Abstract
Foxp3 (forkhead box P3 transcription factor)-expressing regulatory T cells (Tregs) are essential for immunological tolerance, best illustrated by uncontrolled effector T-cell responses and autoimmunity upon loss of Foxp3 expression. Tregs can adopt specific effector phenotypes upon activation, reflecting the diversity of functional demands in the different tissues of the body. Here, we report that Foxp3+CD4+ T cells coexpressing retinoic acid-related orphan receptor-γt (RORγt), the master transcription factor for T helper type 17 (Th17) cells, represent a stable effector Treg lineage. Transcriptomic and epigenetic profiling revealed that Foxp3+RORγt+ T cells display signatures of both Tregs and Th17 cells, although the degree of similarity was higher to Foxp3+RORγt− Tregs than to Foxp3−RORγt+ T cells. Importantly, Foxp3+RORγt+ T cells were significantly demethylated at Treg-specific epigenetic signature genes such as Foxp3, Ctla-4, Gitr, Eos, and Helios, suggesting that these cells have a stable regulatory rather than inflammatory function. Indeed, adoptive transfer of Foxp3+RORγt+ T cells in the T-cell transfer colitis model confirmed their Treg function and lineage stability in vivo, and revealed an enhanced suppressive capacity as compared with Foxp3+RORγt− Tregs. Thus, our data suggest that RORγt expression in Tregs contributes to an optimal suppressive capacity during gut-specific immune responses, rendering Foxp3+RORγt+ T cells as an important effector Treg subset in the intestinal system.
Similar content being viewed by others
INTRODUCTION
CD4+ T cells expressing the forkhead box P3 transcription factor (Foxp3), termed regulatory T cells (Tregs), are crucial for a viable immunological homeostasis by maintaining tolerance against self and foreign antigens. Although a majority of Foxp3+ Tregs originate in the thymus (thymus-derived Tregs), a considerable fraction of Foxp3+ Tregs are de novo induced in the periphery from naive CD4+ T cells.1 Recent findings indicate that under homeostatic conditions, such peripherally induced Tregs (pTregs) are of functional importance mainly at feto–maternal interfaces and for maintaining tolerance to food- and microbiota-derived antigens at mucosal sites.2, 3, 4, 5, 6 In addition to differences in ontogeny, there is growing evidence that Tregs having undergone antigen-specific stimulation may differ significantly from their naive recirculating counterparts in terms of activation status, migratory potential, and regulatory function.7, 8 Together with the notion that Tregs having different origins or anatomical locations display distinct but specific gene expression profiles,9 this has led to the discovery of different functional Treg subpopulations, termed “effector” Treg lineages.10, 11 Interestingly, recent reports indicate that Foxp3+ Tregs can coopt the expression of specific transcription factors that are associated with the differentiation and function of effector CD4+ T-cell lineages. In the context of T helper type 1 (Th1)-mediated inflammation, Tregs can upregulate the Th1-specific transcription factor T-bet, leading to the expression of CXCR3 and the accumulation of Foxp3+ T-bet+ Tregs at sites of inflammation. Importantly, T-bet expression was shown to be essential for the homeostasis and function of Tregs during type-1 inflammation.12 Similarly, the Th2-associated transcription factor GATA3 was found to play an important role for Treg function, as it was shown that GATA3-deficient Tregs display profound defects in peripheral homeostasis and suppressive function.13, 14 In addition, Treg-specific deletion of the transcription factors IRF4 (interferon regulatory factor 4) or STAT3 (signal transducer and activator of transcription 3) resulted in an impaired regulation of Th2- and Th17-dominated immune responses, respectively.15, 16 Recently, a population of germinal center-resident Tregs, termed follicular regulatory T cells, was identified.17, 18 These follicular regulatory T cells suppress germinal center B and T cells and depend, similar to their follicular T helper cell counterparts, on the expression of the transcription factor Bcl6 for their development. Together, these findings support the possibility that the expression of lineage-specific transcription factors may drive the generation of effector Tregs that are specifically suited to regulate immune responses mediated by their corresponding conventional effector CD4+ T-cell lineages.
Recently, we and others have identified a CD4+ T-cell population that simultaneously expresses the transcription factor retinoic acid-related orphan receptor-γt (RORγt), initially described as being essential for Th17 development,19 and Foxp3.20, 21 Naive CD4+ T cells start to express RORγt and Foxp3 when stimulated in vitro under Th17-inducing conditions,20, 21 and it was further suggested that from this intermediate Foxp3+RORγt+ stage, the cells differentiate into either the Treg or Th17 lineage, dependent on the balance of transforming growth factor-β vs. inflammatory cytokines such as interleukin (IL)-6, IL-21, and IL-23.21 Nevertheless, we also demonstrated that in vivo, Foxp3+RORγt+ T cells exist that display the ability to express IL-10 and to suppress the proliferation of effector T cells.20 In contrast to these findings, it has been shown that Foxp3+ Tregs can start to express IL-17 in mice and humans,22, 23 and it was speculated that such cells may contribute to inflammation-associated pathology. Thus, it was not clear so far how Foxp3+RORγt+ T cells relate to bona fide Foxp3+ Tregs and RORγt+ Th17 cells and whether they represent a stable T-cell phenotype with functional relevance in vivo.
In this study, we demonstrate that Foxp3+RORγt+ T cells represent a stable Treg lineage with a highly suppressive phenotype in vivo. Although these cells expressed genes associated with both Tregs and Th17 cells, the majority of mRNAs cluster with Tregs and the epigenetic profile of Treg/Th17-associated genomic regions was indicative of a stable Treg phenotype. The stability of this lineage was further confirmed both in vitro and in vivo upon adoptive transfer in mice. Moreover, Foxp3+RORγt+ T cells showed an enhanced suppressive capacity in the T-cell transfer colitis model as compared with Foxp3+RORγt− Tregs, indicating that Foxp3+RORγt+ T cells represent a potent effector Treg lineage, being highly specialized to regulate immune responses at intestinal sites.
RESULTS
High proportion of Foxp3+RORγt+ T cells in the colon of mice depends on a complex microbiota
We reported previously that 20–30% of Foxp3+ T cells in the lamina propria (LP) of the gut express RORγt.20 In order to better characterize this Foxp3+RORγt+ T-cell population, we first sought to analyze in detail the distribution of these cells in different peripheral and gut-associated organs. In mice containing a complex microflora,24 the proportion of RORγt-expressing cells within the Foxp3+ T-cell pool was ∼10% in lymphoid organs, such as the spleen or peripheral lymph nodes (pLNs). However, in contrast to mice that harbor only a minimal flora,25 the frequency of Foxp3+RORγt+ T cells was clearly elevated in the gut-draining mesenteric lymph nodes (mLNs), Peyer’s patches, and especially the intestinal colonic LP, where ∼80% of all Foxp3+ T cells expressed RORγt (Figure 1a,b). Thus, our data confirm that in the presence of a complex microbiota, a high proportion of Foxp3+ T cells in the intestinal organs, and especially in the LP of the colon, simultaneously express RORγt.
Increased frequency of Foxp3+RORγt+ T cells in mice with complex flora. Spleen, peripheral lymph nodes (pLNs), mesenteric lymph nodes (mLNs), Peyer’s patches (PP), and colonic lamina propria (cLP) were harvested from C57BL/6 mice (minimal flora) or Foxp3RFPRORγtGFP mice (complex flora). Lymphocytes were isolated and the expression levels of RORγt and Foxp3 were analyzed by intracellular staining. (a) Representative fluorescence-activated cell sorting (FACS) plots gated on live CD3+ CD4+ T cells. Numbers indicate the quadrant frequency. (b) Frequency of Foxp3+RORγt+ T cells in different organs isolated from minimal or complex flora-containing Foxp3RFPRORγtGFP mice. Data are shown as mean±s.d. (a, b). Foxp3, forkhead box P3 transcription factor; RORγt, retinoic acid-related orphan receptor-γt. **P<0.01.
