Abstract
Due to a limited range of immunoglobulin (Ig) genes, cattle and several other domestic animals rely on postrecombinatorial amplification of the primary repertoire. We report that activation-induced cytidine deaminase (AID) is strongly expressed in the fetal bovine ileal Peyer’s patch and spleen but not in fetal bone marrow. The numbers of IGHV (immunoglobulin heavy chain variable) mutations correlate with AID expression. The mutational profile in the fetuses is similar to postnatal and immunized calves, with targeting of complementarity-determining region (CDR) over framework region (FR), preference of replacement over silent mutations in CDRs but not in FRs, and targeting of the AID hotspot motif RGYW/WRCY. Statistical analysis indicates negative selection on FRs and positive selection on CDRs. Our results suggest that AID-mediated somatic hypermutation and selection take place in bovine fetuses, implying a role for AID in the diversification of the primary antibody repertoire in the absence of exogenous antigens.
Similar content being viewed by others
Introduction
In comparison to mouse and man, lymphoid differentiation and somatic recombination in several domestic animals, including rabbit, chicken and sheep, yield a relatively small B-cell pool with a restricted antibody repertoire.1 In these latter species, fetal and young animal gut-associated lymphoid tissue hosts an actively proliferating population of B cells, which generates the peripheral B-cell pool and the final primary (pre-immune response) antibody repertoire.2, 3 In ruminants, the amplification of naive B cells and primary repertoire expansion is thought to take place in the ileal Peyer’s patch (IPP).4, 5 Somatic hypermutation (SHM) and gene conversion (GC) have been suggested to expand the limited antibody repertoire in ruminants after recombination but before encountering external antigens,6, 7, 8, 9 although evidence for GC in ruminants is not as strong as in rabbit10, 11 or chicken.12, 13
SHM and GC are well-characterized as secondary diversification processes mediated by activation-induced cytidine deaminase (AID) upon encountering an external antigen. AID attacks single-stranded DNA during transcription and deaminates cytosines to uracils.14 AID preferentially targets C residues in the third position in WRCY hotspot motifs (W=A/T, R=A/G, Y=C/T).15, 16, 17, 18 If the resulting uracil is not recognized and removed by base excision or mismatch repair pathways, it may be used as a template during replication. The outcome is often a mutation regardless of the mechanism.19, 20, 21, 22 Mutations caused by AID activity are targeted to a region extending about 2 kb downstream from the Ig promoter, including the VDJ-coding exon and the J-region intron.23 In SHM, U:G mismatches recruit error-prone DNA repair machinery, which increases the mutation rate by up to a million fold, as recently reviewed by Storck et al.24 The number of mutations is significantly higher in the complementarity-determining regions (CDRs) than in the framework regions (FRs). Mutations in the CDRs enable the generation of high-affinity antibodies.23, 25
To our knowledge, this is the first study in fetal ruminants that directly links AID to IPP or to primary antibody formation. mRNA expression of bovine AID and its mutagenic activity has been analyzed in neonatal and adult lymphoid tissues, although not in IPP.9 In this paper, we demonstrate AID expression in bovine fetal IPP and also in fetal spleen before exposure to exogenous antigens. The mutational profiles of immunoglobulin heavy chain variable (IGHV) genes in fetus are compared with those generated by antigen-induced AID-mediated SHM.
Results
AID is expressed in bovine fetal IPP and spleen
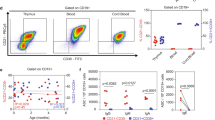
The expression of AID mRNA was analyzed by reverse transcription-quantitative PCR (RT-qPCR) in the fetal ileum, spleen, lymph node and bone marrow, with the muscle as a negative control. The AID expression level [as mean of glyceraldehyde 3-phosphate dehydrogenase (GAPDH)-normalized cycle threshold values±s.d.] was 11.0±3.8 in the ileum (n=21), 9.9±3.3 in the spleen (n=20), 12.8±2.3 in the lymph node (n=16), 16.2±1.5 in the bone marrow (n=14), and 16.5±3.7 in the muscle (n=10). AID expression in the ileum and spleen differed significantly from the expression in the bone marrow and muscle, as assessed using nonparametric Kruskal–Wallis analysis of variance (χ2=35.5, P<2 × 10−16) followed by multiple comparisons using Nemenyi–Damico–Wolfe–Dunn test at α=0.01. Looking at the lymphoid tissues at various stages of development, no statistically significant differences in AID expression were observed in any tissue, apart from the high level of expression in the adult lymph node (Figure 1).
