Abstract
About 50% of all malignant peripheral nerve sheath tumors (MPNSTs) arise as neurofibromatosis type 1 associated lesions. In those patients malignant peripheral nerve sheath tumors are thought to arise through malignant transformation of a preexisting plexiform neurofibroma. The molecular changes associated with this transformation are still poorly understood. We sought to test the hypothesis that dysregulation of expression of kinases contributes to this malignant transformation. We analyzed expression of all 519 kinase genes in the human genome using the nanostring nCounter system. Twelve cases of malignant peripheral nerve sheath tumor arising in a background of preexisting plexiform neurofibroma were included. Both components were separately sampled. Statistical analysis compared global changes in expression levels as well as changes observed in the pairwise comparison of samples taken from the same surgical specimen. Immunohistochemical studies were performed on tissue array slides to confirm expression of selected proteins. The expression pattern of kinase genes can separate malignant peripheral nerve sheath tumors and preexisting plexiform neurofibromas. The majority of kinase genes is downregulated rather than overexpressed with malignant transformation. The patterns of expression changes are complex without simple recurring alteration. Pathway analysis demonstrates that differentially expressed kinases are enriched for kinases involved in the direct regulation of mitosis, and several of these show increased expression in malignant peripheral nerve sheath tumors. Immunohistochemical studies for the mitotic regulators BUB1B, PBK and NEK2 confirm higher expression levels at the protein level. These results suggest that the malignant transformation of plexiform neurofibroma is associated with distinct changes in the expression of kinase genes. The patterns of these changes are complex and heterogeneous. There is no single unifying alteration. Kinases involved in mitotic regulation are particularly enriched in the pool of differentially expressed kinases. Some of these are overexpressed and are therefore possible targets for kinase inhibitors.
Similar content being viewed by others
Main
Most malignant peripheral nerve sheath tumors (MPNSTs) arise as soft tissue malignancies. About half of all malignant peripheral nerve sheath tumors are thought to be sporadic de-novo tumors. The other half are neurofibromatosis type 1 (NF1) associated and develop through malignant transformation of a preexisting plexiform neurofibroma.1, 2
Malignant peripheral nerve sheath tumors often show complex karyotypes with losses and gains of chromosome arms as well as focal amplifications and deletions.3, 4, 5, 6 A number of miRNAs, cell cycle regulators, signaling molecules and transcription factors have been shown to be differentially expressed between neurofibromas and malignant peripheral nerve sheath tumors including CDKN2A, TP53, RB1, EGFR, CD44, PDGFRB, PDGFRA, HGF, MET, IGFR1, SOX9, SOX10, miR-34a and miR-21.3, 7, 8, 9, 10, 11, 12, 13, 14
Neurofibromatosis type 1 patients often have multiple plexiform neurofibromas. Their risk of developing a malignant peripheral nerve sheath tumor is estimated to be in the range from 3–13%.1, 3, 8, 15, 16 For these neurofibromatosis type 1 patients, this often becomes a life-limiting aspect of their condition as the 5-year survival rate of malignant peripheral nerve sheath tumor is ∼40%.10, 17, 18 The biological factors underlying this transformation are still poorly understood.3 Which of a patient’s often numerous plexiform neurofibromas, if any, will transform into a malignant peripheral nerve sheath tumor cannot be predicted.
The starting hypothesis of this study is that the malignant transformation from plexiform neurofibroma to malignant peripheral nerve sheath tumor is associated with changes in the expression profile of kinase genes. A few studies have performed cDNA microarray analysis to identify expression signatures of malignant peripheral nerve sheath tumor.9, 10, 12, 19 These studies have used frozen tissue or cell lines derived from tumors. They showed differences in the gene expression pattern between neurofibromas and unmatched malignant peripheral nerve sheath tumor samples from other patients. Neurofibromatosis type 1-associated and sporadic malignant peripheral nerve sheath tumors could not be distinguished based on their expression profile.10, 12
In this study, we chose a different approach by using a new analysis platform, the nCounter system, to assess the expression of several hundred genes.20, 21 This platform yields good results with small quantities of RNA isolated from formalin fixed paraffin embedded tissue without a prior amplification step. This allowed us to look for differences in gene expression in matched samples of plexiform neurofibroma and malignant peripheral nerve sheath tumor isolated from within the same surgical specimens.
Malignant tumors are often characterized by dysregulation of kinases. This dysregulation can occur because of changes in gene regulation, mutations or altered function. Neurofibromatosis type 1 results from mutations in the neurofibromin 1 gene that encodes a negative regulator of the Ras/MAP kinase signaling pathway. We hypothesized that there may be common global changes in the expression of kinases that occur with the malignant transformation of plexiform neurofibromas because these tumors start off with a shared genetic alteration in neurofibromin 1. Additionally, kinases are a possible target for novel therapeutics. Identification of kinases overexpressed in malignant peripheral nerve sheath tumors could suggest new treatment targets.
Methods
Case Selection
This study was performed with the approval of the institutional review board at the University of Chicago Medical Center. Cases of malignant peripheral nerve sheath tumor arising in known neurofibromatosis type 1 patients were identified. The pathologic slides were reviewed independently by two pathologists (TK and PP) to identify cases that included preexisting plexiform neurofibroma in addition to malignant peripheral nerve sheath tumor. A total of 12 cases containing malignant peripheral nerve sheath tumor and plexiform neurofibroma within the same surgical specimen were included in this study (Supplementary table 1). Representative areas of malignant peripheral nerve sheath tumor and plexiform neurofibroma were marked on the H&E-stained slides. Tissues from the respective areas were sampled with a 4-mm diameter biopsy punch allowing for exact sampling of areas with clearly defined morphologic features.