Foxp3+RORγt+ T cells show a higher overlap in gene regulation with Tregs and display a T-cell receptor (TCR) repertoire indicative of peripherally expanded T cells
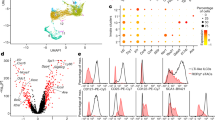
In order to allow for an unequivocal identification and isolation of the different Foxp3- and RORγt-expressing T-cell populations, we crossed BAC-transgenic Rorc(γt)-GfpTG mice, expressing green fluorescent protein (GFP) under control of the Rorc locus,20 with Foxp3-IRES-mRFP (FIR) mice, expressing a red fluorescent protein (RFP) under control of the endogenous Foxp3 locus.26 The strong reporter signals in these Foxp3RFPRORγtGFP double reporter mice enabled us to isolate Foxp3 and RORγt-expressing T-cell populations in high purity for subsequent analyses (Supplementary Figure S1 online). To study the global relationship in their gene expression profiles, we sorted Foxp3−RORγt− CD4+ T cells, Foxp3+RORγt− T cells (further designated as Foxp3+ Tregs), RORγt+Foxp3− T cells (further designated as RORγt+ T cells), and Foxp3+RORγt+ T cells from pooled cells of spleen, pLNs, and mLNs in order to avoid bias due to a specific tissue location. Interestingly, Foxp3+RORγt+ T cells shared expression of ∼99% of their genes with both RORγt+ T cells and Foxp3+ Tregs under homeostatic conditions (Figure 2a). However, hierarchical clustering with specifically expressed genes from RORγt+ T cells and Foxp3+ Tregs revealed a closer relationship between Foxp3+RORγt+ T cells and Foxp3+ Tregs (Figure 2b). In order to better define the phenotypic characteristics of Foxp3+RORγt+ T cells, we performed a more specific candidate comparison between the T-cell subsets. Foxp3+RORγt+ T cells expressed high levels of Th17-assocoiated genes like Il23r, Il1r1, Maf, Irf4, and Ikzf3 (Aiolos) when compared with Foxp3+ Tregs (Figure 2c). However, the expression of Th17-associated effector cytokines like Il17a, Il17f, or Il21 was significantly lower when compared with RORγt+ T cells. Intriguingly, expression of Stat3 was not upregulated in any T-cell subset. When the expression of Treg-associated genes was inspected, we observed a comparably high expression of Ikzf4 (Eos), Ikzf2 (Helios), Nrp1, Il2ra (CD25), and Tnfrsf18 (GITR) in Foxp3+RORγt+ T cells and Foxp3+ Tregs (Figure 2d). In addition, Foxp3+RORγt+ T cells displayed a uniquely high expression of genes that were recently associated with Treg populations resident in mucosal tissues,9, 27 including Il10, Gzmb, Prdm1 (Blimp-1), Itgae (CD103), Ctla4, and Icos (Figure 2e). This was in accordance with an activated CD44int/high CD62Llow phenotype, high CD25 surface expression, and increased ICOS levels as determined by fluorescence-activated cell sorting (FACS) analysis in mLNs and colon (Supplementary Figure S2). Genes involved in the transforming growth factor-β pathway, like Tgfbr1 and Smad3, did not show any differential expression among the subsets analyzed. Similar to RORγt+ T cells, Foxp3+RORγt+ T cells expressed high levels of the chemokine receptors Ccr4, Ccr6, and Ccr9, whereas Ccr7 was expressed at high levels in all subsets (Figure 2f and Supplementary Figure S3). Interestingly, Foxp3+RORγt+ T cells also showed increased expression of Gpr15 and Ffar2, the two receptors with fundamental roles especially in colonic Treg homeostasis.28, 29 To directly assess the tissue-homing capacity of Foxp3+RORγt+ T cells, we performed short-term homing assays using adoptive transfer of radiolabeled T cells into wild-type recipient mice.30 Although Foxp3+RORγt+ T cells showed a slight tendency toward enhanced homing into mucosal tissues, overall the migration behavior was comparable to Foxp3+ Tregs (Supplementary Figure S4). In summary, our data indicate that Foxp3+RORγt+ T cells show a closer relationship to Foxp3+ Tregs in terms of global gene expression. However, comparison of specific profiles revealed a T-cell subset carrying features of both Th17 and particularly intestinal tissue-resident Tregs. Of note, however, Foxp3+RORγt+ T cells did not show unique upregulation of genes specific for effector T cells of the Th1 (Tbx21, Cxcr3 and Ifng), Th2 (Gata3 and Il4), or follicular T helper (Bcl6) lineage (Figure 2g).
Foxp3+RORγt+ T cells show a closer relationship to Foxp3+ regulatory T cells (Tregs). Foxp3+/− RORγt+/− T cells were sorted from pooled spleen and lymph node cells of Foxp3RFPRORγtGFP reporter mice for RNA extraction. Gene expression levels and T-cell receptor (TCR) repertoire of the T-cell populations were examined by microarray analysis and next-generation sequencing, respectively. (a) Comparison of expression profiles between Foxp3+RORγt+ T cells and Foxp3+ Tregs or RORγt+ T cells. Up- and downregulated mRNAs with a fold change (FC) of 1.5 (P<0.05) are numbered and indicated in red or blue. (b) Hierarchical clustering of normalized, mean–centered, and scaled expression values of 858 genes being differentially expressed (log FC>1.5, adjusted P≤0.05) between conditions RORγt+Foxp3− and RORγt−Foxp3+ T cells. (c–g) Heatmaps showing transcription factors and lineage-specific molecules for (c) T helper type 17 (Th17) cells, (d, e) Tregs, (f) Th17- and/or Treg-associated chemokine receptor and gut-specific receptors, and (g) Th1, Th2, and follicular T helper (TFH) cells. The averaged log2 mRNA expression level of two individual experiments is shown with a color code from blue (4) via white up to red (16). (h) Analysis of the TRAV12 TCR diversity of Foxp3+/− RORγt+/− T cells. Diversity (exp(Shannon index)) was calculated from 6,700 productive rearranged TRAV12 CDR3 amino acid sequences. Foxp3, forkhead box P3 transcription factor; RORγt, retinoic acid-related orphan receptor-γt.
Next, we analyzed the TCR repertoire diversity of Foxp3+RORγt+ T cells by next-generation sequencing of the TRAV12 gene segments.31 Consistent with the idea that Tregs in the periphery are continuously expanding by encounter of self-antigen,32, 33 not only Foxp3+ Tregs but also Foxp3+RORγt+ T cells and RORγt+ T cells showed lower TCR diversities compared with Foxp3−RORγt− T cells (Figure 2h). FACS analysis for the cell division marker Ki-67 confirmed that Foxp3+ Tregs, RORγt+ Th17, and especially Foxp3+RORγt+ T cells displayed enhanced frequencies of dividing cells compared with Foxp3−RORγt− T cells (Supplementary Figure S5). However, we found no qualitative differences in the TCR-α repertoire of these four CD4+ T-cell subsets, as numbers of shared clonotypes remained similar when comparing distinct subsets (Supplementary Figure S6a). As a result, 3 to 8 of the 25 most prevalent TCR-α sequences obtained from Foxp3+RORγt+ T cells were consistently shared with the other CD4+ T-cell subsets (Supplementary Figure S6b). Therefore, Foxp3+RORγt+ T cells show a restricted TCR repertoire of cells that have proliferated in response to peripheral stimuli but their TCR-α sequences overlap to a similar degree with Foxp3+ Treg, RORγt+ T cells, and Foxp3−RORγt− CD4+ T cells.
Epigenetic profiling reveals a Treg-specific CpG hypomethylation pattern for Foxp3+RORγt+ T cells
Stable expression of specific T-cell lineage transcription factors and cytokines requires epigenetic modifications within the respective gene loci.34, 35 We had previously demonstrated that demethylation of CpG motifs at the Treg-specific demethylated region (TSDR) within the Foxp3 locus is indicative of a stable Foxp3 expression in Tregs.36, 37, 38 To determine the methylation status of the TSDR in Foxp3+RORγt+ T cells, we purified cells from spleen, LNs (pool of pLNs and mLNs), and colonic LP of Foxp3RFPRORγtGFP mice and compared the TSDR methylation status to that of RORγt+ T cells, Foxp3+ Tregs, and Foxp3−RORγt− T cells isolated from the same organs. As shown in Figure 3a, Foxp3+RORγt+ T cells showed a similar high degree of TSDR demethylation as compared with Foxp3+ Tregs in all organs analyzed, whereas both RORγt+ T cells and Foxp3−RORγt− T cells displayed a highly methylated TSDR. Although these data indicate that Foxp3 is stably expressed in Foxp3+RORγt+ T cells, recent findings suggest that stable expression of Foxp3 alone is not sufficient for conferring and maintaining Treg function and phenotype.39 In addition, a number of Treg-specific epigenetic signature genes (e.g., Ctla4, Tnfrsf18 (GITR) and Ikzf4 (Eos)) need to be fully demethylated to enable Foxp3+ T cells to acquire Treg-specific gene expression, lineage stability, and full suppressive activity.40 Interestingly, Foxp3+RORγt+ T cells also showed substantial demethylation at Ctla4, Tnfrsf18, and Ikzf4, although the degree of demethylation at Tnfrsf18 and Ikzf4 was somewhat lower as compared with Foxp3+ Tregs (Figure 3b). As expected, these loci remained largely methylated in RORγt+ T cells, except for Ctla4 that was substantially demethylated in these cells. We also investigated the methylation status of Ikzf2 (Helios), known as a marker solely expressed by thymus-derived Tregs,41 and observed an almost complete demethylation in Foxp3+ Tregs, whereas Foxp3+RORγt+ T cells displayed a significant degree of methylation (74%) at this site (Figure 3c). To further elucidate this point, we directly determined expression of Helios in cells isolated from different lymphoid organs and the colonic LP. In line with the methylation data, Foxp3+RORγt+ T cells always showed a reduced Helios expression when compared with Foxp3+ Tregs, with the exception of peripheral LN, where a similar expression was observed (Figure 3d). Together, these data reveal substantial organ-specific differences in Helios expression in Foxp3+RORγt+ T cells and Foxp3+ Tregs that might be under epigenetic control.
