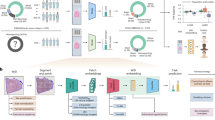
Abstract
A diagnosis of neuroendocrine carcinoma is often morphologically straight-forward; however, the tumor site of origin may remain elusive in a metastatic presentation. Neuroendocrine tumor subtyping has important implications for staging and patient management. In this study, the novel use and performance of a 92-gene molecular cancer classifier for determination of the site of tumor origin are described in a series of 75 neuroendocrine tumors (44 metastatic, 31 primary; gastrointestinal (n=12), pulmonary (n=22), Merkel cell (n=10), pancreatic (n=10), pheochromocytoma (n=10), and medullary thyroid carcinoma (n=11)). Formalin-fixed, paraffin-embedded samples passing multicenter pathologist adjudication were blinded and tested by a 92-gene molecular assay that predicts tumor type/subtype based upon relative quantitative PCR expression measurements for 87 tumor-related and 5 reference genes. The 92-gene assay demonstrated 99% (74/75; 95% confidence interval (CI) 0.93–0.99) accuracy for classification of neuroendocrine carcinomas and correctly subtyped the tumor site of origin in 95% (71/75; 95% CI 0.87–0.98) of cases. Analysis of gene expression subsignatures within the 92-gene assay panel showed 4 genes with promising discriminatory value for tumor typing and 15 genes for tumor subtyping. The 92-gene classifier demonstrated excellent accuracy for classifying and determining the site of origin in tumors with neuroendocrine differentiation. These results show promise for use of this test to aid in classifying neuroendocrine tumors of indeterminate primary site, particularly in the metastatic setting.
Similar content being viewed by others
Main
Neuroendocrine neoplasia encompasses a wide variety of organ sites, tumor grades, and clinicopathogical behaviors. Neuroendocrine differentiation can be recognized by morphological features along with immunohistochemistry (IHC). However, identification of the tumor site of origin in certain clinical contexts may be exceedingly challenging, due to the significant morphological and immunophenotypic similarity within this family of tumors. Clinical situations that may cause diagnostic subtyping challenges include: multiple possible primary tumors (in the context of Multiple Endocrine Neoplasia syndromes), widespread metastatic disease, lack of tumor-expressing site-specific serological markers and small biopsies with insufficient clinical data. The site of origin for neuroendocrine carcinoma has become increasingly important for grading/staging purposes,1, 2 for new clinical management guidelines,3, 4, 5, 6, 7 and for primary site-specific targeted therapy.8, 9, 10 When a primary site cannot be identified, tumors generally are treated according to the presumed aggressiveness of the tumor, as determined by a combination of the tumor grade11, 12, 13, 14 and available clinical and radiographic information about tumor metabolism.14
Because neuroendocrine carcinoma subtyping for site of origin is critical for clinical management, research has been dedicated to finding site and subtype-specific diagnostic markers.15, 16, 17, 18 Several biomarkers have been utilized to determine tumor subtype. eg, calcitonin is expressed by a majority of medullary thyroid carcinomas, and CK20 is characteristically expressed in Merkel cell carcinoma.15, 16 Some authors have proposed a panel of immunohistochemical stains (CDX-2, PDX-1, NESP-55, TTF-1, PAX8) to distinguish between differentiated gastrointestinal, pancreatic, and pulmonary neuroendocrine carcinoma with some success in tumor specificity;17, 18 however, this approach has seen limited utility in clinical practice due to relatively low sensitivities. Functional neuroendocrine carcinoma may have characteristic serology results and clinical presentation (eg, gastrinoma, insulinoma, etc.). Despite extensive clinicopathological investigation, however, it is estimated that up to 10% of patients with differentiated neuroendocrine carcinoma will have an unknown primary site.14
Traditionally, pathologists have used panels of protein-based immunohistochemical stains for tumor classification, but the potential advantage of molecular tests is readily apparent. In recent years, RNA expression-based classification of tumors has become an attractive standardized approach to aiding in the diagnosis and subclassification of tumors19, 20, 21, 22, 23, 24, 25 and for individualized therapy applications.26, 27, 28 In the coming years, gene expression-based analysis may serve as a complementary approach to genome-wide DNA analysis of tumors for individualized therapy.
A cohort of tumors with neuroendocrine differentiation was previously analyzed during a large-scale validation study of the 92-gene assay,20 an RT-PCR-based molecular cancer classifier. The larger validation study was designed to assess the overall performance of the 92-gene assay across 90 different tumor morphologies, comprising 28 different tumor types and 50 subtypes. The aim of the current study is to report detailed results on the performance of the 92-gene molecular classifier in subtyping neuroendocrine tumors and further highlight its potential diagnostic utility. In addition, exploratory analyses were conducted to examine and identify subsets of the 92-gene panel for specific use in neuroendocrine classification.
