics of cancer: the role of physical interactions and mechanical forces in metastasis. Nat Rev Cancer 2011;11:512–522.
Wu JM, Beck AH, Pate LL et al. Endogenous versus tumor-specific host response to breast carcinoma: a study of stromal response in synchronous breast primaries and biopsy site changes. Clin Cancer Res 2011;17:437–446.
Straussman R, Morikawa T, Shee K et al. Tumor microenvironment contributes to innate RAF-inhibitor resistance through HGF secretion. Nature 2012;487:500–504.
Reimers MS, Zeestraten EC, Kuppen PJ, Liefers GJ, van de Velde JH . Biomarkers in precision therapy in colorectal cancer. Gastroenterol Rep 2013;1:166–183.
Yamauchi M, Lochhead P, Morikawa T et al. Colorectal cancer: a tale of two sides or a continuum? Gut 2012;61:794–497.
Leedham S, Tomlinson I . The continuum model of selection in human tumors: general paradigm or niche product? Cancer Res 2012;72:3131–3134.
Stead M, Caswell S, Craigie AM, Eadie D, Anderson AS . Understanding the potential and challenges of adenoma treatment as a prevention opportunity: insights from the BeWEL formative study. Preven Med 2012;54:97–103.
Caswell S, Anderson AS, Steele RJ . Bowel health to better health: a minimal contact lifestyle intervention for people at increased risk of colorectal cancer. Br J Nutr 2009;102:1541–1546.
Kim DH, Smith-Warner SA, Spiegelman D et al. Pooled analyses of 13 prospective cohort studies on folate intake and colon cancer. Cancer Causes Control 2010;21:1919–1930.
Lee JE, Willett WC, Fuchs CS et al. Folate intake and risk of colorectal cancer and adenoma: modification by time. Am J Clin Nutr 2011;93:817–825.
McGlynn AP, Wasson GR, O'Reilly SL et al. Low colonocyte folate is associated with uracil misincorporation and global DNA hypomethylation in human colorectum. J Nutr 2013;143:27–33.
Michor F, Liphardt J, Ferrari M, Widom J . What does physics have to do with cancer? Nat Rev Cancer 2011;11:657–670.
Shanmuganathan R, Nazeema Banu B, Amirthalingam L et al. Conventional and nanotechniques for DNA methylation profiling. J Mol Diagn 2013;15:17–26.
Wild CP, Scalbert A, Herceg Z . Measuring the exposome: a powerful basis for evaluating environmental exposures and cancer risk. Environ Mol Mutagen 2013;54:480–499.
Silviera ML, Smith BP, Powell J, Sapienza C . Epigenetic differences in normal colon mucosa of cancer patients suggest altered dietary metabolic pathways. Cancer Prev Res (Phila) 2012;5:374–384.
Yang Q, Dong Y, Wu W et al. Detection and differential diagnosis of colon cancer by a cumulative analysis of promoter methylation. Nat Commun 2012;3:1206.
Damania D, Roy HK, Subramanian H et al. Nanocytology of rectal colonocytes to assess risk of colon cancer based on field cancerization. Cancer Res 2012;72:2720–2727.
Backman V, Roy HK . Light-scattering technologies for field carcinogenesis detection: a modality for endoscopic prescreening. Gastroenterology 2011;140:35–41.
Roberts NJ, Vogelstein JT, Parmigiani G et al. The predictive capacity of personal genome sequencing. Sci Transl Med 2012;4:133ra58.
Snyder M, Weissman S, Gerstein M . Personal phenotypes to go with personal genomes. Mol Systems Biol 2009;5:273.
Rivera CM, Ren B . Mapping human epigenomes. Cell 2013;155:39–55.
Monick MM, Beach SR, Plume J et al. Coordinated changes in AHRR methylation in lymphoblasts and pulmonary macrophages from smokers. Am J Med Genet B Neuropsychiatr Genet 2012;159B:141–151.
Zhang FF, Cardarelli R, Carroll J et al. Physical activity and global genomic DNA methylation in a cancer-free population. Epigenetics 2011;6:293–299.
Langevin SM, Houseman EA, Christensen BC et al. The influence of aging, environmental exposures and local sequence features on the variation of DNA methylation in blood. Epigenetics 2011;6:908–919.