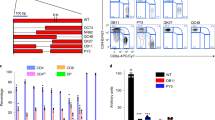
Methylations status of the regulatory T cell (Treg)-specific demethylated region (TSDR) and other Treg-associated epigenetic loci indicates a stable Treg phenotype of Foxp3+RORγt+ T cells. Foxp3+/− RORγt+/− T-cell subsets were fluorescence-activated cell sorted (FACS) from spleen, lymph nodes (LNs), and colon of Foxp3RFPRORγtGFP reporter mice. Analyzed cell subsets from different organs are depicted for (a) TSDR, (b) Treg-specific CpG hypomethylated pattern including Ctla4, Tnfrsf18, and Ikzf4, and (c) Ikzf2. The experiment was performed with two independent sorts. The methylation rates were translated into a color code from yellow (0%) via green (50%) up to blue (100%). Each rectangle represents the methylation of one CpG motif. cLP, colonic lamina propria; Foxp3, forkhead box P3 transcription factor; MFI, mean fluorescence intensity; mLN, mesenteric lymph node; pLN, peripheral lymph node; RORγt, retinoic acid-related orphan receptor-γt. (d) Helios expression of FACS-sorted Foxp3+/−RORγt+/− T-cell subsets was analyzed for different organs by intracellular FACS staining. Representative histograms and a summary of the surface expression of Helios of two independent experiments are depicted. Data are representative for two independent experiments with five mice per experiment and shown as mean±s.d. (e) Correlation of DNA demethylation (left y axis) and gene expression assessed by microarray analysis (right y axis) of indicated Treg and T helper type 17 (Th17) epigenetic signature genes. The graph shows a pool of five to six methylation analyses and three gene expression profiles pooled from spleen and LN for each subset. Data are shown as mean±s.d. F+, Foxp3+RORγt− cells; R+F+, Foxp3+RORγt+ cells; R+, Foxp3−RORγt+ cells. *P<0.05, NS, not significant.
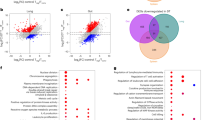
As our transcriptomic analysis revealed expression of several Th17-associated genes in Foxp3+RORγt+ T cells (Figure 2), we also analyzed the methylation status of Th17-specific epigenetic signature genes, including Il17a, Zfp362, Ccr6, Acsbg1, Rora, Dpp4, and Dclk1.42 Foxp3+RORγt+ T cells showed a more pronounced demethylation at Il17a, Zfp362, Dpp4, and Dclk1 when compared with Foxp3+ Tregs (Figure 4a), indicating that Foxp3+RORγt+ T cells also bare specific similarities to Th17 cells in their epigenetic make-up. Importantly, the epigenetic profile of not only Th17-specific but also Treg-specific epigenetic signature genes analyzed here was directly correlated with the respective gene expression levels determined in the transcriptome analyses (Figures 3e and 4b). Together, our data demonstrate that Foxp3+RORγt+ T cells display similarities in their epigenetic profile to both RORγt+ T cells and Foxp3+ Tregs. Yet, the substantial demethylation of Treg-specific epigenetic signature genes in Foxp3+RORγt+ T cells suggests a stable suppressive Treg phenotype for this cell type.
Foxp3+RORγt+ T cells display demethylation of specific T helper type 17 (Th17)-associated epigenetic regions. Foxp3+/− RORγt+/− T-cell subsets were fluorescence-activated cell sorted (FACS) from spleen, lymph nodes (LNs), and colon of Foxp3RFPRORγtGFP reporter mice. (a) Analyzed cell subsets from different organs are depicted for Th17-associated epigenetic regions in genes including IL17a, Dpp4, Zfp362, Ccr6, Acsbg1, Rora, and Dclk1. The experiment was performed with two independent sorts. The methylation rates were translated into a color code from yellow (0%) via green (50%) up to blue (100%). Each rectangle represents the methylation of one CpG motif. cLP, colonic lamina propria; Foxp3, forkhead box P3 transcription factor; RORγt, retinoic acid-related orphan receptor-γt. (b) Correlation of DNA demethylation (left y axis) and gene expression assessed by microarray analysis (right y axis) of indicated Th17 epigenetic signature genes. The graph shows a pool of five to six methylation analyses and three gene-expression profiles pooled from spleen and LN for each subset. Data are shown as mean±s.d. F+, Foxp3+RORγt− cells; R+F+, Foxp3+RORγt+ cells; R+, Foxp3−RORγt+ cells.
Foxp3+RORγt+ T cells represent a stable Treg lineage in vitro and in vivo
We next determined whether the high lineage stability of Foxp3+RORγt+ T cells that was suggested by the epigenetic profiling could be confirmed under in vitro as well as in vivo settings. To this end, we sorted Foxp3+RORγt+ T cells, Foxp3+ Treg, RORγt+ T cells, and Foxp3−RORγt− CD4+ T cells and cultured them under TCR-stimulatory conditions (α-CD3, α-CD28) in the presence of IL-2 and IL-7 for 4 days. As expected, Foxp3−RORγt− T cells did not show a significant upregulation of Foxp3 under these conditions, but they displayed a substantial, although variable, induction of RORγt expression to a similar extent as Foxp3+ Tregs (Figure 5a,b). Foxp3 and RORγt expression were highly stable in Foxp3+ Tregs and RORγt+ T cells, respectively. Interestingly, Foxp3+RORγt+ T cells were also highly stable under these in vitro conditions, and only a small percentage of cells downregulated Foxp3 or RORγt expression (Figure 5a,b). Of note, no Foxp3 induction was observed in RORγt+ T cells, indicating that Foxp3+RORγt+ T cells do not develop from RORγt+ T cells.
Stable expression of Foxp3 and RORγt in Foxp3+RORγt+ T cells under in vitro culture conditions. Foxp3+/−RORγt+/− T cells were sorted from a pool of spleen and lymph node cells from Foxp3RFPRORγtGFP mice and cultured for 4 days in the presence of anti-CD3, anti-CD28 antibodies, and interleukin (IL)-2 and IL-7. Expression levels of Foxp3 and RORγt were assessed by fluorescence-activated cell sorting (FACS). (a) FACS plots show Foxp3 and RORγt expression on day 0 and day 4 of culture in individual live CD3+CD4+ T-cell populations. Numbers indicate the quadrant frequency. (b) Frequency of RORγt+ T cells and Foxp3+ T cells at day 4 of culture. Pooled data from three independent experiments are shown as mean±s.d. Foxp3, forkhead box P3 transcription factor; RORγt, retinoic acid-related orphan receptor-γt. *P<0.05, **P<0.01, ****P<0.0001, NS, not significant.
To analyze the in vivo stability of RORγt+Foxp3+ T cells, we applied an adoptive transfer approach by coinjecting either congenically marked (CD90.2) RORγt+Foxp3+, RORγt+ T cells or Foxp3+ Tregs together with naive CD4+ T cells (CD90.1) into lymphopenic RAG2-deficient mice. At 4 to 7 weeks after transfer, cells were reisolated from different organs and RORγt and Foxp3 expression levels were determined within the CD90.2+ T-cell population. As shown in Figure 6a,b, Foxp3 expression was very stable in transferred Foxp3+ Tregs, with over 90% of CD90.2+ T cells preserving their Foxp3 expression. Although loss of Foxp3 expression was slightly higher in transferred Foxp3+RORγt+ T cells, an average of 70–80% of cells still maintained Foxp3 expression in all organs analyzed (Figure 6a,b). Similarly, RORγt expression was maintained in the majority of transferred RORγt+Foxp3+ and RORγt+ T cells, and no difference in the stability of RORγt expression could be observed between the two subsets (Figure 6a,c). Rather, differences in RORγt expression were correlated to location, with high stability in the colon (80–90%) as compared with organs like the spleen or peripheral LNs (60–70%) (Figure 6a,c). Therefore, our data suggest that Foxp3+RORγt+ T cells do not represent an unstable, intermediate phenotype but rather an effector Treg lineage with high stability, especially in the gut. Of note, we found only a small fraction of unstable Foxp3+RORγt+ T cells that only lost Foxp3, but retained RORγt expression (Supplementary Figure S7), indicating that Foxp3+RORγt+ T cells do not preferentially develop into inflammatory Th17 cells. In accordance with the in vitro data, a significant proportion of adoptively transferred Foxp3+ Tregs upregulated RORγt expression, particularly those reisolated from the colon (Figure 6a,d), although we cannot rule out that the lymphopenic conditions in this transfer model contributed to the strong induction of RORγt in these cells. Nevertheless, we observed only minor upregulation of Foxp3 expression in adoptively transferred RORγt+ T cells (Figure 6a,e), suggesting that most of the Foxp3+RORγt+ T cells originate from Foxp3+ Tregs and not from RORγt+ Th17 cells.