Different grading schemes are used for malignant peripheral nerve sheath tumors. The WHO system recognizes three grades (WHO II, III, IV). The French sarcoma system (FNCLCC) allows for the distinction of three grades on the basis of criteria different from those used in the WHO system. Others use a two-tier system distinguishing low-grade and high-grade malignant peripheral nerve sheath tumors. Owing to these problems with classification, we followed the approach outlined by Rodriguez et al.22 to separate these cases into low-grade and high-grade tumors (Figure 1). Details on the morphologic features are presented in Supplementary table 1. Strict criteria were used for the selection of neurofibroma areas to ensure that only unambiguous areas were included. All showed plexiform growth at low power. Mitotic activity, significant pleomorphism and significant cellularity were all absent (Figure 1).
Case selection. The gross and histologic (all H&E images, 20 × ) findings of representative cases included in the study are illustrated. (a) Case 11 was a patient with mass lesion that grew in a preexisting plexiform neurofibroma of the sciatic nerve (right end of the specimen). (b) and (c) show the H&E morphology of the plexiform neurofibroma (b) and malignant peripheral nerve sheath tumor (c) in the area that was sampled for the analysis with the biopsy punch. Most cases showed features of high-grade malignant peripheral nerve sheath tumor with abundant mitotic activity and necrosis (d). Cases 3 and 4 lacked distinct high grade features. They nevertheless showed unequivocal areas of plexiform neurofibroma (e) and malignant peripheral nerve sheath tumor (f) with several mitotic figures in the shown image, nuclear enlargement and increased cellularity.
RNA Isolation and Ncounter Analysis
Excess paraffin was trimmed from each tissue plug. Two small pieces were cut off each plug. These were processed in parallel and independently. Thus, a total of 48 samples were analyzed—12 malignant peripheral nerve sheath tumors in duplicate and the 12 matched plexiform neurofibromas in duplicate. RNA was purified using standard QIAGEN formalin-fixed paraffin embedded (FFPE) RNeasy kits following the manufacturer’s protocol. RNA concentration was quantified via nanodrop, and RNA quality was assessed with Agilent Bioanalyzer. As expected, the RNA was highly degraded, but the use of the Bioanalyzer allowed assessment of the mean fragment size.
The human kinase nCounter (Nanostring Technologies, Seattle, WA) assay was run according to manufacturers protocol. This probe set assays the expression of all 519 known human kinase genes. Briefly, the samples were randomized. Then, 100 ng of total RNA, the reporter codeset and the capture codeset were mixed and hybridized at 65 °C overnight using a PCR machine. The nCounter sample preparation station was prepared and run according to the manufacturer’s protocol. The samples were then scanned at high resolution (600 fields of view).
Normalization was performed in two separate steps: (1) Before the hybridization step, alien non-natural RNA sequences and probes directed to these were added to the samples at known concentrations. The readings for these control probes were used to normalize the data to account for differences in technical variables like variations in pipetting and hybridization. For this normalization, the geometric mean of levels of the control probes across all samples was calculated. This was then divided by the levels of the control probes found in individual samples to calculate a correction factor that the results of the respective case were multiplied by (Supplementary Figure 1A and B). (2) The utilized probe set also contains probes to eight defined housekeeping genes. These were used to correct for variations that may result from differences in RNA quality. The geometric mean for housekeeping genes within each sample was taken. A correction factor for the individual samples was then again calculated by dividing the overall expression of the housekeeping genes by that of the specific sample (Supplementary Figure 1C). Four samples were clearly outliers in the positive control normalization and were removed (correction factor >5). Four samples were clearly outliers in the housekeeping correction (correction factor >8) and were removed.
Gene Expression Analysis—Combined Sample Set and Pairwise Comparison
Two different types of statistical analysis were run: (1) The combined datasets of malignant peripheral nerve sheath tumors and plexiform neurofibromas were compared to look for differentially expressed genes. For each gene, P-values were calculated by Wilcoxon rank sum test, and false discovery rates were estimated by Storey’s q-value method.23 (2) Pairwise examination of the matched samples from the same case was performed to look for expression changes. For pairwise analysis, all samples were retained and corrected, and empirical Bayes t-tests, using limma, were performed for each individual, comparing plexiform neurofibroma with malignant peripheral nerve sheath tumor. For each gene, fold change was extracted from the table.
Analysis of Outside Gene Expression Data Set
Analysis of outside datasets: GEOquery was used to download GSE14038 into R. Expression levels for 13 plexiform neurofibromas and 6 malignant peripheral nerve sheath tumors were extracted from the dataset.9 To identify differentially expressed genes between plexiform neurofibromas and malignant peripheral nerve sheath tumors, surrogate variables were extracted with sva. Then, for each gene, we constructed a linear fixed effects model: y∼m+TumorType+e, where y is the residual following sva, m is the expected expression level, TumorType is the effect of tumor type and e is the residual error. Fixed effect P-values were estimated by F-test, and false discovery rates were estimated by Storey’s q-value method.23
Pathway Analysis
RefSeq annotations for the differentially expressed genes from both the nanostring dataset and GSE14038 were uploaded to the Database for Annotation, Visualization and Integrated Discovery (DAVID) 6.7 beta Bioinformatics Resources.24, 25, 26 This web-based bioinformatics platform allows the identification of functionally related groups of genes and genes that are part of the same functional pathway. DAVID contains a collection of pathways, ontologies terms and other categories. Given the background gene list (ie the genes assayed), DAVID can calculate which pathways show enrichment for differentially expressed genes. For the nanostring data, the 519 kinases were used as a background gene set, and DAVID’s Functional Annotation Chart and Functional Annotation clustering tools were used, with an ease score=0.1, to calculate fold enrichment. For GSE14038, all genes present on the array were used as a background list, and DAVID’s Functional Annotation Chart and Functional Annotation clustering tools were used, with an ease score=0.1, to calculate fold enrichment, p-value and Bonferroni corrected p-value. Additionally, Gene Set Enrichment Analysis was used to analyze the data from GSE14038.