Activation-induced cytidine deaminase (AID) mRNA expression analyzed by reverse transcription-quantitative PCR. Negatives of GADPH (glyceraldehyde 3-phosphate dehydrogenase)-normalized cycle threshold values (–ΔCt, mean±s.d.) are shown in ileum, spleen, bone marrow and lymph node for three fetal stages and for adults. The dashed lines represent mean±s.d. of AID expression in the muscle used as the negative control (N=12). gd, gestational days.
In the ileum, immunohistochemistry showed strong positive staining for AID in the IPP B-cell follicles (Figure 2a,b). The follicles were AID positive from the stage they become detectable at approximately 195 gestation days (gd). In the fetal lymph nodes, AID-positive follicles were observed only occasionally (Figure 2c,d). In the fetal spleen and bone marrow, the distribution of AID protein could not be reliably analyzed due to non-specific binding of the primary antibody to non-lymphoid cells. Adult lymph node was used as a positive control and displayed the characteristic germinal center staining pattern (not shown).26
Activation-induced cytidine deaminase (AID) expression and B lymphocyte proliferation in fetal bovine lymphoid tissues. (a) AID expression in the ileum of a 238–gestational days (gd) fetus. (b) Higher magnification of the area indicated by the box. (c) AID expression in the lymph node of a 250–gd fetus. (d) Higher magnification of the area indicated by the box. (e–g) Double immunofluorescence for the B-cell marker CD79α (green) and the proliferation marker Ki67 antigen (red) in the ileum of a 260–gd fetus (e), in a lymph node of a 250–gd fetus (f), and in the spleen of a 227–gd fetus (g). Bars=100 μm.
To assess the level of B-cell production in the various lymphoid tissues, we also performed double immunofluorescence staining for the B-cell marker CD79α+ and for the proliferation marker Ki67 antigen (Figure 2e–g). CD79α+ B cells proliferate more actively in the fetal IPP (median 75% Ki67+; s.d. 9.4%; n=7) than in the lymph nodes (cortical follicles; 9.0±6.7%; n=7; P=0.002) and spleen (15±10%; n=9; P=0.002).
Fetal IGHV cDNAs show signs of hypermutation and selection, but the overall mutational load is lower than in postnatal animals
In order to characterize the mutation patterns in fetal IGHV sequences, we constructed IGHV cDNA libraries from the ileum, spleen, and bone marrow of two fetuses (240 and 270 gd). To compare the naive fetal antibody repertoire to that after secondary antibody responses, we also generated cDNA libraries from the ileum of a 51-day-old calf and from a peripheral lymph node of an immunized calf. The germline reference sequences were obtained from the available bovine genome sequencing data27 and by sequencing the functional IGHV genes in four additional animals to a total of 35 functional IGHV genes. The fetal cDNAs aligned to the FR1–FR3 region of the germline sequences, with an average pairwise identity of 99.6%. For the 51-day calf and the immunized calf, the average identities were 97.4% and 92.4%.
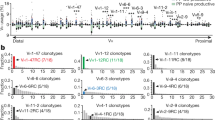
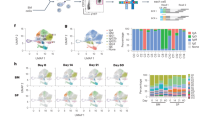
Mutation pattern characteristic of SHM was observed in cDNAs derived from the fetal IPP and spleen as well as from the postnatal samples, although fetal sequences contained fewer mutations (Figure 3 and Table 1). The total mutation load was 5–9 mutations per kilobase in fetal samples from the IPP and spleen, 26 per kilobase in the IPP of the 51-day calf, and 80 per kilobase in the lymph node of the 2-year-old immunized calf. The ratio of mutations in CDRs vs. FRs was 5.6 in fetuses (average of all ileal and splenic samples), 6.7 in the 51-day calf, and 3.4 in the immunized calf. The ratio of replacement over silent mutations was 3.3 in fetuses, 3.1 in the 51-day calf, and 2.1 in the immunized calf. In the fetal bone marrow, few mutations were observed, and the mutations were not concentrated in the CDR regions (Figure 3 and Table 1).
Mutational loads in framework (FR) and complementarity-determining (CDR) regions of immunoglobulin heavy chain variable cDNA sequences from fetal and calf lymphoid tissues. Bars represent mutations per 1 kb FR or CDR sequence analyzed. No CDR mutations were observed in the bone marrow of the 270-day fetus. For numbers of cDNA sequences analyzed, see Table 1.
Negative selection had occurred in FRs, whereas positive selection had occurred in the CDRs in both the fetus and calf data sets, based on statistical tests developed specifically to estimate selection in Ig sequences (Table 2).28, 29 In the bone marrow, selection could not be reliably analyzed owing to the small numbers of mutations.