Foxp3+RORγt+ T cells display high stability in vivo. Fluorescence-activated cell sorted (FACS) CD90.2+CD4+ Foxp3+RORγt−, Foxp3+RORγt+, and Foxp3−RORγt+ T cells from Foxp3RFPRORγtGFP mice were cotransferred with CD90.1+CD4+CD45RBhigh T cells from CD90.1 mice into RAG2-deficient mice in a ratio of 1:4. The mice were killed at weeks 4–7 after transfers. Cells from spleen, peripheral lymph nodes (pLNs), mesenteric lymph nodes (mLNs), and colon were analyzed for Foxp3 and RORγt expression in CD4+CD90.2+ cells by FACS. (a) Representative FACS plots of Foxp3 and RORγt expression in CD4+CD90.2+ cells in spleen and colon. Numbers indicate the quadrant frequency. (b) Stability of Foxp3 expression in Foxp3+RORγt− (▴) and Foxp3+RORγt+ (●) cells. (c) Stability of RORγt expression in Foxp3+RORγt+ (●) and Foxp3−RORγt+ (▪) cells. (d) Induction of RORγt in Foxp3+RORγt− T cells. (e) Induction of Foxp3 expression in Foxp3−RORγt+ T cells. Frequencies in live CD3+CD4+CD90.2+ T cells are displayed for different organs. FACS plots from one representative out of four independently performed experiments are shown. The graphs show pooled data from four independent experiments with 3–5 mice per group. cLP, colonic lamina propria; Foxp3, forkhead box P3 transcription factor; RORγt, retinoic acid-related orphan receptor-γt. *P<0.05, ***P<0.001, ****P<0.0001, NS, not significant.
Foxp3+RORγt+ T cells display superior suppressive capacity in T cell-mediated intestinal inflammation
Transfer of naive CD4+ T cells into lymphopenic mice induces a T cell-mediated autoimmune inflammation of the gut that can be prevented by cotransfer of Foxp3+ Tregs.43 Thus, the adoptive transfer approach described in Figure 6 enabled us to investigate the relative potency of Foxp3+RORγt+ T cells in regulating the inflammatory response induced by transfer of naive CD45RBhigh CD4+ T cells into RAG2-deficient mice. To monitor the suppressive capacities of the different Treg populations, we chose a suboptimal 1:4 ratio between Foxp3+ and naive T cells. This ratio does not prevent colitis, but induces a certain degree of inflammation suitable to detect differences in the suppressive capacity of Treg subsets.44 When transferred alone, naive T cells induced severe signs of colitis and weight loss in RAG2-deficient mice, starting from week 3 after transfer (Figure 7a). Cotransfer of Foxp3+ Tregs (Treg/T naive ratio of 1:4) led to a reduction in signs of colitis and weight loss; however, these mice still displayed a colitis phenotype as compared with phosphate-buffered saline-treated control mice. Intriguingly, mice that were cotransferred in the same 1:4 ratio with Foxp3+RORγt+ T cells were completely protected from colitis induction (Figure 7a). In accordance with these results, we observed pronounced inflammatory cellular infiltration in the mucosa and submucosa, as well as significant epithelial destruction, in the group of mice that were transferred with naive CD4+ T cells only (Figure 7b). Whereas mice that were cotransferred with Foxp3+ Tregs showed reduced but nonetheless substantial colonic pathology, cotransfer of Foxp3+RORγt+ T cells almost completely prevented colonic tissue damage (Figure 7b,c), together demonstrating a superior suppressive capacity of Foxp3+RORγt+ T cells as compared with Foxp3+ Tregs during T cell-mediated intestinal inflammation. We next assessed whether the presence of Foxp3+RORγt+ T cells influence the induction of RORγt+ IL-17-producing effector T cells in the adoptively transferred naive CD4+ T-cell population (CD90.1). As shown in Figure 7d, there was no significant difference in the frequency of RORγt+ T cells, as well as in the frequencies of IL-17- or interferon-γ-producing effector T cells in the colon of mice cotransferred with Foxp3+RORγt+ T cells as compared with Foxp3+ Tregs (Figure 7d), suggesting that a direct effect on Th17 induction could not explain the enhanced anti-inflammatory function of Foxp3+RORγt+ T cells. Of note, a small but sizable fraction of transferred CD90.2+ Foxp3+RORγt+ T cells expressed IL-17 at the time point of analysis, confirming that RORγt+ expression also correlates with the ability to express this cytokine in Foxp3+ cells (Supplementary Figure S8a). Nevertheless, the total number of IL-17+Foxp3+RORγt+ remained low in all organs, including the colonic LP (Supplementary Figure S8b).
Foxp3+RORγt+ T cells show increased suppressive capacity in comparison with Foxp3+ regulatory T cells (Tregs) in a transfer colitis model. CD90.1+CD4+CD45RBhigh T cells were sorted from CD90.1 mice and injected into RAG2-deficient mice to induce colitis. In addition, CD90.2+CD4+ Foxp3+RORγt− Tregs and Foxp3+RORγt+ T cells were sorted from Foxp3RFPRORγtGFP mice and were cotransferred in a ratio of 1:4. Mice were monitored over 4–7 weeks of the experiment. (a) Body weight changes over the time course of the experiment, (b) histopathology in colonic tissue, and (c) a summary graph of histopathological scores are shown for different experimental groups. Scale bars=200 μm. (d) Expression of the transcription factor RORγt and cytokine production in CD3+CD4+CD90.1+ T cells. Representative fluorescence-activated cell sorting (FACS) plots and frequencies in the different experimental groups are shown. (e) Total number of lamina propria lymphocytes isolated from the colon. Data are shown as mean±s.d. Experiments are representative for three independent experiments (a, b, d, upper panel) or pooled data from four independent experiments with 3–5 mice per experimental group. cLP, colonic lamina propria; Foxp3, forkhead box P3 transcription factor; IFN-γ, interferon-γ; IL-17, interleukin-17; PBS, phosphate-buffered saline; RORγt, retinoic acid-related orphan receptor-γt. *P<0.05, **P<0.01, ***P<0.001.
Moreover, we did not observe major differences in the expansion capacity of CD90.2+ Foxp3+RORγt+ T cells and Foxp3+ Tregs. More than 90% of cells from both populations stained positive for the proliferation marker Ki-67 in spleen and pLNs 2 weeks after transfer, resulting in low, but comparable, total amounts of CD90.2+ Foxp3+RORγt+ T cells and Foxp3+ Tregs in lymphoid organs at early (2 weeks) and late (4–7 weeks) time points after transfer (Supplementary Figures S9 and S10). Importantly, the superior suppressive capacity of Foxp3+RORγt+ T cells was also not caused by an enhanced general inhibitory effect on the expansion of transferred naive CD90.1+ T cells, as illustrated by similar effector T-cell/Treg ratios and comparable amounts of total CD90.1+ T cells recovered from the lymphoid organs of the different groups (Supplementary Figures S9 and S10).
However, cotransfer of Foxp3+RORγt+ T cells led to a strong reduction in total cellular infiltration into the colon (Figure 7e), decreased total numbers of CD90.1+ T cells, and a reduction in the total amount of IL-17- and interferon-γ-producing effector T cells in this organ (Supplementary Figures S10 and S11), suggesting that Foxp3+RORγt+ T cells exhibit a specific suppressive effect on inflammatory cell expansion/recruitment in the gut. Viewed as a whole, our data revealed a significantly enhanced suppressive capacity of Foxp3+RORγt+ T cells as compared with Foxp3+ Tregs under these proinflammatory conditions in the gut, suggesting that RORγt expression in Tregs contributes to an optimal regulatory function during gut-specific immune responses.