Immunohistochemistry on Tissue Array Slides
Immunohistochemical staining was performed to validate differences in the expression on the protein level for selected candidates. Tissue array slides from a previously reported study were used.7 These include samples from 34 malignant peripheral nerve sheath tumors and 105 neurofibromas including 39 plexiform neurofibromas. The tissue array slides sample a separate set of tumors different from those cases used for the RNA expression studies. Each case was represented by at least two different cores in separate locations on the array. Commercially available antibodies against PBK, BUB1B and NEK2 were used (PBK: Cell Signaling, polyclonal rabbit antibody, cat. # 4942, 1:50; BUB1B: BD Transduction Laboratories, monoclonal mouse antibody, cat. # 612503, 1:50; NEK2: Abcam, monoclonal mouse antibody, cat. # ab55550, 1:75). The staining intensity was scored on a scale of 0–3.7 The staining of all cores representing a given case was averaged. Statistical analysis of the staining results was performed by t-test.
Results
Differentially Expressed Kinase Genes Result in Distinctive Signatures of Malignant Peripheral Nerve Sheath Tumors and Plexiform Neurofibromas
Ten of the twelve included malignant peripheral nerve sheath tumors were classified as high-grade and two as low-grade (Figure 1).22 Expression data for 519 kinases in the human genome was collected using the nanostring nCounter platform. The paired neurofibromas and malignant peripheral nerve sheath tumors were analyzed in duplicate, and data was normalized using both positive control probes as well as housekeeping genes. Unsupervised clustering and principal component analysis (Figure 2a) of the normalized expression data demonstrate that the expression pattern of the analyzed kinase genes can separate the malignant peripheral nerve sheath tumor samples from the plexiform neurofibroma samples. Two of the malignant peripheral nerve sheath tumor samples (case 3 and case 4) cluster together with the plexiform neurofibroma samples. Interestingly, these were the two malignant peripheral nerve sheath tumors that lacked high-grade features on the histopathologic analysis. The distance matrix (Figure 2b) also illustrates the same separation of the data set into two distinct groups—the high-grade malignant peripheral nerve sheath tumors and the plexiform neurofibromas. The two lower grade tumors again cluster together with the plexiform neurofibromas.
Distinct clustering of malignant peripheral nerve sheath tumors (MPNST, MP) and plexiform neurofibromas (NF, N) in principal component analysis and distance matrix. (a) The principal component analysis shows that the global expression profile of the samples distinguishes between malignant peripheral nerve sheath tumors (shown in blue) and plexiform neurofibromas (shown in red). Two diagrams showing separation with principal component 1 and 2 as well as principal component 2 and 3 are included. The two outliers are the two lower grade malignant peripheral nerve sheath tumors, cases 3 and 4, that cluster together with the plexiform neurofibromas in this analysis. Duplicates run on the same tissue core often cluster closely together. (b) The distance matrix plot is based on the pairwise comparison of samples, and groups them together depending on how closely they resemble each other in their gene expression profile. This analysis is again able to separate malignant peripheral nerve sheath tumors from plexiform neurofibromas. The two lower grade malignant peripheral nerve sheath tumors again cluster with the plexiform neurofibromas.
A Distinct Set of Genes is Differentially Expressed between Malignant Peripheral Nerve Sheath Tumors and Plexiform Neurofibromas Including Genes Implicated in Mitotic Regulation
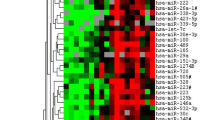
281 genes were differentially expressed in the tumor samples, 267 of these were downregulated in malignant peripheral nerve sheath tumors, 16 were upregulated (FDR ≤0.005; Supplementary table 2). The expression of these genes shows distinctly different signatures in the malignant peripheral nerve sheath tumors compared with the plexiform neurofibromas. A heatmap generated using all samples remaining in this analysis visually illustrates how most of these 281 differentially expressed kinase genes are downregulated in malignant peripheral nerve sheath tumors, not overexpressed (Figure 3a). However, there was a small set of upregulated kinase genes in the malignant peripheral nerve sheath tumors. These almost exclusively included kinases involved in mitotic regulation. Indeed, pathway analysis using the Database for Annotation, Visualization and Integrated Discovery (DAVID) identified genes important for the regulation of mitotic cycle/chromosome segregation as the biggest functional group among the differentially expressed genes. This smaller set of genes provided similar separation of malignant peripheral nerve sheath tumors and plexiform neurofibromas as that observed for all differentially expressed genes (Figure 3b). The majority of these genes again showed downregulation with progression to malignant peripheral nerve sheath tumor. However, several mitosis checkpoint kinases were upregulated with malignant transformation, including BUB1, BUB1B, PBK, TTK, CHEK1 and NEK2. Interestingly, several of these genes (BUB1, BUB1B, TTK and NEK2) are involved in the kinetochore checkpoint and are frequently overexpressed in many cancers, particularly those with significant aneuploidy.