Fetal mutations correlate with AID expression and are targeted to well-characterized AID hotspot motifs
In the fetal tissues analyzed (ileum, spleen, and bone marrow at 240 and 270 gd), AID expression as measured by RT-qPCR correlated significantly and positively with the overall mutational load (r=0.922, P=0.027, n=6) and specifically with the numbers of CDR mutations (r=0.946, P=0.016) and missense mutations (r=0.917, P=0.027). AID expression also correlated with silent mutations (r=0.845, P=0.034), which are not affected by selection on the protein level.
AID is known to preferentially target WRCY and its complementary RGYW motif in the genome.17, 30 In particular, Cs at the third position of WRCY and the corresponding Gs at the second position of RGYW are targeted. We searched for these hotspot motifs at the FR1–FR3 region and analyzed mutations within the motifs. As shown in Figure 4, the target G/C nucleotides in RGYW/WRCY motifs were preferentially mutated as compared with G/C nucleotides outside the motifs in both fetal and postnatal calf sequences.
Targeting of mutations to activation-induced cytidine deaminase (AID) hotspot motifs (RGYW/WRCY) in immunoglobulin heavy chain variable cDNA sequences from fetal and calf lymphoid tissues. Numbers of mutated AID target G/C nucleotides and numbers of mutated G/C nucleotides outside AID target motifs are shown per 1,000 target and non-motif G/C nucleotides analyzed. A total of 11,039 target G nucleotides, 10,931 target C nucleotides, 63,078 non-target G nucleotides, and 68,486 non-target C nucleotides were analyzed.
Discussion
In ruminants, the fetal IPP is characterized by SHM of Ig genes in the absence of an external antigen.6 To our knowledge, no molecular studies addressing the mechanism have been reported. We studied the role of AID in this process. AID is an obligatory factor in all known mechanisms that shape the antibody repertoire in response to antigenic challenge, e.g., SHM, GC, and class switch recombination.31, 32, 33
We report that AID is strongly expressed in the fetal bovine IPP and spleen (Figures 1 and 2). The mutational profile in IGHV cDNA sequences derived from these tissues is similar to the profile generated by AID-mediated SHM in secondary antibody response. Mutations are preferentially located in CDRs (Figure 3), generally resulting in an amino-acid change (Table 1), and are enriched in the AID hotspot motifs (Figure 4). By contrast, AID expression is low in the fetal bone marrow, correlating with small numbers of IGHV mutations. The data are highly suggestive of AID-dependent SHM in the fetal bovine IPP and spleen before exposure to exogenous antigens and imply a specific molecular mechanism to diversify the limited germline Ig repertoire.27, 34
Statistical analysis indicated strong selection of the mutated IGHV sequences in the fetal IPP and spleen (Table 2). Negative selection on the FRs can be explained by the adverse effects replacement mutations have on protein structure. The CDR sequences were positively selected, although the fetus is sterile and there is no appreciable external antigen present to account for the selective forces. In sequences derived from the fetal IPP and spleen, mutations were ∼5.6 times as frequent in CDRs as in FRs. The RGYW/WRCY preference of AID explains at most about 34% of this difference, as the frequency of these motifs in CDRs was ∼1.9 times that in FRs. The remainder can be attributed to selection, although other components of the mutation machinery may also contribute, such as the WA/TW mutator DNA polymerase η recruited by AID-induced U:G mismatches.17, 30, 35 The positive selection may be driven by unknown endogenous antigen(s). Even if not antigen driven, the products of unmutated IGHV genes may be unable to provide for proliferative signals as efficiently as those encoded by genes carrying favorable mutations in their CDR-encoding regions.
Unknown IGHV germline sequences potentially present in the analyzed animals or technical sequence errors did not likely affect our conclusions. To complement the limited information on the structure of the bovine Ig heavy chain locus, we recently characterized the bovine IGHV repertoire based on the available genome sequencing data.27 In addition, we now cloned and sequenced functional IGHV genes in individual animals analyzed in this work. To improve the accuracy of cDNA data sets, short or low quality cDNA sequences were excluded from the analyses. The combined error rate for PCR and sequencing, based on analysis of the vector-derived sequences, was 0.784 per kilobase or about 5% of the average observed mutational load. We also tested for the generation of chimeric IGHV sequences in the PCR reactions, using a 1:1 mixture of two known IGHV clones. We cloned and sequenced 109 PCR products out of which only one clone was chimeric, indicating that the polymerase used was highly processive.
The bovine genome contains a limited number of functional Ig genes, which are restricted to few families or subgroups.27, 34 Antibody-diversifying mechanisms in addition to recombination are therefore likely to exist. Previous studies have suggested that these may include SHM,6, 36 GC,7 and junctional diversity.37 Our study provides extensive molecular and mutational data suggesting that AID-associated SHM and selection are taking place in the IPP and spleen of bovine fetuses before exposure to external antigens. These processes are apparently spatially separated from de novo B lymphopoiesis and recombination, which occur in the fetal bovine bone marrow and lymph nodes,38, 39 as AID expression and/or the numbers of IGHV mutations are low in these tissues.