DISCUSSION
Foxp3+RORγt+ T cells were identified in both mice and humans. Initially, we reported that in mice, Foxp3+RORγt+ T cells secrete the anti-inflammatory cytokine IL-10 in vivo and are capable of suppressing activated CD4+ T cells in vitro.20 Although these findings pointed towards a regulatory function of these cells, subsequent studies demonstrated that Foxp3+ T cells can start to produce the proinflammatory cytokine IL-17, hence suggesting that these cells might contribute to inflammatory conditions. In humans, IL-17-producing Foxp3+ T cells were identified in peripheral blood as well as in the microenvironments of chronic inflammation and cancer.23, 45, 46, 47 Although most of these studies indicated that human Foxp3+IL-17+ cells retain regulatory capacity as long as they express Foxp3, it was suggested that they can foster the expression of proinflammatory cytokines and thus behave as “inflammatory” Tregs.23 In addition, in vitro studies showed that human RORγt+ Th17 cells preferentially differentiate from naïve Foxp3+ T cells that lose their Foxp3 expression48 that may eventually lead to conversion into pathogenic Th17 cells under inflammatory conditions.49 Additional studies in mice have shown that Foxp3+RORγt+ T cells have a regulatory function during autoimmune diabetes;50 nevertheless, it was proposed that these cells represent intermediates that can differentiate either toward Foxp3+RORγt− Tregs or RORγt+Foxp3− Th17 cells. Further studies suggested that in mice, Foxp3+ T cells also can lose Foxp3 expression and convert into Th17 cells or pathogenic “exFoxp3” cells.51, 52 With regard to these findings, it was therefore unclear whether those Foxp3+RORγt+ T cells that can be found in lymphoid organs and intestinal LP of mice represent a stable Treg subpopulation or just an intermediate stage during Treg or Th17 cell development. To answer this question, we generated Foxp3RFPRORγtGFP double reporter mice that enabled us to perform a comprehensive phenotypical and functional analysis of the Foxp3+RORγt+ T-cell population and to study their lineage stability.
Our data show that the frequency of Foxp3+RORγt+ T cells is strongly increased in the colon, dependent on the presence of a complex microbiota. Although we did not assess the frequency of these cells in other nonimmune tissues in this study, we have shown previously that Foxp3+RORγt+ T cells also exist in other tissues such as the lung, although at frequencies that were comparable to that found in the spleen, whereas the skin contained per se only very little frequencies of RORγt+ CD4+ T cells.20 In addition, recent work confirmed that Tregs isolated from intestinal LP express high levels of RORγt+, whereas fat tissue-resident Tregs as well as other peripheral Treg populations displayed only low levels of RORγt expression.9 This demonstrates that although low frequencies of Foxp3+RORγt+ T cells can be observed under normal physiologic conditions in different peripheral lymphoid and nonlymphoid tissues, a majority of these cells are clearly located within intestinal tissues.
Our transcriptome studies showed a 99% overlap in gene expression between Foxp3+RORγt+ T cells, Foxp3+ Tregs, and RORγt+ T cells, illustrating a high similarity between these CD4+ T-cell subsets. However, moving the focus onto differentially expressed genes revealed a closer relationship between Foxp3+RORγt+ T cells and Foxp3+ Tregs. However, a higher overlap between these two subsets did not result in a complete inhibition of RORγt+ T cell-associated gene expression. Interestingly, some Th17 lineage-defining genes were highly expressed in Foxp3+RORγt+ T cells (e.g., Rorc or Il23r), whereas the expression of characteristic cytokines (Il17a, Il17f, or Il21) was low compared with Th17 cells, indicating that these cells are functionally different. The low IL-17 expression in Foxp3+RORγt+ T cells might be explained by the only incomplete demethylation of the Il17a locus (when compared with RORγt+ T cells) that has been previously shown to play an important role for IL-17 expression and Th17 lineage specificity.53 Nonetheless, the expression pattern of specific chemokine receptors was very similar between Foxp3+RORγt+ and RORγt+ T cells, indicating that these cell types share a similar migration pattern. However, the short-term in vivo homing assay indicated that Foxp3+RORγt+ T cells show a migratory pattern being more similar to Foxp3+ Tregs than to RORγt+ T cells. Despite this, it is still possible that the tendencies toward an enhanced homing into gut and other mucosa-associated tissues would only become significant at time points later than 24 h after cell transfer or under inflammatory conditions. Intriguingly, Foxp3+RORγt+ T cells showed high expression of Irf4, Prdm1, and Il10. In a recent report, Cretney et al.27 demonstrated that Blimp-1 (encoded by Prdm1) expression was restricted to a population of effector Tregs. Furthermore, they could show that Blimp-1 and IRF4 acted together to induce IL-10 expression. Similar to Foxp3+RORγt+ T cells, these Blimp-1+ effector Tregs also expressed high levels of CD103, ICOS, and GITR. In addition, Foxp3+RORγt+ T cells, even when isolated from nonintestinal lymphoid tissues, expressed high levels of genes that were recently described to be overrepresented in small intestine LP Tregs, e.g., Gzmb and Ccr9.9 Thus, Foxp3+RORγt+ T cells express a set of effector Treg-specific “core-genes” plus a panel of genes that can confer gut specificity, together clearly indicating that these cells represent a highly suppressive gut-specific effector Treg lineage. Whether RORγt expression itself is of importance for the establishment of this unique phenotype remains to be studied.
In order to assess the stability of the Treg phenotype of Foxp3+RORγt+ T cells, we focused on the CpG methylation status in different gene loci that have been described to be of significance for Treg lineage stability.36, 38, 40 Our data showed full demethylation of the TSDR in Foxp3+RORγt+ T cells and substantial demethylation of Ctla4, Tnfrsf18, and Ikzf4, a clear indication of full Treg lineage stability and function according to Ohkura et al.40 Of note, the degree of demethylation in Ikzf4 was lower in Foxp3+RORγt+ T cells compared with Foxp3+ Tregs. It has been argued that downmodulation of Eos (encoded by Ikzf4) transcription is indicative of Tregs undergoing reprogramming into T helper-like cells.54 However, the overall methylation pattern in the Treg-specific epigenetic signature genes in Foxp3+RORγt+ T cells may rather support the idea that a fraction of Foxp3+RORγt+ T cells are peripherally induced and are on the way to acquire full demethylation in their Tnfrsf18 and Ikzf4 loci, as recently suggested.40 Further supporting this hypothesis, we also found a mixed pattern of Helios expression in Foxp3+RORγt+ T cells, indicating that the Foxp3+RORγt+ T cells observed under physiologic conditions may be a mixture of not only thymic but also peripherally induced origin, especially in the gut-associated tissues. It should be noted however that Helios expression may not unambiguously distinguish thymus-derived Tregs from pTregs, as several studies demonstrated that Foxp3+ pTregs may under certain conditions also express Helios.55
Regarding the stability of Foxp3 expression we observed a substantial degree of Foxp3 downregulation in Foxp3+RORγt+ T cells upon adoptive transfer into lymphopenic recipients. This observation was somewhat surprising as Foxp3+RORγt+ T cells displayed a fully demethylated TSDR that safeguards stable Foxp3 expression.34, 36 However, it is possible that some Foxp3+RORγt+ T cells only transiently lose Foxp3 expression in the lymphopenic environment. As recently shown by Miyao et al.,37 such latent Tregs retain a “Treg memory” and can rapidly reexpress Foxp3 expression upon provision of appropriate stimulatory signals. Another hypothesis for the observed instability is supported by the finding that Foxp3+RORγt+ T cells contain peripherally induced Tregs. We could recently show that Foxp3 expression in thymus-derived Tregs is more stable in an adoptive transfer model when compared with pTregs.44 Therefore, these pTregs, which presumably originate from mLNs in our transfer approach, might account for the increased loss of Foxp3 expression that we observed after transfer of Foxp3+RORγt+ T cells. Importantly, we did not observe substantial reprogramming of Foxp3+RORγt+ T cells into RORγt+ Th17 cells, even under the proinflammatory conditions in the transfer colitis model. In contrast, our findings demonstrate that RORγt+Foxp3+ T cells represent a rather stable effector Treg lineage with a highly potent suppressive capacity in the transfer colitis model. Of note, it is possible that the pool of Foxp3+RORγt+ T cells that can be found in other models of inflammation22, 50 may contain, in addition to the stable RORγt+ effector Treg lineage that we described here, an increased proportion of developmental Foxp3+RORγt+ intermediates that eventually differentiate into Foxp3− Th17 cells.
In summary, our phenotypic and functional characterization of the microbiota-induced RORγt+Foxp3+ T cells revealed a hybrid phenotype of this T-cell population, showing both transcriptional and epigenetic profiles of Th17 cells and Tregs. Their specific molecular patterns provide them with stable and unique functional properties of effector Tregs for the very efficient suppression of gut-specific inflammatory responses.