Materials and methods
Case Selection and Adjudication
Case selection criteria and methodology for the larger 92-gene classifier validation trial have been reported elsewhere; this trial included 1017 tumors from 28 tumor types and over 50 tumor subtypes.20 For the current study, we included all tumors (n=75) from the original trial that were considered to have neuroendocrine differentiation, including: Merkel cell carcinoma, medullary thyroid carcinoma, pheochromocytoma, paraganglioma, pulmonary neuroendocrine carcinoma (carcinoid, small cell carcinoma, large cell neuroendocrine carcinoma), pancreatic neuroendocrine carcinoma (all grades), and gastrointestinal neuroendocrine carcinoma (all grades; stomach, small intestine, appendix, and colorectum). Both primary and metastatic cases were included. Excluded were some sites of ‘epithelial’29 neuroendocrine tumors (thymus, pituitary, kidney, bladder, cervix, ovary), carcinomas with occult/mixed neuroendocrine differentiation, and most of the rarer “neural”29 types of neuroendocrine tumors (neuroblastoma, olfactory neuroblastoma, central nervous system primitive neuroectodermal tumors). Each case had been reviewed for diagnostic accuracy by consensus of two pathologists at different institutions. Case adjudication was performed by a primary pathologist through evaluation of clinical glass slides and available medical records and by a second pathologist who viewed a selected slide(s) by online whole slide digital imaging (Spectrum and ImageScope, Aperio Technologies, Vista, CA, USA) with clinicopathological information provided by the originating pathologist. Only adjudicated cases in which pathologists at both institutions agreed upon a consensus diagnosis for tumor type and subtype were included in the study (K=1). Cases were graded according to the grading criteria for each subtype as outlined in Klimstra et al1 and Hochwald et al30 using mitotic rate and tumor necrosis as applicable. These cases were selected and analyzed before the recent inclusion of Ki-67 in neuroendocrine tumor grading schemes, and thus this marker was not performed. Merkel cell carcinomas were considered grade 3. Grade 1 and 2 tumors were considered to be well-differentiated tumors, while grade 3 tumors were considered to be poorly differentiated. Medullary carcinomas and pheochromocytomas/paragangliomas were not graded.
Molecular Testing with the 92-Gene Assay
After passing diagnostic adjudication, a selected formalin-fixed, paraffin-embedded block was sectioned in RNAase-free conditions to produce one hematoxylin and eosin-stained section and three unstained 7-μ sections for molecular testing. The freshly prepared slides included only a research ID and were shipped to bioTheranostics (San Diego, CA, USA) for testing with only information on patient gender and biopsy site. Samples were macrodissected using the H&E-stained template or laser capture microdissected for tumor enrichment. Tumor enrichment after manual microdissection and laser-guided microdissection were >80% and >90%, respectively. Total RNA was extracted and DNase treated. First-strand cDNA was synthesized and then was pre-amplified (PreAmp, Life Technologies, Carlsbad, CA, USA). Real-time PCR was then performed using an ABI 7900HT instrument quantitatively measuring the expression of 87 tumor-associated genes and 5 reference genes as previously described21 and quality control parameters applied.20 The raw quantitative data was compared with a reference set of 2094 tumors (representing 28 main tumor types and 50 histological subtypes) for prediction of tumor type and subtype by proprietary statistical algorithm. Neuroendocrine tumor types/subtypes in the 92-gene assay panel include Adrenal—pheochromocytoma/paraganglioma, Neuroendocrine—skin (Merkel cell carcinoma), Neuroendocrine—lung low grade (pulmonary carcinoid), Neuroendocrine—lung high grade (pulmonary small cell carcinoma or large cell neuroendocrine carcinoma), Neuroendocrine—intestine (neuroendocrine tumors of all grades from the alimentary tract), Neuroendocrine—pancreas (pancreatic endocrine tumors), and Neuroendocrine—thyroid (medullary thyroid carcinoma). Quality control parameters for the 92-gene assay were described previously.20
Analysis of 92-Gene Panel Subsets
To select a gene subset for typing of neuroendocrine tumors, receiver operating characteristic analysis was performed for each of the 87 tumor-associated genes using 2094 tumors from the 92-gene assay reference database to assess their discriminatory power to differentiate neuroendocrine tumors (N=290) from non-neuroendocrine tumors (N=1804). Genes with the highest area under curve were chosen, and their performance in 957 cases from a blinded validation study was examined.20 The 75 neuroendocrine cases validated in this study do not overlap at all with the 290 cases from the training set in the reference database.
To identify a gene subset for subtyping neuroendocrine tumors, analysis of variance (ANOVA) was conducted for each of 87 genes using the 290 neuroendocrine tumors in the reference set. Genes with smallest P values were the ones that best distinguish the subtypes of neuroendocrine tumors and were thus selected as candidates for subtyping. The performance of the selected genes in the 75 neuroendocrine tumors from the validation study cohort was assessed by principal component analysis (PCA) and visualized in a three-dimensional plot using the first three principal components to examine the separation of different neuroendocrine subtypes.