Using hematopoietic chimeras, we have previously shown that peripheral B lymphocytes in young cattle are generated from a self-sufficient pool of developing B cells independent of bone marrow hematopoietic stem cells.40 In these animals, the ratio of donor cells among circulating B lymphocytes stably reflected that in the IPP. These observations and our current data suggest two phases of B-cell generation: first, differentiation in the bone marrow and lymph nodes; second, migration to the IPP for additional diversification of the limited germline repertoire by SHM. In addition to IPP, SHM in cattle fetuses occurs in the spleen, and occasional AID-positive follicles were also observed in the fetal lymph nodes (Figure 2c,d). The proliferation rate of B cells is markedly higher in the IPP than in the other lymphoid tissues (Figure 2e–g),41 whereas the total number of lymphocytes in IPP is roughly comparable with the spleen and lymph nodes.42, 43 Thus, our observations are compatible with previous studies indicating that IPP produces the majority of the diversified peripheral B-cell pool.44
Methods
RNA extraction. Tissue samples from 15 bovine fetuses (190–290 gd) were collected from a local abattoir, snap frozen in liquid nitrogen, and conserved at −80 °C. Total RNA was extracted from the ileum, spleen, lymph node, bone marrow, and muscle using Eurozol (EuroClone, Milano, Italy) according to the manufacturer’s instructions. Briefly, 100–300 mg of frozen tissue was ground in a pre-cooled mortar, suspended in 1 ml Eurozol, and homogenized using Polytron PT1200 homogenizer (Kinematica AB, Lucerne, Switzerland). In all, 2.5 M lithium chloride was used to precipitate RNA followed by resuspension in RNase-free water. Remaining genomic DNA was removed with DNase treatment (Turbo DNase, Life Technologies, Carlsbad, CA) according to the manufacturer’s instructions. The RNA concentration was determined on Nanodrop spectrophotometer (Thermo Scientific, Asheville, NC) and the quality was assessed using RNA 6000 Nano kit on Agilent 2100 Bioanalyser (Agilent Technologies, Santa Clara, CA).
RT-qPCR. First-strand cDNA was synthesized from 1 μg of DNase-treated total RNA using RevertAid M-MuLV Reverse Transcriptase (Fermentas, St Leon-Rot, Germany). cDNAs were prepared as previously described.38 The cDNA mixture was diluted 1:2 in RNase-free water and 2 μl of this was used as a template in the qPCR reaction.
qPCR was performed using the SYBR green technology. The primers bAIDfw3, bAIDrev2 and QGAPDfw, QGAPDrev (Table 3) were designed on the basis of known sequences of bovine AID and bovine GAPDH (GeneBank accession number NM_001038682.1 (AID) and NM_001034034.1 (GAPDH)). Each reaction contained 1 × SYBR Premix Ex Taq mixture (TaKaRa, Otsu, Japan), 0.2 μM of forward and reverse primers and 1 × ROX II Reference Dye in a final volume of 20 μl. PCR conditions were adjusted with serial dilutions (107–101) of plasmid templates confirming that the PCR efficiency was satisfactory (90–110%). Templates used in these standard curves were cloned into pBluescript II SK (AID) vector (Fermentas) and pSTBlue-1 (GAPDH) vector (Life Technologies). The cDNA was transcribed from total RNA obtained from a young calf. The cloning primers were Aicda 5′ and Aicda 3′ yielding a 651 bp product (AID) and QGAPDfw and QGAPDrev yielding a 145 bp product (GAPDH; Table 3). Amplicons from the templates and cDNA samples were sequenced. Amplification was carried out using Stratagene’s Mx3005P real-time PCR system (Agilent Technologies). Cycling conditions were: 95 °C for 10 s, followed by 40 cycles of 95 °C for 5 s, 60 °C for 35 s and 80 °C for 15 s. The fluorescence signal was collected at 80 °C, with two replicates per each sample. The dissociation curves of the final products were analyzed with following conditions: 95 °C for 60 s, 55 °C for 30 s and 95 °C for 30 s, where the data were collected during all points from 55 °C to 95 °C.
Cycle threshold (Ct) values were determined with MxPro software based on a threshold line that was automatically defined above the non-informative data. To control the possible variation in the amount of cDNA in different samples, the level of gene expression of the target genes was normalized in relation to the reference gene GAPDH.