METHODS
Mice. C57BL/6 mice aged 14–16 weeks, female RAG2-deficient mice aged 3–8 weeks, and female CD90.1 C57BL/6 mice aged 4–8 weeks were bred in-house (Helmholtz Centre for Infection Research, Braunschweig, Germany; TWINCORE, Hanover, Germany) or purchased (C57BL/6) from Charles River (Sulzfeld, Germany). RORc(gt)-GfpTG mice20 were crossed to Foxp3-IRES-mRFP (FIR) mice26 to obtain Foxp3RFPRORγtGFP double reporter mice. The mice were bred and maintained under specific pathogen-free conditions at our animal facilities (Helmholtz Center for Infection Research). The mice used for this study were rederived to possess a complex microbiota inclusive of, among others, segmented filamentous bacteria.24 Where indicated, mice with a minimal microflora with close similarity to the altered Schaedler flora were used.25 All animal experiments were performed in compliance with the German animal protection law (TierSchG BGBl. I S. 1105; 25.05.1998). All mice were housed and handled in accordance with good animal practice as defined by FELASA (Federation of European Laboratory Animal Science Associations) and the national animal welfare body GV-SOLAS (Gesellschaft für Versuchstierkunde/Society for Laboratory Animal Science). All animal experiments were approved by the Lower Saxony Committee on the Ethics of Animal Experiments as well as the responsible state office (Lower Saxony State Office of Consumer Protection and Food Safety) under the permit numbers 33.9-42502-04-10/0039 and 33.9-42502-04-12/1012.
Reagents and antibodies. The following antibodies were purchased from Affymetrix (Frankfurt am Main, Germany): CD3 APC-eFluor780 (17A2); CD3 PerCPCy5.5 (145-2C11); CD4 Alexa488 (GK1.4); CD11c APC (N418); CD44 PerCPCy5.5 (IM7); CD45RB PE (363.16A); CD62L APC (MEL-14); CD90.2 PE-Cy7 (53-2.1); CD199 (CCR9) PE (eBioCW-1.2 eBio); Foxp3 Alexa488 (FJK-16s); Foxp3 eFluor450 (FJK-16s); IL-17 APC (eBio17B7); interferon-γ FITC (XMG1.2); Helios Alexa647 (22F6); RORγt PE (B2D). The following antibodies were purchased from BioLegend (Fell, Germany): CD3 Pacific Blue (17A2); CD4 BV605 (RM4-5); CD4 Pacific Blue (RM4-5); CD4 APC (RM4-5); CD4 PE-Cy7 (RM4-5); CD8 APC (53-6.7); CD25 BV605 (PC61); CD194 (CCR4) APC (2G12 BL); ICOS PerCPCy5.5 (C398.4A); Ki-67 APC (16A8). The following antibodies were purchased from BD Biosciences (Heidelberg, Germany): CD19 APC (1D3) and CD196 (CCR6) Alexa647 (140706 BD). To discriminate live and dead cells, the following live/dead staining reagents were used: 4′,6-diamidino-2-phenylindole (DAPI; Sigma-Aldrich, Schnelldorf, Germany); LIVE/DEAD fixable Aqua dead stain kit (Life Technologies, Darmstadt, Germany), and LIVE/DEAD fixable Blue dead stain kit (Life Technologies). The Foxp3/Transcription Factor Staining Buffer Set (Affymetrix) was used for fixation and intracellular staining.
Cell isolation from organs. Single-cell suspensions were prepared from spleen, pLN, mLN, or Peyer’s patches by smashing organs through a 100 μm mesh. Lysis of erythrocytes was performed for spleens. Colon and small intestine were physically emptied, opened longitudinally, and cut into 2–3 cm pieces. Tissue pieces were incubated in PBS containing 30 mM EDTA (Sigma-Aldrich) for 30 min, washed vigorously with PBS to remove remaining mucus, cut into 1–2 mm pieces, and digested in prewarmed Iscove’s modified Dulbecco’s medium (Life Technologies/Gibco, Darmstadt, Germany) containing 1 mg ml−1 Collagenase D (Roche, Diagnostics, Mannheim, Germany) and 100 μg ml−1 DNaseI (Roche). The supernatant was filtered and the remaining tissue was smashed through a 100 μm mesh. LP cells were separated using a 40%/80% gradient (Percoll solution, GE Healthcare, Freiburg, Germany; or Easycoll, Merck Millipore/Biochrom, Berlin, Germany; 900 g, 20 min, 20 °C, no break). The interphase was harvested and washed. Cells were resuspended in PBS containing 0.2% bovine serum albumin or medium containing fetal calf serum before extra/intracellular staining.
Flow cytometry. Cell suspensions were incubated with PBS containing 0.2% bovine serum albumin and 1% anti-mouse CD16/CD32 antibody (BioXcell, West Lebanon, NH) for 5 min on ice. Live/dead staining was performed in PBS using the LIVE/DEAD fixable Aqua or Blue dead staining kit according to the manufacturer’s recommendations, or DAPI prior measurement. Surface staining was performed for 15 min on ice in PBS containing 0.2% bovine serum albumin. Intracellular staining was performed using Foxp3/Transcription Factor Staining Buffer Set according to the manufacturer’s recommendations. To block unspecific antibody bindings, 40 μg ml−1 rat IgG (Dianova, Hamburg, Germany) was used. Cells were washed, resuspended in PBS containing 0.2% bovine serum albumin, and measured using BD LSRII SORP (BD Biosciences). Cell sorting was performed on FACSAria II (BD Biosciences) in the cell sorting facility of Helmholtz Centre for Infection Research. Data were analyzed using the FlowJo software (Tree Star, Ashland, OR).
Sorting of ex vivo -isolated CD4+ Foxp3+/−RORγt+/− subsets. CD4+ T cells were enriched by CD4 (L3T4) MicroBeads (Miltenyi Biotec, Bergisch Gladbach, Germany) followed by AutoMACS separation from spleen and/or LN of Foxp3RFPRORγtGFP mice and stained for CD3, CD4, CD8, CD11c, and CD19. Foxp3+/−RORγt+/− cell subsets were sorted from the CD3+CD4+ or CD4+CD8−CD11c−CD19− population according to the reporter signals.
Bisulfite pyrosequencing. Genomic DNA was isolated from ex vivo-isolated Foxp3+/−RORγt+/− subsets of different organs using NucleoSpin Tissue/Tissue XS kit (Macherey-Nagel, Düren, Germany) and bisulfite converted using the EZ DNA Methylation Kit (Zymo Research, Freiburg, Germany) according to the manufacturer’s instructions. Differentially methylated regions were amplified by PCR containing 10 ng of bisulfite-converted genomic DNA, HotStar Taq PCR buffer (Qiagen, Hilden, Germany), 1 U HotStar Taq DNA polymerase, 2.5 mM MgCl2, and 0.38 μM forward and reverse primers in a final volume of 50 μl reaction (cycle: 95 °C for 15 min; 50 × 95 °C for 30 s, 57–62 °C for 1 min, 72 °C for 1 min; 72 °C for 7 min). The PCR product was analyzed by gel electrophoresis. Then, 20–40 μl of the PCR product, Pyromark Gold Q96 reagents (Qiagen), Pyromark buffers (Qiagen), Streptavidin Sepharose (GE Healthcare), and sequencing primers were used for pyrosequencing on a PSQ96MA (Qiagen) according to the manufacturer’s protocol. The information of primers is listed in Supplementary Table S1. Male mice were used for DNA methylation analysis to avoid artificial recalculation because of X-chromosome inactivation in female mice.
Microarray analysis, hierarchical clustering, and heatmap generation. RNA was isolated from ex vivo-isolated Foxp3+/−RORγt+/− T cell subsets of pooled spleen and LN using RNeasy Plus Mini Kit (Qiagen) according to the manufacturer’s instructions. DNA microarray hybridization was performed by the genome facility of the Helmholtz Centre for Infection Research. Quality and integrity of total RNA was controlled on Agilent Technologies 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA). RNA samples were loaded on a SurePrint G3 Mouse GE 8x60K one-color array (Agilent). Data processing was done using the Limma package56 from the R/Bioconductor framework.57 After passing array quality assessment/control, unprocessed array intensity values were read-in using function read.maimages and background corrected with the improved saddle-point approximation to maximum likelihood method58 using intensity offset of 16. Array intensities have been normalized between arrays using quantile normalization59 to obtain similar distributions of expression intensities. Normalized intensities of probes targeting the same gene were mean averaged with function avereps to compute a single expression value per gene. Differentially expressed genes were determined using the lmFit function for linear modeling and by computation of moderated t-statistics and log-odds with function eBayes.60 P-values from the moderated t-tests have been corrected for multiple testing using the Benjamini–Hochberg procedure for controlling the false discovery rate. Finally, the set of differentially expressed genes was filtered by fold change (|log2 FC|≥1.5) and corrected P-value (P≤0.05). All array data generated in this study were deposited in the Gene Expression Omnibus database and are available under accession number GSE60059.