Results
Performance Characteristics of the 92-Gene Classifier in Neuroendocrine Tumors
The performance of the 92-gene classifier in accurately classifying 28 tumor types and 50 subtypes has been reported.20 All 75 neuroendocrine tumors in this study met quality control parameters and were classified by the assay. The cohort included 44 females and 31 males, with a mean age of 62 years (range 29–86). Tumor characteristics are provided in Table 1. Cases were comprised of 59% metastatic tumors and 41% primary tumors. The most common biopsy site was liver, followed by lung and lymph node (Figure 1). The performance characteristics for the 92-gene assay predictions of neuroendocrine subtype are shown in Table 2. Assay sensitivities were 99% (95% CI: 0.93–0.99) for accurate classification of neuroendocrine tumors and 95% (95% CI:0.87–0.98) for identification of tumor subtype for the site of origin. Positive predictive values ranged from 0.83 to 1.00 for individual subtypes. A confusion matrix comparing the reference diagnosis with the 92-assay results is shown in Table 3; this highlights areas of concordance and discordance between the 92-gene classifier subtyped cases and reference diagnosis. The concordance rate of the molecular results with the reference diagnoses in poorly differentiated neuroendocrine carcinoma (grade 3 tumors) was 87% (20/23), whereas for well-differentiated neuroendocrine carcinoma (grade 1 and 2 tumors from the gastrointestinal tract, pancreas, or lung) it was 97% (30/31, see examples in Figure 2).
Distribution of biopsy sites. The height of the bars represents the number of cases included from each biopsy site. Dark gray bars indicate metastatic tumors to the sample site and light gray bars indicate primary tumors at the sample site. The most common sites were liver metastases and lung primaries followed by lymph node metastases.
Examples of well-differentiated neuroendocrine carcinomas classified correctly by the 92-gene assay, H&E, × 200. Top: Primary pulmonary carcinoid, prediction strength 96%. Middle: Pancreatic endocrine tumor metastatic to liver, prediction strength 96%. Bottom: Ileal carcinoid metastatic to the liver, prediction strength 96%.
Four cases had discordant 92-gene assay predictions compared with the reference diagnosis (Table 4). Three of the four cases were correctly predicted as neuroendocrine carcinoma but were discordant at the subtype (site of origin) level. Case 1 was adjudicated as an endobronchial pulmonary well-differentiated neuroendocrine (carcinoid) tumor with liver metastases at the time of primary diagnosis that was predicted by the assay to be a pancreatic endocrine primary. Case 2 was a pulmonary small cell carcinoma predicted to be Merkel cell carcinoma. Case 3 was a poorly differentiated pancreatic neuroendocrine carcinoma predicted to be a Merkel cell carcinoma. Case 4 was adjudicated as a poorly differentiated pancreatic neuroendocrine carcinoma and predicted to be a non-seminomatous germ cell tumor; however, a neuroendocrine tumor type was not ruled out by the assay (data not shown).
Exploratory Analysis of Neuroendocrine Gene Subsets
Further analysis was explored to potentially define a smaller subset of genes within the 92-gene assay panel with high sensitivities and specificities for neuroendocrine classification and subtyping. Of note, the genes encoding synaptophysin and chromogranin are not part of the 92-gene assay panel.21 Four genes demonstrated high discriminatory ability for distinguishing neuroendocrine from non-neuroendocrine tumor types in the assay reference set (N=2094), based on an area under curve cutoff of ≥0.8 from the receiver operating characteristic analysis. Consistently, area under curve values for these 4 genes were >0.8 in the 957 cases from the validation study cohort (Table 5). Biomarker utility for discrimination of neuroendocrine from non-neuroendocrine tumors can also be seen in the stripchart plots showing the distribution of expression values of each gene in each of the subtypes of neuroendocrine cases, as well as in non-neuroendocrine cases in the validation cohort (Figure 3).
The top 15 genes with significant P values from ANOVA analysis were selected as candidate genes to best distinguish different subtypes of neuroendocrine tumors in the reference set. These genes are described in more detail in Table 6 and include KIF2C,31, 32 SFTA3, CDCA3,33 KIF12,31, 34 CDH17,35, 36, 37, 38, 39 LOC100130899 (uncharacterized), NBLA00301,40, 41 HOXD11,42, 43, 44, 45, 46 EPS8L3, IRX3,47, 48 WWC1,49, 50, 51, 52, 53 HOXB8,54, 55, 56 FOXG1,57, 58, 59, 60, 61, 62 BCL11B,63, 64, 65 and LOC100506088 (uncharacterized). To visualize how well these 15 genes can distinguish neuroendocrine subtypes in the validation cohort, PCA were performed and the first three principal components were used to produce a three-dimensional plot showing the unsupervised clustering pattern of the different neuroendocrine subtypes (Figure 4). The PCA plot shows distinct separation of each neuroendocrine subtype and provides preliminary evidence that neuroendocrine subtyping may be feasible through optimization of the collective expression of the 15-gene set.