Brody JS . Transcriptome alterations induced by cigarette smoke. Int J Cancer 2012;131:2754–2762.
De Coster S, van Leeuwen DM, Jennen DG et al. Gender-specific transcriptomic response to environmental exposure in Flemish adults. Environ Mol Mutagen 2013;54:574–588.
Beineke P, Fitch K, Tao H et al. A whole blood gene expression-based signature for smoking status. BMC Med Genomics 2012;5:58.
Pagmantidis V, Meplan C, van Schothorst EM, Keijer J, Hesketh JE . Supplementation of healthy volunteers with nutritionally relevant amounts of selenium increases the expression of lymphocyte protein biosynthesis genes. Am J Clin Nutr 2008;87:181–189.
Rappaport SM . Biomarkers intersect with the exposome. Biomarkers 2012;17:483–489.
Gonzalez LC . Protein microarrays, biosensors, and cell-based methods for secretome-wide extracellular protein-protein interaction mapping. Methods 2012;57:448–458.
Roy HK, Gomes A, Turzhitsky V et al. Spectroscopic microvascular blood detection from the endoscopically normal colonic mucosa: biomarker for neoplasia risk. Gastroenterology 2008;135:1069–1078.
Roy HK, Turzhitsky V, Kim Y et al. Association between rectal optical signatures and colonic neoplasia: potential applications for screening. Cancer Res 2009;69:4476–4483.
Shaheen NJ . Editorial: what is behind the remarkable increase in esophageal adenocarcinoma? Am J Gastroenterol 2014;109:345–347.
Brandstedt J, Wangefjord S, Nodin B et al. Associations of hormone replacement therapy and oral contraceptives with risk of colorectal cancer defined by clinicopathological factors, beta-catenin alterations, expression of cyclin D1, p53, and microsatellite-instability. BMC Cancer 2014;14:371.
Coppede F . The role of epigenetics in colorectal cancer. Expert Rev Gastroenterol Hepatol 2014, (e-pub ahead of print).
Bishehsari F, Mahdavinia M, Vacca M et al. Epidemiological transition of colorectal cancer in developing countries: environmental factors, molecular pathways, and opportunities for prevention. World J Gastroenterol 2014;20:6055–6072.
Acknowledgements
This work was supported by USA National Institute of Health (NIH) (R01 CA151993 (to SO), R01 CA137178 (to ATC), and P50 CA127003 (to CSF)). PL is a Scottish Government clinical academic fellow, and was supported by a Frank Knox Memorial Fellowship from Harvard University. ATC is a Damon Runyon Clinical Investigator. The content is solely the responsibility of the authors and does not necessarily represent the official views of NIH. Funding agencies did not have any role in the decision to submit the manuscript for publication, or the writing of the manuscript. The following standardized official symbols were used: HUGO (Human Genome Organization)-approved official symbols for genes and gene products, including BRAF, ERBB2, ESR1, FASN, KRAS, MGMT, MLH1, PGR, and PTGS2, all of which are described at www.genenames.org.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
ATC has consulted for Bayer Healthcare, Millennium Pharmaceuticals, and Pfizer. This work was not supported by Bayer Healthcare, Millennium Pharmaceuticals, or Pfizer. All other authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Lochhead, P., Chan, A., Nishihara, R. et al. Etiologic field effect: reappraisal of the field effect concept in cancer predisposition and progression. Mod Pathol 28, 14–29 (2015). https://doi.org/10.1038/modpathol.2014.81
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2014.81
This article is cited by
-
Early detection of lung cancer using artificial intelligence-enhanced optical nanosensing of chromatin alterations in field carcinogenesis
Scientific Reports (2023)
-
On Explaining Peto’s Paradox.
European Journal of Epidemiology (2023)
-
Nanomolar EP4 receptor potency and expression of eicosanoid-related enzymes in normal appearing colonic mucosa from patients with colorectal neoplasia
BMC Gastroenterology (2022)
-
MiR-509-3p is oncogenic, targets the tumor suppressor PHLPP2, and functions as a novel tumor adjacent normal tissue based prognostic biomarker in colorectal cancer
BMC Cancer (2022)
-
Promoter hypermethylation of GALR1 acts as an early epigenetic susceptibility event in colorectal carcinogenesis
Journal of Human Genetics (2022)