Averaged, normalized, scaled and mean centered expression values of 858 genes that are differentially expressed (|log2 FC| ≥1.5; P≤0.05) in pairwise comparison Foxp3+RORγt− vs. Foxp3−RORγt+ T cells (two independent experiments) served as input for hierarchical clustering and heatmap generation using the R package (www.r-project.org) pheatmap.
TCR repertoire analysis. RNA was isolated from 2.5 × 104–7.2 × 104 ex vivo-isolated Foxp3+/−RORγt+/− T cell subsets of pooled spleen and LN from individual Foxp3RFPRORγtGFP double reporter mice using RNeasy Plus Mini Kit (Qiagen) according to the manufacturer’s instructions. Complementary DNA templates were synthesized using Transcriptor First Strand cDNA Synthesis Kit (Roche) according to the manufacturer’s recommendation. Amplicon libraries of rearranged TRAV12 CDR3 regions for the Roche/454 Genome Sequencer FLX+ System were generated as described before.31 Amplicons were purified by agarose gel electrophoresis and QIAquick Gel Extraction Kit (Qiagen), and quantified by Quant-iT dsDNA HS Assay Kit (Life Technologies/Invitrogen, Darmstadt, Germany). High-throughput sequencing of amplicons was performed on a Genome Sequencer FLX+ system (Roche) according to the manufacturer’s recommendations. Productive rearrangements, J usage, and CDR3α regions were defined by comparing nucleotide sequences with the reference sequences from IMGTs, the international ImMunoGeneTics information systems (http://www.imgt.org; IMGT/HighV-QUEST).61, 62 Diversity was calculated as “effective number of sequences” by the exponential of Shannon index.
In vitro stability assay. 1 × 105 ex vivo-isolated Foxp3+/−RORγt+/− cells from pooled spleen and LN were cultured in a flat-bottom plate coated with anti-CD3 (3 μg ml−1; 145-2C11; Affymetrix) and anti-CD28 (5 μg ml−1; 37.51; Affymetrix) antibodies in Iscove’s modified Dulbecco’s medium (Gibco) containing 10% fetal calf serum (Biochrom), 50 U ml−1 penicillin/streptomycin (Gibco), 1 mM sodium pyruvate (Gibco), 50 μM 2-mercaptoethanol (Gibco), 25 mM HEPES (Biochrom), and 1% NEAA (Gibco) in the presence of recombinant mouse IL-2 (10 ng ml−1; R&D Systems, Wiesbaden, Germany) and recombinant mouse IL-7 (10 ng ml−1; R&D) for 4 days. Cells were harvested, stained for CD3 and CD4, and analyzed for expression of Foxp3mRFP and RORγtGFP.
Adoptive transfer and in vivo stability assay. 7 × 104 ex vivo-isolated Foxp3+RORγt+, Foxp3−RORγt+, or Foxp3+RORγt− CD4+ T cells of pooled spleen and LN from CD90.2 mice were mixed with 2.8 × 105 FACS-sorted naive CD4+ T cells (CD4+CD45RBhighCD8−CD11c−CD19−) from CD90.1 congenic mice in a 1 to 4 ratio in PBS. 3.5 × 105 cells were adoptively transferred to RAG2-deficient mice aged 3–8 weeks by intraperitoneal injection. Control groups received the same amount of naive T cells only or PBS. Cells from spleen, pLN, mLN, and colon were isolated 2 weeks or 4–7 weeks after transfer. Living cell count was analyzed on BD Accuri C6 (BD Biosciences). The stability of Foxp3 and RORγt expression was assessed based on their reporter signals in living cells stained for CD3, CD4, and CD90.2. Detailed analysis of the cytokine production and transcription factor expression in CD90.1+ or CD90.2+ cell populations was carried out with cells stimulated with phorbol 12-myristate 13-acetate (10 ng ml−1; Sigma-Aldrich) and ionomycin (0.5 μg ml−1; Sigma-Aldrich) for 4 h. Brefeldin A (10 μg ml−1; Sigma-Aldrich) was added for the last 2 h. Stimulated cells were stained extracellularly for CD3, CD4, and CD90.2 and intracellularly for RORγt, Foxp3, IL-17A, and interferon-γ. Living cells were discriminated using LIVE/DEAD fixable Blue dead cell stain kit (Life Technologies) or DAPI (Sigma-Aldrich) for unfixed cells and measured by FACS.
Histology. Colon tissue from RAG2-deficient mice described above was sampled and fixed in paraformaldehyde at the date of end-point analysis. Samples were processed to paraffin blocks, sectioned, and stained with hematoxylin and eosin at the Mouse Pathology Department of Helmholtz Centre for Infection Research. Samples were scored as described previously.24 In brief, the presence of rare inflammatory cells in the LP were counted as: 0, increased numbers of inflammatory cells; 1, confluence of inflammatory cells; 2, extending into the submucosa; and 3, transmural extension of the inflammatory cell infiltrate. For epithelial damage, absence of mucosal damage was counted as 0, discrete focal lymphoepithelial lesions were counted as 1, mucosal erosion/ulceration was counted as 2, and a score of 3 was given for extensive mucosal damage and extension through deeper structures of the bowel wall. The two subscores were added and the combined histological score ranged from 0 (no changes) to 6 (extensive cell infiltration and tissue damage).
Statistics. Statistical analysis was performed with GraphPad Prism software (La Jolla, CA) using the Mann–Whitney two-tailed t-test and Bonferroni correction (homing assay). For in vivo stability, statistical analysis was performed using two-way analysis of variance with Bonferroni’s multiple comparison test. Error bars indicate the s.d. in all the figures. P-values for significance were indicated with *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001, or NS (not significant).
Accession codes
References
Bilate, A.M. & Lafaille, J.J. Induced CD4+Foxp3+ regulatory T cells in immune tolerance. Annu. Rev. Immunol. 30, 733–758 (2012).
Cording, S. et al. The intestinal micro-environment imprints stromal cells to promote efficient Treg induction in gut-draining lymph nodes. Mucosal Immunol. 7, 359–368 (2014).
Haribhai, D. et al. A central role for induced regulatory T cells in tolerance induction in experimental colitis. J. Immunol. 182, 3461–3468 (2009).
Josefowicz, S.Z. et al. Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature 482, 395–399 (2012).
Lathrop, S.K. et al. Peripheral education of the immune system by colonic commensal microbiota. Nature 478, 250–254 (2011).
Samstein, R.M., Josefowicz, S.Z., Arvey, A., Treuting, P.M. & Rudensky, A.Y. Extrathymic generation of regulatory T cells in placental mammals mitigates maternal-fetal conflict. Cell 150, 29–38 (2012).
Huehn, J. & Hamann, A. Homing to suppress: address codes for Treg migration. Trends Immunol. 26, 632–636 (2005).
Liston, A. & Gray, D.H. Homeostatic control of regulatory T cell diversity. Nat. Rev. Immunol. 14, 154–165 (2014).
Feuerer, M., Hill, J.A., Kretschmer, K., von Boehmer, H., Mathis, D. & Benoist, C. Genomic definition of multiple ex vivo regulatory T cell subphenotypes. Proc. Natl. Acad. Sci. USA 107, 5919–5924 (2010).
Cretney, E., Kallies, A. & Nutt, S.L. Differentiation and function of Foxp3(+) effector regulatory T cells. Trends Immunol. 34, 74–80 (2013).
Delacher, M., Schreiber, L., Richards, D.M., Farah, C., Feuerer, M. & Huehn, J. Transcriptional control of regulatory T cells. Curr. Top. Microbiol. Immunol. 381, 83–124 (2014).
Koch, M.A., Tucker-Heard, G., Perdue, N.R., Killebrew, J.R., Urdahl, K.B. & Campbell, D.J. The transcription factor T-bet controls regulatory T cell homeostasis and function during type 1 inflammation. Nat. Immunol. 10, 595–602 (2009).
Wang, Y., Su, M.A. & Wan, Y.Y. An essential role of the transcription factor GATA-3 for the function of regulatory T cells. Immunity 35, 337–348 (2011).
Wohlfert, E.A. et al. GATA3 controls Foxp3(+) regulatory T cell fate during inflammation in mice. J. Clin. Invest. 121, 4503–4515 (2011).
Chaudhry, A. et al. CD4+ regulatory T cells control TH17 responses in a Stat3-dependent manner. Science 326, 986–991 (2009).
Zheng, Y. et al. Regulatory T-cell suppressor program co-opts transcription factor IRF4 to control T(H)2 responses. Nature 458, 351–356 (2009).
Chung, Y. et al. Follicular regulatory T cells expressing Foxp3 and Bcl-6 suppress germinal center reactions. Nat. Med. 17, 983–988 (2011).