Discussion
Results from this blinded study demonstrate excellent performance of a 92-gene expression-based molecular classifier for prediction of tumor site of origin in a heterogeneous cohort of both primary and metastatic neuroendocrine tumors. These findings are particularly noteworthy, because neuroendocrine cases included in the current study were part of a larger validation trial20 that was completely blinded to tumor diagnosis, and especially pertinent here, neuroendocrine differentiation.
The 97% accuracy of the 92-gene assay for well-differentiated neuroendocrine tumors reported here is superior to published findings using IHC panels.17, 18 All well-differentiated neuroendocrine tumors from the gastrointestinal tract (12/12) and pancreas (8/8) and 91% (10/11) of pulmonary well-differentiated neuroendocrine tumors were correctly classified for the site of origin in our study; this included both metastatic and primary tumors. Correct identification of primary site in the metastatic setting is important, as treatment options and prognosis differ for thoracic-, pancreatic-, and gastrointestinal tract-based neuroendocrine tumors.4, 6, 7, 8, 9, 10, 66 Sangoi et al17 showed that IHC for PAX8 had only a 65% sensitivity for identifying pancreatic origin in well-differentiated neuroendocrine tumors metastatic to the liver, although it was 100% specific. Several cases of primary gastrointestinal neuroendocrine tumors in this study expressed PAX8. Long et al66 found similar results, with positive staining for PAX8 in only 50% of pancreatic neuroendocrine tumors metastatic to the liver, and with positive staining of all duodenal, 85% of rectal, and approximately 20% of appendiceal and gastric primary neuroendocrine tumors. In this study, the only gastrointestinal tumors metastatic to the liver that were tested for PAX8 were ileal tumors, which never showed any positive staining for PAX8 in the primary tumors. Srivastava et al18 demonstrated that an IHC panel including CDX2, PDX-1, NESP-55, and TTF-1 had limited performance for accurately predicting the primary site of gastrointestinal and pulmonary primary tumors, although it showed a sensitivity and specificity of 97% and 91%, respectively, for predicting pancreatic origin. In poorly differentiated tumors, the 92-gene assay showed rare discordant cases, but even in these diagnostically challenging cases the assay displayed an excellent overall performance overall of 87%.
Numerous pre-analytic and analytic factors can affect the reproducibility of protein IHC between and within labs. Some of the same pre-analytic factors affect both mRNA and proteins, including cold ischemic time, fixation methods and timings, tissue processing, and storage methods.67 Although mRNA degradation has been seen as more of a concern for mRNA than protein, both targets have been successfully adapted for formalin-fixed, paraffin-embedded processing. Protein IHC has additional variation in antibody retrieval methods and antibody clone differences, as well as wide variation in techniques and controls used between labs; all these factors can all affect the rate of false positive and false negative IHC results.68, 69, 70, 71, 72, 73, 74 In addition, selection of specific IHC markers and the interpretation of the protein expression patterns can be subjective and is dependent on the individual pathologist.
The strength of molecular diagnostics for tumor classification, including the 92-gene assay, lies both in standardized testing methods and in the comparison of gene expression between tumor samples and a well-adjudicated and robust expression database, provided rigorous quality controls are built into the assay process to evaluate specimens for appropriate mRNA integrity. Real-time, quantitative PCR for measurement of RNA expression is a standardized, highly reproducible, multiplexed panel of expression markers but with a logarithmically extended dynamic range of gene expression measurement not available in protein IHC. Because the signals are not directly visualized on tumor tissue, however, this assay is most effectively used with careful guidance by a pathologist to ensure sample selection for enrichment of tumor and exclusion of interfering normal cells (lymphocytes, fibroblasts, etc), a process of which is already growing rapidly within laboratories performing molecular oncology testing. The 92-gene assay routinely includes laser microdissection of tumor tissue, which is a key step that contributes to classification accuracy.19, 20
Feasibility analysis was performed to determine whether a subset of genes within the 92-gene biomarker panel could have value in predicting the site of origin for neuroendocrine tumors. Specifically, 15 genes were identified that showed reasonable discrimination between neuroendocrine tumors from different anatomic sites in this set of tested tumor samples. As their initial discovery was part of a data-driven process looking at differential gene expression across a diverse and wide variety of tumor types,21 and not for neuroendocrine typing in particular, mechanistic links to neuroendocrine differentiation or specific neuroendocrine tumor types are currently unknown. Many of the genes have been implicated in embryonic development, particularly in neuronal development (Table 6). Some genes have been implicated in minimally differentiated/embryonic-type tumors such as neuroblastoma and acute leukemias. The demonstration that the 15-gene subset of the 92-assay panel can discriminate differentiating tumor subtypes that are seemingly closely related supports the strength of using a collective gene expression profile for tumor subclassification. Future research on the specific function of these genes may provide additional insight into neuroendocrine tumor pathogenesis.
A limitation of this study was that it is a post hoc analysis from a larger validation trial that was powered to classify tumors from a much wider variety of tumor types and subtypes. For this reason, rarer neuroendocrine tumor types (eg, ovarian carcinoid, olfactory neuroblastoma, tumors with mixed/occult neuroendocrine differentiation) were not included. Additionally, this subset analysis also did not account for the prevalence of tumor subtypes in clinical practice. Further extension of the classifier to answer the question of prevalence effect may be particularly important for prospective trials of clinical and diagnostic utility in metastatic neuroendocrine carcinoma of unknown primary.