Malignant peripheral nerve sheath tumors (MPNST) and plexiform neurofibromas (NF) are characterized by distinctive expression profiles. Kinases involved in mitotic regulation are differentially expressed. (a) 281 genes were identified as differentially expressed at statistically significant levels between malignant peripheral nerve sheath tumor and plexiform neurofibroma samples based on the Wilcoxon test (with FDR ≤0.005). This heatmap illustrates the expression of these genes (green: upregulation, red: downregulation) and shows clear separations of the sample sets into malignant peripheral nerve sheath tumors and plexiform neurofibromas based on the expression level of these 281 genes. The majority of kinase genes are downregulated in malignant peripheral nerve sheath tumors rather than overexpressed. (b) Pathway analysis identified kinases involved in mitotic regulation as the most prominent functional group of differentially expressed genes. This smaller gene set shows similar separation of the samples into malignant peripheral nerve sheath tumors and plexiform neurofibromas (green: upregulation, red: downregulation). Again, downregulation of kinase genes rather than upregulation is the dominant change.
Pairwise Comparison Identifies Heterogeneous Changes in Gene Expression Profiles with Recurring Patterns
The design of the experiment, using matched pairs of plexiform neurofibromas and malignant peripheral nerve sheath tumors, offered the opportunity to investigate how each individual tumor altered kinase expression with the malignant transformation from plexiform neurofibroma to malignant peripheral nerve sheath tumor.
Fifty-five genes were significantly downregulated (FDR 0.05) in at least 10 of 12 samples (Supplementary table 2) in the pairwise comparison of plexiform neurofibroma and malignant peripheral nerve sheath tumor components obtained from the same surgical specimen. These include the tumor suppressor TGFB2R that is strongly downregulated in all 12 tumor samples. Additionally, several receptor tyrosine kinases, including ERBB2, ERBB3 and NTRK3, were strongly downregulated in most if not all tumors. Receptor tyrosine kinases are an important class of kinases that bind a ligand and mediate signaling across the cell membrane. Expression of receptor tyrosine kinases is frequently altered in cancers, and many of them represent potential therapeutic targets. 54 of the 58 receptor tyrosine kinases in the human genome were expressed at measurable levels in our data set. Malignant peripheral nerve sheath tumors showed downregulation of most receptor tyrosine kinases. However, a few of these were overexpressed in each of the samples (Supplementary Figure 2), with each pair showing a different pattern of upregulation. Fourteen of these fifty-four receptor tyrosine kinases were differentially expressed between malignant peripheral nerve sheath tumors and plexiform neurofibromas at statistically significant levels when analyzing all samples (Figure 4). The vast majority of these receptor tyrosine kinases were strongly downregulated in most samples.
Most differentially expressed receptor tyrosine kinases are downregulated in malignant peripheral nerve sheath tumors (MPNST). This diagram illustrates the changes in expression of those 14 receptor tyrosine kinase genes that are differentially expressed at a statistically significant level between malignant peripheral nerve sheath tumor and plexiform neurofibroma in the pairwise comparison. For each of the 12 included malignant peripheral nerve sheath tumors, the relative fold change in expression compared with the respective plexiform neurofibroma is plotted (downregulation to the left/light grey, overexpression to the right/dark grey).
Neurofibromatosis type 1 is defined by mutation in neurofibromin 1, which encodes a GTPase activating protein that negatively regulates the Ras signaling pathway. Thus, we hypothesized that expression of kinases in the MAP kinase cascade may be altered in the development of malignant peripheral nerve sheath tumor arising from plexiform neurofibroma. This analysis showed that most of the kinases that are part of the MAP kinase signaling cascade tend to be downregulated in malignant peripheral nerve sheath tumors compared with the preexisting neurofibromas. 15 kinase genes of this signaling cascade were differentially expressed between plexiform neurofibromas and malignant peripheral nerve sheath tumors at statistically significant levels in the pairwise comparison, when analyzing all samples (Figure 5). Interestingly, the ERK kinases (MAPK1 and MAPK3) are downregulated in malignant peripheral nerve sheath tumors, although still highly expressed. MAP3K6 and MAP3K8, which are MAP kinase kinase kinases that signal through the stress-activated JNK cascade, were also strongly downregulated in most tumors.
Most differentially expressed genes encoding components of the MAP kinase signaling cascade are downregulated in malignant peripheral nerve sheath tumors (MPNST). This diagram illustrates the changes in expression of those 15 kinase genes of the MAP kinase signaling cascade that are differentially expressed at a statistically significant level between malignant peripheral nerve sheath tumors and plexiform neurofibroma in the pairwise comparison. For each of the 12 included malignant peripheral nerve sheath tumors, the relative fold change in expression compared with the respective plexiform neurofibroma is plotted (downregulation to the left/light grey, overexpression to the right/dark grey).
Only a small set of kinase genes were identified as being consistently overexpressed in several samples (Supplementary table 2). Only 12 genes were upregulated in at least five samples and showed downregulation in no more than two of the other samples. Interestingly this short list includes several of those kinases that were also found to be overexpressed in malignant peripheral nerve sheath tumors based on the Wilcoxon test (eg PBK, PTK7, BUB1B, CHEK1 and NEK2).
Analysis of Outside Data Set
We next analyzed another, previously produced malignant peripheral nerve sheath tumor expression dataset. In this dataset, we found strong downregulation of TGFBR2 (data not shown) and upregulation of BUB1B, BUB1, NEK2 and PBK (Figure 6a). The expansion of the gene lists from the microarray allowed deeper analysis of differential gene expression by Gene Set Enrichment Analysis and pathways by the Database for Annotation, Visualization and Integrated Discovery (DAVID). Indeed, we found that mitotic genes, particularly those involved in the spindle checkpoint, were differentially expressed between malignant peripheral nerve sheath tumor and plexiform neurofibromas (Figure 7). Moreover, Gene Set Enrichment Analysis identified several gene modules involving mitotic checkpoint proteins as differentially expressed between plexiform neurofibroma and malignant peripheral nerve sheath tumor.