Immunostaining. AID immunohistochemistry was performed using the anti-human AID monoclonal antibody EK2-5G9 (Ascenion, Munich, Germany).26 The immunogenic peptide sequence of the anti-AID antibody is 100% identical to the bovine AID, and it produced an expected staining pattern in the adult lymph nodes used as positive controls. Paraffin sections of paraformaldehyde-fixed tissues were subjected to heat-induced antigen retrieval in 10 mM Tris-HCl pH 9.5, 1 mM EDTA pH 8.0, and blocked for endogenous peroxidase and biotin. The primary antibody was detected using a biotinylated goat anti-rat Ig secondary antibody (Jackson Immunoresearch, Suffolk, UK) and tyramide signal amplification as described previously.45
Double immunofluorescence for CD79α and the Ki67 antigen was performed using the mouse anti-human CD79α monoclonal antibody HM57 (Dako, Glostrup, Denmark) and the rabbit anti-human Ki67 antigen monoclonal antibody SP6 (Thermo Scientific). Antigen retrieval was performed as above. The primary antibodies were detected using an Alexa Fluor 488–conjugated goat anti-mouse Ig secondary antibody and an Alexa Fluor 546–conjugated goat anti-rabbit Ig secondary antibody (Life Technologies). The sections were counterstained with DAPI (4,6-diamidino-2-phenylindole) and the formaldehyde-induced autofluorescence was suppressed by staining in 0.1% Sudan Black B in 70% ethanol for 20 min.
The immunostained sections were observed and photographed using a Leica DM4000 microscope (Leica Microsystems, Wetzlar, Germany) equipped with an Olympus DP70 (Olympus, Tokyo, Japan) camera. CD79α single-positive and CD79α/Ki67 double-positive cells were counted using the ImageJ software (Bethesda, MD).46
Preparation and sequencing of fetal and calf IGHV cDNA libraries Samples of ileum were collected from fetuses of 240 and 270 gd and from a 51-days-old calf. Spleen and bone marrow samples were also collected from the fetuses. First-strand cDNA was synthesized using SuperScript III First-Strand Synthesis SuperMix (Life Technologies) according to the manufacturer’s instructions. cDNA was primed with target-specific primers (IgH rev1 for ileum of 240-days-old fetus) or with equal amounts of oligo(dT)20 and random primer mixes for the rest of the samples. Phusion High-Fidelity PCR Master Mix (Thermo Scientific) was used for PCR. Reaction contained 1 × Phusion Master Mix and 0.5 μM of forward and reverse primers (IgH fwd1 and IgH rev1 for samples from the ileum and IgH fwd2 and IgH rev2 for samples from the spleen and bone marrow, Table 3) in a final volume of 50 μl. Primers were designed to span the IGHV region from leader sequence to IGHμ region. In all, 2 μl of first-strand cDNA was used as the template. Cycling conditions consisted of an initial denaturation of 98 °C for 30 s, followed by 18 cycles of 98 °C for 10 s, 60 °C for 30 s, 72 °C for 20 s, and a final extension of 72 °C for 7 min. PCR products were then electrophoresed and purified. Approximately 100 ng of the purified PCR product was ligated into the pCR 4Blunt-TOPO vector and transformed into TOP10 Escherichia coli (Life Technologies). The TOP10 E. coli were grown overnight at 37 °C on LB-ampicillin (100 μg ml−1) plates. In all, 1 ml of LB-ampicillin medium was inoculated with a single bacterial clone.
For the 240 gd ileum, a total number of 384 single clones were isolated for sequencing. Purified DNA was amplified for 35 cycles using Dynazyme II polymerase (Thermo Scientific) with T7 and T3 primers (Oligomer, Helsinki, Finland). The PCR products were sequenced at the DNA sequencing and Genomics laboratory (Institute of Biotechnology, University of Helsinki, Finland).
For the rest of the samples, 48–144 single colonies were picked, purified, and sequenced. Plasmid purification and sequencing was conducted by GATC Biotech AG (Konstanz, Germany).
Preparation and sequencing of a post-immunisation IGHV cDNA library An inactivated whole-cell Streptococcus uberis vaccine was used for vaccination of a calf, which developed a strong antibody response. The draining lymph node of the vaccination site was removed at slaughter 9 months after vaccination, cut into small pieces, and stored in RNAlater (Life Technologies). Total RNA was extracted from the lymph node using Trizol (Life Technologies) according to the manufacturer’s instructions. RNA was DNase treated (DNase I Recombinant, Roche, Basel, Switzerland) and first-strand cDNA was synthesized using the iScript Select cDNA synthesis kit (Bio-Rad, Hercules, CA) with equivalent concentrations of the Oligo(dT)20 and random primer mixes. The bovine VH repertoire was amplified in PCR reactions containing 20 ng cDNA, 0.5 μM of each of the three reverse primers (Bov VH-R1–Bov VH-R3, Table 3) in combination with 0.5 μM of each of one of the eight forward primers (Bov VH-F1–Bov VH-F8), 0.2 μM each dNTP, 2.5 mM MgCl2, 0.05 U Platinum Taq DNA Polymerase (Life Technologies), and 1 × PCR buffer in a final volume of 20 μl. Cycling conditions were: 95 °C for 10 min, followed by 30 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 45 s followed by a final 10 min extension at 72 °C. The eight heavy chain reactions were combined and column purified using the DNA Clean and Concentrator kit (Zymo Research, Irvine, CA) before digestion with AscI and SpeI. The digested DNA was ligated into the similarly digested vector JSC (GenBank Accession EU109715). DH5α-E ElectroMax (Life Technologies) were transformed with the ligated DNA and plated on 2YT-amp plates (100 μg ml−1 ampicillin) containing 2% glucose. There were 4.3 × 1010 primary transformants of which 88% contained an insert. A total of 384 clones were isolated from the library for sequencing using primers specific to plasmid JSC (JSCmcsFwd and JSCmcsRev1, Table 3) and analyzed as above.