Findings from this study showed that there were four discrepancies between the 92-gene assay prediction and reference diagnosis. Review of these cases supports the reference diagnosis in all four cases based upon the adjudicated clinicopathological evidence by two pathologists at different institutions. In such situations, there always remains a question whether some of these cases could be incorrectly classified by current clinicopathological criteria or whether misclassification is due to n aberrant gene expression in the tested tumor. Practically, incorporation of both clinicopathological and molecular data (standardized gene expression data and targeted mutational analysis) will be required in order to properly classify patients for treatment with standard or targeted chemotherapies.
In conclusion, the 92-gene assay showed excellent accuracy for identifying neuroendocrine carcinomas and the neuroendocrine tumor site of origin in both well-differentiated and poorly differentiated tumors. This novel finding may have an immediate clinical application for distinguishing metastatic pancreatic endocrine tumors from other well-differentiated neuroendocrine carcinomas given that there are therapies (tyrosine kinase and mTOR inhibitors) specifically approved for pancreatic tumors.8, 9, 10 In addition, a subset of 15 genes within the 92-gene assay panel showed promising discriminatory ability to subclassify the tumor site of origin of neuroendocrine tumors. Further investigation of these genes may have diagnostic and theranostic value.
References
Klimstra DS, Modlin IR, Coppola D et al. The pathologic classification of neuroendocrine tumors: a review of nomenclature, grading, and staging systems. Pancreas 2010;39:707–712.
Scarpa A, Mantovani W, Capelli P et al. Pancreatic endocrine tumors: improved TNM staging and histopathological grading permit a clinically efficient prognostic stratification of patients. Mod Pathol 2010;23:824–833.
Anthony LB, Strosberg JR, Klimstra DS et al. The NANETS consensus guidelines for the diagnosis and management of gastrointestinal neuroendocrine tumors (nets): well-differentiated nets of the distal colon and rectum. Pancreas 2010;39:767–774.
Boudreaux JP, Klimstra DS, Hassan MM et al. The NANETS consensus guideline for the diagnosis and management of neuroendocrine tumors: well-differentiated neuroendocrine tumors of the Jejunum, Ileum, Appendix, and Cecum. Pancreas 2010;39:753–766.
Chen H, Sippel RS, O'Dorisio MS et al. The North American Neuroendocrine Tumor Society consensus guideline for the diagnosis and management of neuroendocrine tumors: pheochromocytoma, paraganglioma, and medullary thyroid cancer. Pancreas 2010;39:775–783.
Kulke MH, Anthony LB, Bushnell DL et al. NANETS treatment guidelines: well-differentiated neuroendocrine tumors of the stomach and pancreas. Pancreas 2010;39:735–752.
Phan AT, Oberg K, Choi J et al. NANETS consensus guideline for the diagnosis and management of neuroendocrine tumors: well-differentiated neuroendocrine tumors of the thorax (includes lung and thymus). Pancreas 2010;39:784–798.
Raymond E, Hobday T, Castellano D et al. Therapy innovations: tyrosine kinase inhibitors for the treatment of pancreatic neuroendocrine tumors. Cancer Metastasis Rev 2011;30:19–26.
Yao JC, Shah MH, Ito T et al. Everolimus for advanced pancreatic neuroendocrine tumors. N Engl J Med 2011;364:514–523.
Yao JC, Lombard-Bohas C, Baudin E et al. Daily oral everolimus activity in patients with metastatic pancreatic neuroendocrine tumors after failure of cytotoxic chemotherapy: a phase II trial. J Clin Oncol 2010;28:69–76.
Stoyianni A, Pentheroudakis G, Pavlidis N . Neuroendocrine carcinoma of unknown primary: a systematic review of the literature and a comparative study with other neuroendocrine tumors. Cancer Treat Rev 2011;37:358–365.
Strosberg JR, Coppola D, Klimstra DS et al. The NANETS consensus guidelines for the diagnosis and management of poorly differentiated (high-grade) extrapulmonary neuroendocrine carcinomas. Pancreas 2010;39:799–800.
Van Eeden S, Quaedvlieg PF, Taal BG et al. Classification of low-grade neuroendocrine tumors of midgut and unknown origin. Hum Pathol 2002;33:1126–1132.
Zuetenhorst JM, Taal BG . Metastatic carcinoid tumors: a clinical review. Oncologist 2005;10:123–131.
Cheuk W, Kwan MY, Suster S et al. Immunostaining for thyroid transcription factor 1 and cytokeratin 20 aids the distinction of small cell carcinoma from Merkel cell carcinoma, but not pulmonary from extrapulmonary small cell carcinomas. Arch Pathol Lab Med 2001;125:228–231.