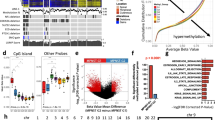
Additional studies validate differential expression of BUB1B, NEK2 and PBK—outside data set and immunohistochemical staining. (a) Analysis of previously published gene expression microarray data also shows significantly higher expression of BUB1B, NEK2 and PBK in those unmatched samples of malignant peripheral nerve sheath tumor and neurofibroma. For all three, the p-value was P<0.01. Immunohistochemical staining for BUB1B (b–e): Representative images of malignant peripheral nerve sheath tumors positive for BUB1B (b and c) are shown as well as a neurofibroma negative for BUB1B (d). (e) The intensity score shows significantly stronger expression in the malignant peripheral nerve sheath tumors compared with all neurofibromas (P<0.001) as well as compared with only plexiform neurofibromas (P<0.001). Immunohistochemical staining for NEK2 (f, i): Representative images of malignant peripheral nerve sheath tumors with positive nuclear staining for NEK2 (f), positive cytoplasmic staining for NEK2 (g) and mixed nuclear/cytoplasmic staining for NEK 2 are shown (h). (i) The intensity score shows significantly stronger expression in the malignant peripheral nerve sheath tumors compared with all neurofibromas (P<0.001), as well as compared with only plexiform neurofibromas (P<0.001). Immunohistochemical staining for PBK (j–m): Representative images of malignant peripheral nerve sheath tumors positive for PBK (j and k) are shown as well as a neurofibroma with positive staining for PBK (l). (m) The intensity score shows significantly stronger expression in the malignant peripheral nerve sheath tumors compared with all neurofibromas (P<0.001) as well as compared to only plexiform neurofibromas (P<0.001).
Pathway analysis. Gene Set Enrichment Analysis (GSEA) of malignant peripheral nerve sheath tumors (MPNSTs) and neurofibromas (NFs) from GSE14038 demonstrates enrichment for higher expression of genes related the G2-M checkpoint and chromosomal condensation in malignant peripheral nerve sheath tumors. For example, genes in the expression neighborhood of BUB1B and CENPF are highly upregulated in malignant peripheral nerve sheath tumors. Additionally, many genes in the G2-M checkpoint and the reactome mitotic prometaphase are upregulated in malignant peripheral nerve sheath tumors.
Progression to Malignant Peripheral Nerve Sheath Tumor Goes Along with the Increased Expression of BUB1B, NEK2 and PBK at the Protein Level
Immunohistochemical studies were performed as an additional validation step to assess the expression of selected differentially expressed genes at the protein level. These immunohistochemical studies focused on kinase implicated in mitotic regulation that we found to be upregulated in malignant peripheral nerve sheath tumors based on the expression data. They confirmed that malignant peripheral nerve sheath tumors showed significantly more expression of NEK2, PBK and BUB1B compared with plexiform neurofibromas on a separate and significantly larger set of tumors (Figure 6 B through M). The plexiform neurofibroma samples only exhibited minimal immunoreactivity for BUB1B, whereas many of the malignant peripheral nerve sheath tumors showed distinct positivity that varied from focal to diffuse. There was weak focal expression of PBK in plexiform neurofibromas whereas the malignant peripheral nerve sheath tumors showed stronger and often more diffuse staining. NEK2 staining was also minimal in the plexiform neurofibromas. Malignant peripheral nerve sheath tumors showed variable staining with negative cases, cases showing cytoplasmic reactivity, cases with mixed nuclear and cytoplasmic reactivity and a few cases showing only nuclear immunoreactivity. Different patterns of immunohistochemical staining with variable nuclear and cytoplasmic labeling have previously been observed in studies of NEK2 expression.27, 28
Discussion
This study provides a unique comparison of gene expression profiling on matched paired samples of malignant peripheral nerve sheath tumor and the underlying plexiform neurofibroma they arise from. It utilizes a new technique for the expression profiling. The technical features of the nCounter platform20, 21 made this study feasible, because this system works with small amounts of RNA extracted from formalin-fixed paraffin tissue. Extraction of tissue from paraffin blocks matched to H&E slides allowed accurate sampling of areas of interest based on histomorphologic features. The often tight clustering of the two duplicate samples ran on one tissue core and the separation of tumors into their two diagnostic categories argues for the robustness of the expression data. Immunohistochemical studies for selected markers confirmed the results of the gene expression analysis at the protein level. The study design allowed two complimentary approaches for the data analysis: (1) Identification of differentially expressed genes based on the combined dataset and (2) pairwise comparison of matched plexiform neurofibroma/malignant peripheral nerve sheath tumors from the same surgical specimen.
The expression signature of kinase genes provides good separation between malignant peripheral nerve sheath tumors and plexiform neurofibromas in the principal component analysis and the distance matrix. This finding supports that the malignant transformation of plexiform neurofibroma to malignant peripheral nerve sheath tumor is associated with distinct changes in the expression profile of kinase genes. Most of the differentially expressed kinase genes show downregulation rather than overexpression. This result mirrors the changes described by Subramanian et al10 in their study of cDNA microarray data from frozen tumor samples. Interestingly, our approach did not clearly separate the low-grade malignant peripheral nerve sheath tumors from the plexiform neurofibromas. There has not yet been a systematic study of the gene expression profile found in low-grade malignant peripheral nerve sheath tumors. Subramanian et al10 also mention that an outlier case of malignant peripheral nerve sheath tumor with expression profile indistinguishable from neurofibroma represented low-grade malignant peripheral nerve sheath tumor. Defining the earliest molecular alterations of malignant transformation found in low-grade malignant peripheral nerve sheath tumors is therefore still an elusive goal.