Cloning and sequencing of bovine germline IGHV genes. Muscle genomic DNA was extracted from three fetuses (aged 182, 240, and 270 gd) and a 51-days-old calf using GenElute Mammalian genomic DNA Miniprep kit (Sigma-Aldrich, St Louis, MO). Primers GenIGHV fwd1 and GenIGHVrev1 (Table 3) were used with Phusion High-Fidelity PCR Master Mix. Purified PCR product was then cloned into pCR 4 Blunt-TOPO vector as above. A total number of 384 colonies was picked, purified, and sequenced by GATC Biotech AG, yielding 35–70 sequences per animal. Sequences that were observed at least twice were accepted, to a total of 214 reads.
Analysis of mutations. The sequence data from the cDNA libraries were analyzed with Geneious Pro software version 5.6.6 (Biomatters, Auckland, New Zealand) and R.47 High-quality cDNA sequences spanning the FR1–FR3 region were matched to the bovine IGHV gene sequences and aligned on the best matching genes using the “map-to-reference” function of Geneious with high sensitivity. Replacement and silent mutations in the cDNA alignments and specifically in the AID-target motifs RGYW/WRCY17, 30 in the IGHV genes were detected in Geneious. RGYW and WRCY target motifs were detected separately in the forward strand. The numbers of mutations in FR and CDR regions, mutations of the AID target nucleotides (the second G in motif RGYW and the third C in motif WRCY), and mutations of G/C nucleotides outside the AID motifs were counted in R.47 The ratios of mutated FR/CDR/AID target/non-target nucleotides were then calculated as

where mi=the number of mutated FR/CDR/AID target/non-target nucleotides in the ith cDNA sequence, ni=the total number of FR/CDR/AID target/non-target nucleotides in the corresponding reference gene sequence, and k=the number of cDNA/reference gene sequence pairs analyzed from the tissue sample.
Statistical analysis. RT-qPCR data from different fetal tissues were compared in R using the nonparametric approximate Kruskal–Wallis one-way analysis of variance followed by multiple comparisons using the nonparametric Nemenyi–Damico–Wolfe–Dunn test at α=0.01.
Cell counts from immunofluorescence analyses were compared using the nonparametric Mann–Whitney U test in the IBM SPSS Statistics software version 20 (IBM Corporation, Armonk, NY).
For detecting selection, the BASELINe framework (Bayesian estimation of antigen-driven selection) was applied.28, 29 The tests were run at the web server at http://selection.med.yale.edu/baseline/ using focused selection statistics and CDR and FR boundaries as defined by Kabat et al.48
The Pearson correlations between AID expression data and the mutation counts were calculated in IBM SPSS Statistics version 20.
P values for cell counts and correlations were corrected for multiple testing using Holm’s method.49
Accession codes
References
Ratcliffe, M.J.H. B cell development in gut associated lymphoid tissues. Vet. Immunol. Immunopathol. 87, 337–340 (2002).
Weill, J.C. & Reynaud, C.A. Galt versus bone marrow models of B cell ontogeny. Dev. Comp. Immunol. 22, 379–385 (1998).
Lanning, D., Zhu, X., Zhai, S.K. & Knight, K.L. Development of the antibody repertoire in rabbit: gut-associated lymphoid tissue, microbes, and selection. Immunol. Rev. 175, 214–228 (2000).
Reynolds, J. The genesis, tutelage and exodus of B cells in the ileal Peyer’s patch of sheep. Int. Rev. Immunol. 15, 265–299 (1997).
Yasuda, M., Jenne, C.N., Kennedy, L.J. & Reynolds, J.D. The sheep and cattle Peyer’s patch as a site of B-cell development. Vet. Res. 37, 401–415 (2006).