Bobos M, Hytiroglou P, Kostopoulos I et al. Immunohistochemical distinction between merkel cell carcinoma and small cell carcinoma of the lung. Am J Dermatopathol 2006;28:99–104.
Sangoi AR, Ohgami RS, Pai RK et al. PAX8 expression reliably distinguishes pancreatic well-differentiated neuroendocrine tumors from ileal and pulmonary well-differentiated neuroendocrine tumors and pancreatic acinar cell carcinoma. Mod Pathol 2011;24:412–424.
Srivastava A, Hornick JL . Immunohistochemical staining for CDX-2, PDX-1, NESP-55, and TTF-1 can help distinguish gastrointestinal carcinoid tumors from pancreatic endocrine and pulmonary carcinoid tumors. Am J Surg Pathol 2009;33:626–632.
Erlander MG, Ma XJ, Kesty NC et al. Performance and clinical evaluation of the 92-gene real-time PCR assay for tumor classification. J Mol Diagn 2011;13:493–503.
Kerr SE, Schnabel CA, Sullivan PS et al. Multisite validation study to determine performance characteristics of a 92-gene molecular cancer classifier. Clin Cancer Res 2012;30:30.
Ma XJ, Patel R, Wang X et al. Molecular classification of human cancers using a 92-gene real-time quantitative polymerase chain reaction assay. Arch Pathol Lab Med 2006;130:465–473.
Perou CM, Sorlie T, Eisen MB et al. Molecular portraits of human breast tumours. Nature 2000;406:747–752.
Pillai R, Deeter R, Rigl CT et al. Validation and reproducibility of a microarray-based gene expression test for tumor identification in formalin-fixed, paraffin-embedded specimens. J Mol Diagn 2011;13:48–56.
Sorlie T, Perou CM, Tibshirani R et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 2001;98:10869–10874.
van Laar RK, Ma XJ, de Jong D et al. Implementation of a novel microarray-based diagnostic test for cancer of unknown primary. Int J Cancer 2009;125:1390–1397.
Eiermann W, Rezai M, Kummel S et al. The 21-gene recurrence score assay impacts adjuvant therapy recommendations for ER-positive, node-negative and node-positive early breast cancer resulting in a risk-adapted change in chemotherapy use. Ann Oncol 2012;7:7.
Mathieu MC, Mazouni C, Kesty NC et al. Breast Cancer Index predicts pathological complete response and eligibility for breast conserving surgery in breast cancer patients treated with neoadjuvant chemotherapy. Ann Oncol 2012;23:2046–2052.
van't Veer LJ, Dai H, van de Vijver MJ et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 2002;415:530–536.
Wick MR . Neuroendocrine neoplasia. Current concepts. Am J Clin Pathol 2000;113:331–335.
Hochwald SN, Zee S, Conlon KC et al. Prognostic factors in pancreatic endocrine neoplasms: an analysis of 136 cases with a proposal for low-grade and intermediate-grade groups. J Clin Oncol 2002;20:2633–2642.
Nakagawa T, Tanaka Y, Matsuoka E et al. Identification and classification of 16 new kinesin superfamily (KIF) proteins in mouse genome. Proc Natl Acad Sci USA 1997;94:9654–9659.
Wang W, Jiang Q, Argentini M et al. Kif2C minimal functional domain has unusual nucleotide binding properties that are adapted to microtubule depolymerization. J Biol Chem 2012;287:15143–15153.
Yoshida K . Cell-cycle-dependent regulation of the human and mouse Tome-1 promoters. FEBS Lett 2005;579:1488–1492.
Katoh M . Characterization of KIF12 gene in silico. Oncol Rep 2005;13:367–370.
Lee NP, Poon RT, Shek FH et al. Role of cadherin-17 in oncogenesis and potential therapeutic implications in hepatocellular carcinoma. Biochim Biophys Acta 2010;1806:138–145.
Morimatsu K, Aishima S, Kayashima T et al. Liver-intestine cadherin expression is associated with intestinal differentiation and carcinogenesis in intraductal papillary mucinous neoplasm. Pathobiology 2012;79:107–114.
Park JH, Seol JA, Choi HJ et al. Comparison of cadherin-17 expression between primary colorectal adenocarcinomas and their corresponding metastases: the possibility of a diagnostic marker for detecting the primary site of metastatic tumour. Histopathology 2011;58:315–318.
Wang J, Yu JC, Kang WM et al. The predictive effect of cadherin-17 on lymph node micrometastasis in pN0 gastric cancer. Ann Surg Oncol 2012;19:1529–1534.
Xu Y, Zhang J, Liu QS et al. Knockdown of liver-intestine cadherin decreases BGC823 cell invasiveness and metastasis in vivo. World J Gastroenterol 2012;18:3129–3137.
Voth H, Oberthuer A, Simon T et al. Identification of DEIN, a novel gene with high expression levels in stage IVS neuroblastoma. Mol Cancer Res 2007;5:1276–1284.