Receptor tyrosine kinases are central elements of cell signaling pathways and potential therapeutic targets. Overall, the 54-receptor tyrosine kinases with informative readings show a quite diverse pattern of altered expression in the plexiform neurofibroma/malignant peripheral nerve sheath tumor sets. In all cases, transformation towards malignant peripheral nerve sheath tumors goes along with upregulation of a few receptor tyrosine kinases and downregulation of most others. The pattern of this change is very diverse. EGFR has previously been described as expressed in malignant peripheral nerve sheath tumors and as a potential target for treatment.29, 30, 31 In this analysis, EGFR is expressed at significant levels in most samples. The pairwise comparison between preexisting plexiform neurofibroma and malignant peripheral nerve sheath tumor shows upregulation of EGFR expression in four malignant peripheral nerve sheath tumors and downregulation in eight malignant peripheral nerve sheath tumors. Expression changes in EGFR were therefore not a uniform feature of malignant transformation in this data set. Previous reports have shown expression of EGFR in subsets of malignant peripheral nerve sheath tumors and neurofibromas as well as amplification of the EGFR locus in some 26–28% of malignant peripheral nerve sheath tumors.10, 29, 30, 31, 32 Gene dosage reduction and downregulation of ERBB2 and ERBB3 as seen here has previously been reported in malignant peripheral nerve sheath tumors.29, 33
Neurofibromatosis type 1-associated neurofibromas and malignant peripheral nerve sheath tumors arise in a background of a ‘rasopathy’ with altered Ras/MAP kinase signaling.3, 14, 34, 35 We therefore looked for changes occurring with the transformation from plexiform neurofibroma to malignant peripheral nerve sheath tumor in the expression level of kinases in this signaling cascade. Most kinases of this set are again downregulated in malignant peripheral nerve sheath tumors. ERK kinases (MAPK1 and MAPK3) are expressed in the malignant peripheral nerve sheath tumors but at comparatively lower levels compared with the preexisting plexiform neurofibroma.
The results on expression level changes in the receptor tyrosine kinases and the Ras/MAP kinase signaling pathway both suggest that the expression changes in these kinase genes are complex and diverse. Many of these kinases are expressed in neurofibromas and malignant peripheral nerve sheath tumors. There may be only subtle expression level changes. The effect of such graded changes on tumor behavior is difficult to predict in these complex networks. The analysis does not identify any single candidate gene that would act as a simple on–off switch in the malignant transformation towards malignant peripheral nerve sheath tumor. None of these kinases shows uniform changes that could make it a diagnostic marker or a therapeutic target.
Overall, only relatively few kinases were found to show significant upregulation with the malignant transformation towards malignant peripheral nerve sheath tumor. The performed pathway analysis identified kinases implicated in mitotic regulation as a distinct set of differentially expressed genes including PBK, BUB1B, NEK2, CHEK1 and BUB1. Some of these mitotic regulators including NEK2, PBK and BUB1 were also found to be differentially expressed in the study by Subramanian et al.10 The overexpression of BUB1B, PBK and NEK2 in malignant peripheral nerve sheath tumors at the protein level was confirmed by immunohistochemistry. All three of these kinases have recently been shown to have a role in other malignancies and are interesting candidates for genes in the context of the malignant transformation of plexiform neurofibromas:
-
BUB1B and BUB1 are part of the spindle assembly checkpoint that ensures proper segregation of the chromosomes.36, 37, 38, 39 Alterations in BUB1B have been documented in a number of cancers including breast cancer, lung cancer, colon cancer and gastric cancer.37, 38, 40, 41, 42, 43, 44 In these, BUB1B overexpression appears associated with more aggressive behavior, increased proliferative activity, genomic complexity, chromosomal instability and DNA aneuploidy.37, 40, 41, 42, 43, 45, 46 Interestingly, aneuploidy and complex karyotype are common finding in malignant peripheral nerve sheath tumors.
-
NEK2 is one of several kinases including CDK1/cyclin B and Aurora A that are associated with the centrosome during cell cycle progression. NEK2 is involved in centrosome cycle as well as spindle assembly.27 NEK2 promotes centrosome separation at the onset of mitosis and is involved in cell division and mitotic regulation.47, 48 Overexpression of NEK2 induces premature centrosome separation and nuclear defects, which are indicative of mitotic errors.49 High levels of NEK2 expression are found in seminomas27 and ductal carcinoma of the breast.28
-
PBK is overexpressed in some hematopoetic neoplasms including acute lymphoblastic leukemia, multiple myeloma and Burkitt’s lymphoma but also in colon cancer, breast cancer and cholangiocarcinoma.50, 51, 52 It is thought to be involved in cytokinetic function and DNA damage repair.
In summary, this study shows that there are distinct changes in kinase expression that occur with malignant transformation from plexiform neurofibroma to malignant peripheral nerve sheath tumor. Downregulation of kinases rather than overexpression appears to be the dominant change of this malignant transformation. Changes in MAP kinase signaling and receptor tyrosine kinases like EGFR have been suggested as possible additional biological changes driving malignant transformation of plexiform neurofibromas towards malignant peripheral nerve sheath tumor beyond the shared neurofibromin mutation.14 Receptor tyrosine kinases have also been proposed as possible therapeutic targets.53 The presented data suggests that this malignant transformation is associated with rather complex diverse changes and that it does not rely on a single biological switch. In many cases, the expression of these kinase genes shows graded changes rather than a simple alteration between on–off states. Regulators of mitotic progression including PBK, BUB1B and NEK2 stand out as distinct set of overexpressed genes. These could represent potential targets for the treatment of malignant peripheral nerve sheath tumors.
References
Tonsgard JH . Clinical manifestations and management of neurofibromatosis type 1. Semin Pediatr Neurol 2006;13:2–7.