Reynaud, C.A., Garcia, C., Hein, W.R. & Weill, J.C. Hypermutation generating the sheep immunoglobulin repertoire is an antigen-independent process. Cell 80, 115–125 (1995).
Parng, C.L., Hansal, S., Goldsby, R.A. & Osborne, B.A. Gene conversion contributes to Ig light chain diversity in cattle. J. Immunol. 157, 5478–5486 (1996).
Berens, S.J., Wylie, D.E. & Lopez, O.J. Use of a single V(H) family and long CDR3s in the variable region of cattle Ig heavy chains. Int. Immunol. 9, 189–199 (1997).
Verma, S., Goldammer, T. & Aitken, R. Cloning and expression of activation induced cytidine deaminase from Bos taurus. Vet. Immunol. Immunopathol. 134, 151–159 (2010).
Becker, R.S. & Knight, K.L. Somatic diversification of immunoglobulin heavy chain VDJ genes: evidence for somatic gene conversion in rabbits. Cell 63, 987–997 (1990).
Knight, K.L. & Becker, R.S. Molecular basis of the allelic inheritance of rabbit immunoglobulin VH allotypes: implications for the generation of antibody diversity. Cell 60, 963–970 (1990).
Reynaud, C.A., Anquez, V., Grimal, H. & Weill, J.C. A hyperconversion mechanism generates the chicken light chain preimmune repertoire. Cell 48, 379–388 (1987).
Thompson, C.B. & Neiman, P.E. Somatic diversification of the chicken immunoglobulin light chain gene is limited to the rearranged variable gene segment. Cell 48, 369–378 (1987).
Maul, R.W. et al. Uracil residues dependent on the deaminase AID in immunoglobulin gene variable and switch regions. Nat. Immunol. 12, 70–76 (2011).
Pham, P., Bransteitter, R., Petruska, J. & Goodman, M.F. Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation. Nature 424, 103–107 (2003).
Yu, K., Huang, F.T. & Lieber, M.R. DNA substrate length and surrounding sequence affect the activation-induced deaminase activity at cytidine. J. Biol. Chem. 279, 6496–6500 (2004).
Longo, N.S., Satorius, C.L., Plebani, A., Durandy, A. & Lipsky, P.E. Characterization of Ig gene somatic hypermutation in the absence of activation-induced cytidine deaminase. J. Immunol. 181, 1299–1306 (2008).
Wang, M., Rada, C. & Neuberger, M.S. Altering the spectrum of immunoglobulin V gene somatic hypermutation by modifying the active site of AID. J. Exp. Med. 207, 141–153 (2010).
Rada, C., Jarvis, J.M. & Milstein, C. AID-GFP chimeric protein increases hypermutation of Ig genes with no evidence of nuclear localization. Proc. Natl. Acad. Sci. USA 99, 7003–7008 (2002).
Wilson, T.M. et al. MSH2-MSH6 stimulates DNA polymerase eta, suggesting a role for A:T mutations in antibody genes. J. Exp. Med. 201, 637–645 (2005).
Di Noia, J.M. & Neuberger, M.S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 76, 1–22 (2007).
Neuberger, M.S. & Rada, C. Somatic hypermutation: activation-induced deaminase for C/G followed by polymerase eta for A/T. J. Exp. Med. 204, 7–10 (2007).
Peled, J.U. et al. The biochemistry of somatic hypermutation. Annu. Rev. Immunol. 26, 481–511 (2008).
Storck, S., Aoufouchi, S., Weill, J.C. & Reynaud, C.A. AID and partners: for better and (not) for worse. Curr. Opin. Immunol. 23, 337–344 (2011).
Cowell, L.G., Kim, H.J., Humaljoki, T., Berek, C. & Kepler, T.B. Enhanced evolvability in immunoglobulin V genes under somatic hypermutation. J. Mol. Evol. 49, 23–26 (1999).
Greiner, A. et al. Differential expression of activation-induced cytidine deaminase (AID) in nodular lymphocyte-predominant and classical Hodgkin lymphoma. J. Pathol. 205, 541–547 (2005).
Niku, M., Liljavirta, J., Durkin, K., Schroderus, E. & Iivanainen, A. The bovine genomic DNA sequence data reveal three IGHV subgroups, only one of which is functionally expressed. Dev. Comp. Immunol. 37, 457–461 (2012).
Uduman, M. et al. Detecting selection in immunoglobulin sequences. Nucleic Acids Res. 39, W499–W504 (2011).
Yaari, G., Uduman, M. & Kleinstein, S.H. Quantifying selection in high-throughput immunoglobulin sequencing data sets. Nucleic Acids Res. 40, e134 (2012).