Voth H, Oberthuer A, Simon T et al. Co-regulated expression of HAND2 and DEIN by a bidirectional promoter with asymmetrical activity in neuroblastoma. BMC Mol Biol 2009;10:28.
Cai LY, Abe M, Izumi S et al. Identification of PRTFDC1 silencing and aberrant promoter methylation of GPR150, ITGA8 and HOXD11 in ovarian cancers. Life Sci 2007;80:1458–1465.
Goodman FR . Limb malformations and the human HOX genes. Am J Med Genet 2002;112:256–265.
Patterson LT, Pembaur M, Potter SS . Hoxa11 and Hoxd11 regulate branching morphogenesis of the ureteric bud in the developing kidney. Development 2001;128:2153–2161.
Taketani T, Taki T, Shibuya N et al. The HOXD11 gene is fused to the NUP98 gene in acute myeloid leukemia with t(2;11)(q31;p15). Cancer Res 2002;62:33–37.
Zha Y, Ding E, Yang L et al. Functional dissection of HOXD cluster genes in regulation of neuroblastoma cell proliferation and differentiation. PLoS One 2012;7:e40728.
Bellefroid EJ, Kobbe A, Gruss P et al. Xiro3 encodes a Xenopus homolog of the Drosophila Iroquois genes and functions in neural specification. EMBO J 1998;17:191–203.
Ragvin A, Moro E, Fredman D et al. Long-range gene regulation links genomic type 2 diabetes and obesity risk regions to HHEX, SOX4, and IRX3. Proc Natl Acad Sci USA 2010;107:775–780.
Hill VK, Dunwell TL, Catchpoole D et al. Frequent epigenetic inactivation of KIBRA, an upstream member of the Salvador/Warts/Hippo (SWH) tumor suppressor network, is associated with specific genetic event in B-cell acute lymphocytic leukemia. Epigenetics 2011;6:326–332.
Ji M, Yang S, Chen Y et al. Phospho-regulation of KIBRA by CDK1 and CDC14 phosphatase controls cell-cycle progression. Biochem J 2012;447:93–102.
Kauppi K, Nilsson LG, Adolfsson R et al. KIBRA polymorphism is related to enhanced memory and elevated hippocampal processing. J Neurosci 2011;31:14218–14222.
Makuch L, Volk L, Anggono V et al. Regulation of AMPA receptor function by the human memory-associated gene KIBRA. Neuron 2011;71:1022–1029.
Shinawi T, Hill V, Dagklis A et al. KIBRA gene methylation is associated with unfavorable biological prognostic parameters in chronic lymphocytic leukemia. Epigenetics 2012;7:211–215.
Greer JM, Capecchi MR . Hoxb8 is required for normal grooming behavior in mice. Neuron 2002;33:23–34.
Knoepfler PS, Sykes DB, Pasillas M et al. HoxB8 requires its Pbx-interaction motif to block differentiation of primary myeloid progenitors and of most cell line models of myeloid differentiation. Oncogene 2001;20:5440–5448.
Vider BZ, Zimber A, Hirsch D et al. Human colorectal carcinogenesis is associated with deregulation of homeobox gene expression. Biochem Biophys Res Commun 1997;232:742–748.
Adesina AM, Nguyen Y, Guanaratne P et al. FOXG1 is overexpressed in hepatoblastoma. Hum Pathol 2007;38:400–409.
Adesina AM, Nguyen Y, Mehta V et al. FOXG1 dysregulation is a frequent event in medulloblastoma. J Neurooncol 2007;85:111–122.
Dastidar SG, Narayanan S, Stifani S et al. Transducin-like enhancer of Split-1 (TLE1) combines with Forkhead box protein G1 (FoxG1) to promote neuronal survival. J Biol Chem 2012;287:14749–14759.
Kortum F, Das S, Flindt M et al. The core FOXG1 syndrome phenotype consists of postnatal microcephaly, severe mental retardation, absent language, dyskinesia, and corpus callosum hypogenesis. J Med Genet 2011;48:396–406.
Striano P, Paravidino R, Sicca F et al. West syndrome associated with 14q12 duplications harboring FOXG1. Neurology 2011;76:1600–1602.
Tian C, Gong Y, Yang Y et al. Foxg1 has an essential role in postnatal development of the dentate gyrus. J Neurosci 2012;32:2931–2949.
Gutierrez A, Kentsis A, Sanda T et al. The BCL11B tumor suppressor is mutated across the major molecular subtypes of T-cell acute lymphoblastic leukemia. Blood 2011;118:4169–4173.
Obata M, Kominami R, Mishima Y . BCL11B tumor suppressor inhibits HDM2 expression in a p53-dependent manner. Cell Signal 2012;24:1047–1052.
Zhang LJ, Vogel WK, Liu X et al. Coordinated regulation of transcription factor Bcl11b activity in thymocytes by the mitogen-activated protein kinase (MAPK) pathways and protein sumoylation. J Biol Chem 2012;287:26971–26988.