Tucker T, Wolkenstein P, Revuz J et al Association between benign and malignant peripheral nerve sheath tumors in NF1. Neurology 2005;65:205–211.
Upadhyaya M . Genetic basis of tumorigenesis in NF1 malignant peripheral nerve sheath tumors. Front Biosci 2011;16:937–951.
Mantripragada KK . Diaz de Stahl T, Patridge C, et al. Genome-wide high-resolution analysis of DNA copy number alterations in NF1-associated malignant peripheral nerve sheath tumors using 32 K BAC array. Genes Chromosomes Cancer 2009;48:897–907.
Upadhyaya M, Spurlock G, Thomas L et al Microarray-based copy number analysis of neurofibromatosis type-1 (NF1)-associated malignant peripheral nerve sheath tumors reveals a role for Rho-GTPase pathway genes in NF1 tumorigenesis. Hum Mutat 2012;33:763–776.
Bridge RS, Bridge JA, Neff JR et al Recurrent chromosomal imbalances and structurally abnormal breakpoints within complex karyotypes of malignant peripheral nerve sheath tumour and malignant triton tumour: a cytogenetic and molecular cytogenetic study. J Clin Pathol 2004;57:1172–1178.
Pytel P, Karrison T, Can G et al Neoplasms with schwannian differentiation express transcription factors known to regulate normal schwann cell development. Int J Surg Pathol 2010;18:449–457.
Spurlock G, Knight SJ, Thomas N et al Molecular evolution of a neurofibroma to malignant peripheral nerve sheath tumor (MPNST) in an NF1 patient: correlation between histopathological, clinical and molecular findings. J Cancer Res Clin Oncol 2010;136:1869–1880.
Miller SJ, Jessen WJ, Mehta T et al Integrative genomic analyses of neurofibromatosis tumours identify SOX9 as a biomarker and survival gene. EMBO Mol Med 2009;1:236–248.
Subramanian S, Thayanithy V, West RB et al Genome-wide transcriptome analyses reveal p53 inactivation mediated loss of miR-34a expression in malignant peripheral nerve sheath tumours. J Pathol 2010;220:58–70.
Yang J, Ylipaa A, Sun Y et al Genomic and molecular characterization of malignant peripheral nerve sheath tumor identifies the IGF1R pathway as a primary target for treatment. Clin Cancer Res 2011;17:7563–7573.
Miller SJ, Rangwala F, Williams J et al Large-scale molecular comparison of human schwann cells to malignant peripheral nerve sheath tumor cell lines and tissues. Cancer Res 2006;66:2584–2591.
Itani S, Kunisada T, Morimoto Y et al MicroRNA-21 correlates with tumorigenesis in malignant peripheral nerve sheath tumor (MPNST) via programmed cell death protein 4 (PDCD4). J Cancer Res Clin Oncol 2012;138:1501–1509.
Carroll SL . Molecular mechanisms promoting the pathogenesis of Schwann cell neoplasms. Acta Neuropathol 2012;123:321–348.
King AA, Debaun MR, Riccardi VM et al Malignant peripheral nerve sheath tumors in neurofibromatosis 1. Am J Med Genet 2000;93:388–392.
Evans DG, Baser ME, McGaughran J et al Malignant peripheral nerve sheath tumours in neurofibromatosis 1. J Med Genet 2002;39:311–314.
Stucky CC, Johnson KN, Gray RJ et al Malignant peripheral nerve sheath tumors (MPNST): the Mayo Clinic experience. Ann Surg Oncol 2012;19:878–885.
Longhi A, Errani C, Magagnoli G et al High grade malignant peripheral nerve sheath tumors: outcome of 62 patients with localized disease and review of the literature. J Chemother 2010;22:413–418.
Lee PR, Cohen JE, Tendi EA et al Transcriptional profiling in an MPNST-derived cell line and normal human Schwann cells. Neuron Glia Biol 2004;1:135–147.
Geiss GK, Bumgarner RE, Birditt B et al Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol 2008;26:317–325.
Reis PP, Waldron L, Goswami RS et al mRNA transcript quantification in archival samples using multiplexed, color-coded probes. BMC Biotechnol 2011;11:46.
Rodriguez FJ, Folpe AL, Giannini C et al Pathology of peripheral nerve sheath tumors: diagnostic overview and update on selected diagnostic problems. Acta Neuropathol 2012;123:295–319.
Storey JD, Tibshirani R . Statistical significance for genomewide studies. Proc Natl Acad Sci USA 2003;100:9440–9445.
Dennis G, Sherman BT, Hosack DA et al DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol 2003;4:P3.
Huang da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009;4:44–57.
Huang da W, Sherman BT, Zheng X et al Extracting biological meaning from large gene lists with DAVID. Curr Protoc Bioinformatics 2009;27 (Suppl):13.11.1–13.11.13.
Barbagallo F, Paronetto MP, Franco R et al Increased expression and nuclear localization of the centrosomal kinase Nek2 in human testicular seminomas. J Pathol 2009;217:431–441.
Wang S, Li W, Lv S et al Abnormal expression of Nek2 and beta-catenin in breast carcinoma: clinicopathological correlations. Histopathology 2011;59:631–642.
Holtkamp N, Malzer E, Zietsch J et al EGFR and erbB2 in malignant peripheral nerve sheath tumors and implications for targeted therapy. Neuro Oncol 2008;10:946–957.
Perrone F, Da Riva L, Orsenigo M et al PDGFRA, PDGFRB, EGFR, and downstream signaling activation in malignant peripheral nerve sheath tumor. Neuro Oncol 2009;11:725–736.