Linterman, M.A. et al. Foxp3+ follicular regulatory T cells control the germinal center response. Nat. Med. 17, 975–982 (2011).
Ivanov II et al. The orphan nuclear receptor RORgammat directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell 126, 1121–1133 (2006).
Lochner, M. et al. In vivo equilibrium of proinflammatory IL-17+ and regulatory IL-10+ Foxp3+ RORgamma t+ T cells. J. Exp. Med. 205, 1381–1393 (2008).
Zhou, L. et al. TGF-beta-induced Foxp3 inhibits T(H)17 cell differentiation by antagonizing RORgammat function. Nature 453, 236–240 (2008).
Esposito, M. et al. IL-17- and IFN-gamma-secreting Foxp3+ T cells infiltrate the target tissue in experimental autoimmunity. J. Immunol. 185, 7467–7473 (2010).
Kryczek, I. et al. IL-17+ regulatory T cells in the microenvironments of chronic inflammation and cancer. J. Immunol. 186, 4388–4395 (2011).
Lochner, M. et al. Microbiota-induced tertiary lymphoid tissues aggravate inflammatory disease in the absence of RORgamma t and LTi cells. J. Exp. Med. 208, 125–134 (2011).
Stehr, M. et al. Charles River altered Schaedler flora (CRASF) remained stable for four years in a mouse colony housed in individually ventilated cages. Lab. Anim. 43, 362–370 (2009).
Wan, Y.Y. & Flavell, R.A. Identifying Foxp3-expressing suppressor T cells with a bicistronic reporter. Proc. Natl. Acad. Sci. USA 102, 5126–5131 (2005).
Cretney, E. et al. The transcription factors Blimp-1 and IRF4 jointly control the differentiation and function of effector regulatory T cells. Nat. Immunol. 12, 304–311 (2011).
Kim, S.V. et al. GPR15-mediated homing controls immune homeostasis in the large intestine mucosa. Science 340, 1456–1459 (2013).
Smith, P.M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Hamann, A., Klugewitz, K., Austrup, F. & Jablonski-Westrich, D. Activation induces rapid and profound alterations in the trafficking of T cells. Eur. J. Immunol. 30, 3207–3218 (2000).
Fohse, L. et al. High TCR diversity ensures optimal function and homeostasis of Foxp3+ regulatory T cells. Eur. J. Immunol. 41, 3101–3113 (2011).
Fisson, S. et al. Continuous activation of autoreactive CD4+ CD25+ regulatory T cells in the steady state. J. Exp. Med. 198, 737–746 (2003).
Fohse, L. et al. Differential postselection proliferation dynamics of alphabeta T cells, Foxp3+ regulatory T cells, and invariant NKT cells monitored by genetic pulse labeling. J. Immunol. 191, 2384–2392 (2013).
Huehn, J., Polansky, J.K. & Hamann, A. Epigenetic control of FOXP3 expression: the key to a stable regulatory T-cell lineage? Nat. Rev. Immunol. 9, 83–89 (2009).
Wilson, C.B., Rowell, E. & Sekimata, M. Epigenetic control of T-helper-cell differentiation. Nat. Rev. Immunol. 9, 91–105 (2009).
Floess, S. et al. Epigenetic control of the foxp3 locus in regulatory T cells. PLoS Biol. 5, e38 (2007).
Miyao, T. et al. Plasticity of Foxp3(+) T cells reflects promiscuous Foxp3 expression in conventional T cells but not reprogramming of regulatory T cells. Immunity 36, 262–275 (2012).
Polansky, J.K. et al. DNA methylation controls Foxp3 gene expression. Eur. J. Immunol. 38, 1654–1663 (2008).
Ohkura, N., Kitagawa, Y. & Sakaguchi, S. Development and maintenance of regulatory T cells. Immunity 38, 414–423 (2013).
Ohkura, N. et al. T cell receptor stimulation-induced epigenetic changes and Foxp3 expression are independent and complementary events required for Treg cell development. Immunity 37, 785–799 (2012).
Thornton, A.M. et al. Expression of Helios, an Ikaros transcription factor family member, differentiates thymic-derived from peripherally induced Foxp3+ T regulatory cells. J. Immunol. 184, 3433–3441 (2010).
Yang, B.H. et al. Development of a unique epigenetic signature during in vivo Th17 differentiation. Nucleic Acids Res. 43, 1537–1548 (2015).
Izcue, A., Coombes, J.L. & Powrie, F. Regulatory lymphocytes and intestinal inflammation. Annu. Rev. Immunol. 27, 313–338 (2009).
Huang, Y.J. et al. Induced and thymus-derived Foxp3(+) regulatory T cells share a common niche. Eur. J. Immunol. 44, 460–468 (2014).
Ayyoub, M. et al. Human memory FOXP3+ Tregs secrete IL-17 ex vivo and constitutively express the T(H)17 lineage-specific transcription factor RORgamma t. Proc. Natl. Acad. Sci. USA 106, 8635–8640 (2009).
Beriou, G. et al. IL-17-producing human peripheral regulatory T cells retain suppressive function. Blood 113, 4240–4249 (2009).
Voo, K.S. et al. Identification of IL-17-producing FOXP3+ regulatory T cells in humans. Proc. Natl. Acad. Sci. USA 106, 4793–4798 (2009).
Valmori, D., Raffin, C., Raimbaud, I. & Ayyoub, M. Human RORgammat+ TH17 cells preferentially differentiate from naive FOXP3+Treg in the presence of lineage-specific polarizing factors. Proc. Natl. Acad. Sci. USA 107, 19402–19407 (2010).
Komatsu, N. et al. Pathogenic conversion of Foxp3+ T cells into TH17 cells in autoimmune arthritis. Nat. Med. 20, 62–68 (2014).
Tartar, D.M. et al. FoxP3+RORgammat+ T helper intermediates display suppressive function against autoimmune diabetes. J. Immunol. 184, 3377–3385 (2010).
Yang, X.O. et al. Molecular antagonism and plasticity of regulatory and inflammatory T cell programs. Immunity 29, 44–56 (2008).
Zhou, X. et al. Instability of the transcription factor Foxp3 leads to the generation of pathogenic memory T cells in vivo. Nat. Immunol. 10, 1000–1007 (2009).
Thomas, R.M., Sai, H. & Wells, A.D. Conserved intergenic elements and DNA methylation cooperate to regulate transcription at the il17 locus. J. Biol. Chem. 287, 25049–25059 (2012).
Sharma, M.D. et al. An inherently bifunctional subset of Foxp3+ T helper cells is controlled by the transcription factor eos. Immunity 38, 998–1012 (2013).
Shevach, E.M. & Thornton, A.M. tTregs, pTregs, and iTregs: similarities and differences. Immunol. Rev. 259, 88–102 (2014).
Smyth, G.K. Limma: linear models for microarray data In Bioinformatics and Computational Biology Solutions using R and Bioconductor (Gentleman, R.V.C., Dudoit, S., Irizarry, R. & Huber, W., eds) 397–420 (Springer, New York, 2005).
Gentleman, R.C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Silver, J.D., Ritchie, M.E. & Smyth, G.K. Microarray background correction: maximum likelihood estimation for the normal-exponential convolution. Biostatistics 10, 352–363 (2009).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3 Article 3 (2004).
Brochet, X., Lefranc, M.P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Lefranc, M.P. et al. IMGT, the international ImMunoGeneTics information system. Nucleic Acids Res. 37, D1006–D1012 (2009).
Acknowledgements
We thank Lothar Gröbe, Beate Pietzsch, Maria Ebel, Zharah Fiebig, and Sabrina Woltemate for technical assistance. This work was supported by grants from the German Research Foundation (KFO 250 to T.S., J.H., and M.L. and SFB 900/Z1 to S.S.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
Supplementary information
Rights and permissions
About this article
Cite this article
Yang, BH., Hagemann, S., Mamareli, P. et al. Foxp3+ T cells expressing RORγt represent a stable regulatory T-cell effector lineage with enhanced suppressive capacity during intestinal inflammation. Mucosal Immunol 9, 444–457 (2016). https://doi.org/10.1038/mi.2015.74
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2015.74
This article is cited by
-
Microbiota-assisted iron uptake promotes immune tolerance in the intestine
Nature Communications (2023)
-
Early-life interactions between the microbiota and immune system: impact on immune system development and atopic disease
Nature Reviews Immunology (2023)
-
Regulatory T cells in the face of the intestinal microbiota
Nature Reviews Immunology (2023)
-
Mining the microbiota to identify gut commensals modulating neuroinflammation in a mouse model of multiple sclerosis
Microbiome (2022)
-
TLR4 regulates RORγt+ regulatory T-cell responses and susceptibility to colon inflammation through interaction with Akkermansia muciniphila
Microbiome (2022)