Long KB, Srivastava A, Hirsch MS et al. PAX8 Expression in well-differentiated pancreatic endocrine tumors: correlation with clinicopathologic features and comparison with gastrointestinal and pulmonary carcinoid tumors. Am J Surg Pathol 2010;34:723–729.
Medeiros F, Rigl CT, Anderson GG et al. Tissue handling for genome-wide expression analysis: a review of the issues, evidence, and opportunities. Arch Pathol Lab Med 2007;131:1805–1816.
Guerriere-Kovach PM, Hunt EL, Patterson JW et al. Primary melanoma of the skin and cutaneous melanomatous metastases: comparative histologic features and immunophenotypes. Am J Clin Pathol 2004;122:70–77.
Montone KT, van Belle P, Elenitsas R et al. Proto-oncogene c-kit expression in malignant melanoma: protein loss with tumor progression. Mod Pathol 1997;10:939–944.
Leong TY, Cooper K, Leong AS . Immunohistology—past, present, and future. Adv Anat Pathol 2010;17:404–418.
Bussolati G, Leonardo E . Technical pitfalls potentially affecting diagnoses in immunohistochemistry. J Clin Pathol 2008;61:1184–1192.
Leong AS . Pitfalls in diagnostic immunohistology. Adv Anat Pathol 2004;11:86–93.
Agoff SN, Lamps LW, Philip AT et al. Thyroid transcription factor-1 is expressed in extrapulmonary small cell carcinomas but not in other extrapulmonary neuroendocrine tumors. Mod Pathol 2000;13:238–242.
Kubba LA, McCluggage WG, Liu J et al. Thyroid transcription factor-1 expression in ovarian epithelial neoplasms. Mod Pathol 2008;21:485–490.
D'Alessandro V, Muscarella LA, la Torre A et al. Molecular analysis of the HuD gene in neuroendocrine lung cancers. Lung Cancer 2010;67:69–75.
Lee EK, Kim W, Tominaga K et al. RNA-binding protein HuD controls insulin translation. Mol Cell 2012;45:826–835.
Binda AV, Kabbani N, Levenson R. . Regulation of dense core vesicle release from PC12 cells by interaction between the D2 dopamine receptor and calcium-dependent activator protein for secretion (CAPS). Biochem Pharmacol 2005;69:1451–1461.
Hooks SB, Callihan P, Altman MK et al. Regulators of G-Protein signaling RGS10 and RGS17 regulate chemoresistance in ovarian cancer cells. Mol Cancer 2010;9:289.
James MA, Lu Y, Liu Y et al. RGS17, an overexpressed gene in human lung and prostate cancer, induces tumor cell proliferation through the cyclic AMP-PKA-CREB pathway. Cancer Res 2009;69:2108–2116.
Sun Y, Fang R, Li C et al. Hsa-mir-182 suppresses lung tumorigenesis through down regulation of RGS17 expression in vitro. Biochem Biophys Res Commun 2010;396:501–507.
Bonnefond A, Philippe J, Durand E et al. Whole-exome sequencing and high throughput genotyping identified KCNJ11 as the thirteenth MODY gene. PLoS One 2012;7:e37423.
Dupont J, Pereira C, Medeira A et al. Permanent neonatal diabetes mellitus due to KCNJ11 mutation in a Portuguese family: transition from insulin to oral sulfonylureas. J Pediatr Endocrinol Metab 2012;25:367–370.
Rubio-Cabezas O, Flanagan SE, Damhuis A et al. KATP channel mutations in infants with permanent diabetes diagnosed after 6 months of life. Pediatr Diabetes 2012;13:322–325.
Acknowledgements
We wish to thank Mary Till and David Dvorak for their efforts in data management and study coordination. We also thank W. Edward Highsmith, PhD and Veena Singh, MD for their expertise and guidance, and Brittany Carey, BS for her work during the primary 92-gene assay validation study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
CAS, YZ, VH, and ME are employees at bioTheranostics and have ownership interest (including patents) for bioTheranostics. PSS, SMD, and EFB have commercial research funding for bioTheranostics.
Rights and permissions
About this article
Cite this article
Kerr, S., Schnabel, C., Sullivan, P. et al. A 92-gene cancer classifier predicts the site of origin for neuroendocrine tumors. Mod Pathol 27, 44–54 (2014). https://doi.org/10.1038/modpathol.2013.105
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.105
Keywords
This article is cited by
-
Liver metastases from pituitary carcinomas mimicking visceral well-differentiated neuroendocrine tumors: a series of four cases
Diagnostic Pathology (2020)
-
Carcinoid Tumors: Past, Present, and Future
Indian Journal of Surgical Oncology (2020)
-
Ki67 Proliferative Index in Carcinoid Tumors Involving Ovary
Endocrine Pathology (2018)
-
Towards a new classification of gastroenteropancreatic neuroendocrine neoplasms
Nature Reviews Clinical Oncology (2016)
-
Poorly Differentiated Neoplasms of Unknown Primary Site: Diagnostic Usefulness of a Molecular Cancer Classifier Assay
Molecular Diagnosis & Therapy (2015)