Keizman D, Issakov J, Meller I et al Expression and significance of EGFR in malignant peripheral nerve sheath tumor. J Neurooncol 2009;94:383–388.
Perry A, Kunz SN, Fuller CE et al Differential NF1, p16, and EGFR patterns by interphase cytogenetics (FISH) in malignant peripheral nerve sheath tumor (MPNST) and morphologically similar spindle cell neoplasms. J Neuropathol Exp Neurol 2002;61:702–709.
Levy P, Vidaud D, Leroy K et al Molecular profiling of malignant peripheral nerve sheath tumors associated with neurofibromatosis type 1, based on large-scale real-time RT-PCR. Mol Cancer 2004;3:20.
Rauen KA, Banerjee A, Bishop WR et al Costello and cardio-facio-cutaneous syndromes: Moving toward clinical trials in RASopathies. Am J Med Genet C Semin Med Genet 2011;157:136–146.
Kraniak JM, Sun D, Mattingly RR et al The role of neurofibromin in N-Ras mediated AP-1 regulation in malignant peripheral nerve sheath tumors. Mol Cell Biochem 2010;344:267–276.
Logarinho E, Bousbaa H . Kinetochore-microtubule interactions ‘in check’ by Bub1, Bub3 and BubR1: the dual task of attaching and signalling. Cell Cycle 2008;7:1763–1768.
Rizzardi C, Torelli L, Barresi E et al BUBR1 expression in oral squamous cell carcinoma and its relationship to tumor stage and survival. Head Neck 2011;33:727–733.
Ando K, Kakeji Y, Kitao H et al High expression of BUBR1 is one of the factors for inducing DNA aneuploidy and progression in gastric cancer. Cancer Sci 2010;101:639–645.
Lee YK, Choi E, Kim MA et al BubR1 as a prognostic marker for recurrence-free survival rates in epithelial ovarian cancers. Br J Cancer 2009;101:504–510.
Du J, Du Q, Zhang Y et al Expression of cell-cycle regulatory proteins BUBR1, MAD2, Aurora A, cyclin A and cyclin E in invasive ductal breast carcinomas. Histol Histopathol 2011;26:761–768.
Scintu M, Vitale R, Prencipe M et al Genomic instability and increased expression of BUB1B and MAD2L1 genes in ductal breast carcinoma. Cancer Lett 2007;254:298–307.
Yuan B, Xu Y, Woo JH et al Increased expression of mitotic checkpoint genes in breast cancer cells with chromosomal instability. Clin Cancer Res 2006;12:405–410.
Grabsch H, Takeno S, Parsons WJ et al Overexpression of the mitotic checkpoint genes BUB1, BUBR1, and BUB3 in gastric cancer—association with tumour cell proliferation. J Pathol 2003;200:16–22.
Shichiri M, Yoshinaga K, Hisatomi H et al Genetic and epigenetic inactivation of mitotic checkpoint genes hBUB1 and hBUBR1 and their relationship to survival. Cancer Res 2002;62:13–17.
Pinto M, Vieira J, Ribeiro FR et al Overexpression of the mitotic checkpoint genes BUB1 and BUBR1 is associated with genomic complexity in clear cell kidney carcinomas. Cell Oncol 2008;30:389–395.
Yamamoto Y, Matsuyama H, Chochi Y et al Overexpression of BUBR1 is associated with chromosomal instability in bladder cancer. Cancer Genet Cytogenet 2007;174:42–47.
Fry AM . The Nek2 protein kinase: a novel regulator of centrosome structure. Oncogene 2002;21:6184–6194.
Fletcher L, Cerniglia GJ, Nigg EA et al Inhibition of centrosome separation after DNA damage: a role for Nek2. Radiat Res 2004;162:128–135.
Hayward DG, Newbatt Y, Pickard L et al Identification by high-throughput screening of viridin analogs as biochemical and cell-based inhibitors of the cell cycle-regulated nek2 kinase. J Biomol Screen 2010;15:918–927.
He F, Yan Q, Fan L et al PBK/TOPK in the differential diagnosis of cholangiocarcinoma from hepatocellular carcinoma and its involvement in prognosis of human cholangiocarcinoma. Hum Pathol 2010;41:415–424.
Zhu F, Zykova TA, Kang BS et al Bidirectional signals transduced by TOPK-ERK interaction increase tumorigenesis of HCT116 colorectal cancer cells. Gastroenterology 2007;133:219–231.
Park JH, Lin ML, Nishidate T et al PDZ-binding kinase/T-LAK cell-originated protein kinase, a putative cancer/testis antigen with an oncogenic activity in breast cancer. Cancer Res 2006;66:9186–9195.
Torres KE, Liu J, Young E et al Expression of 'drugable’ tyrosine kinase receptors in malignant peripheral nerve sheath tumour: potential molecular therapeutic targets for a chemoresistant cancer. Histopathology 2011;59:156–159.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Stricker, T., Henriksen, K., Tonsgard, J. et al. Expression profiling of 519 kinase genes in matched malignant peripheral nerve sheath tumor/plexiform neurofibroma samples is discriminatory and identifies mitotic regulators BUB1B, PBK and NEK2 as overexpressed with transformation. Mod Pathol 26, 930–943 (2013). https://doi.org/10.1038/modpathol.2012.242
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2012.242
Keywords
This article is cited by
-
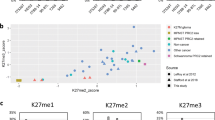
Loss of H3K27 tri-methylation is a diagnostic marker for malignant peripheral nerve sheath tumors and an indicator for an inferior survival
Modern Pathology (2016)
-
Treatment of Neurofibromatosis Type 1
Current Treatment Options in Neurology (2015)