Rogozin, I.B., Pavlov, Y.I., Bebenek, K., Matsuda, T. & Kunkel, T.A. Somatic mutation hotspots correlate with DNA polymerase eta error spectrum. Nat. Immunol. 2, 530–536 (2001).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Revy, P. et al. Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the hyper-IgM syndrome (HIGM2). Cell 102, 565–575 (2000).
Arakawa, H., Hauschild, J. & Buerstedde, J.M. Requirement of the activation-induced deaminase (AID) gene for immunoglobulin gene conversion. Science 295, 1301–1306 (2002).
Ekman, A., Niku, M., Liljavirta, J. & Iivanainen, A. Bos taurus genome sequence reveals the assortment of immunoglobulin and surrogate light chain genes in domestic cattle. BMC Immunol. 10, 22 (2009).
Zeng, X. et al. DNA polymerase eta is an A-T mutator in somatic hypermutation of immunoglobulin variable genes. Nat. Immunol. 2, 537–541 (2001).
Verma, S. & Aitken, R. Somatic hypermutation leads to diversification of the heavy chain immunoglobulin repertoire in cattle. Vet. Immunol. Immunopathol. 145, 14–22 (2012).
Koti, M., Kataeva, G. & Kaushik, A.K. Novel atypical nucleotide insertions specifically at VH-DH junction generate exceptionally long CDR3H in cattle antibodies. Mol. Immunol. 47, 2119–2128 (2010).
Ekman, A., Pessa-Morikawa, T., Liljavirta, J., Niku, M. & Iivanainen, A. B-cell development in bovine fetuses proceeds via a pre-B like cell in bone marrow and lymph nodes. Dev. Comp. Immunol. 34, 896–903 (2010).
Ekman, A., Ilves, M. & Iivanainen, A. B lymphopoiesis is characterized by pre-B cell marker gene expression in fetal cattle and declines in adults. Dev. Comp. Immunol. 37, 39–49 (2012).
Pessa-Morikawa, T., Niku, M. & Iivanainen, A. Persistent differences in the level of chimerism in B versus T cells of Freemartin cattle. Dev. Comp .Immunol. 28, 77–87 (2004).
Reynolds, J.D. Mitotic rate maturation in the Peyer’s patches of fetal sheep and in the bursa of Fabricius of the chick embryo. Eur. J. Immunol. 17, 503–507 (1987).
Trepel, F. Number and distribution of lymphocytes in man. A critical analysis. Klin. Wochenschr. 52, 511–515 (1974).
Reynolds, J.D. & Morris, B. The evolution and involution of Peyer’s patches in fetal and postnatal sheep. Eur. J. Immunol. 13, 627–635 (1983).
Reynaud, C.A., Mackay, C.R., Müller, R.G. & Weill, J.C. Somatic generation of diversity in a mammalian primary lymphoid organ: the sheep ileal Peyer’s patches. Cell 64, 995–1005 (1991).
Niku, M., Ekman, A., Pessa-Morikawa, T. & Iivanainen, A. Identification of major cell types in paraffin sections of bovine tissues. BMC Vet. Res. 2, 5 (2006).
Schneider, C.A., Rasband, W.S. & Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
R Development Core Team. R: A Language and Environment for Statistical Computing, R Foundation for Statistical Computing: Vienna, Austria, (2012) Available from: http://www.R-project.org.
Kabat, E.A., Wu, T.T., Perry, H.M., Gottesman, K.S. & Foeller, C. Sequences of Proteins of Immunological Interest 5th edn National Institutes of Health: Bethesda, MD, USA, (1991).
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979).
Acknowledgements
This study was supported by grants from the Academy of Finland (122540/2007 to A.I.), Ministry of Agriculture and Forestry (828/312/2009 to A.I.), Finnish Veterinary Foundation and Finnish Foundation of Veterinary Research (to J.L.), and AgResearch internal funding (ARC project A13579). J.L. and A.E. were students in the Viikki Graduate School in Molecular Biosciences. We thank Kirsi Lahti, Tuire Pankasalo, Santeri Suokas, and Charlotte Bouchet (AgResearch) for expert technical assistance, Bryce Buddle for providing the calf tissue, and Kirsti Sihto and Else Anttila for help in collecting the fetal material. We also thank Stewart Bisset for critically reviewing the manuscript and for helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Rights and permissions
About this article
Cite this article
Liljavirta, J., Ekman, A., Knight, J. et al. Activation-induced cytidine deaminase (AID) is strongly expressed in the fetal bovine ileal Peyer’s patch and spleen and is associated with expansion of the primary antibody repertoire in the absence of exogenous antigens. Mucosal Immunol 6, 942–949 (2013). https://doi.org/10.1038/mi.2012.132
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2012.132
This article is cited by
-
Immunogenetic factors driving formation of ultralong VH CDR3 in Bos taurus antibodies
Cellular & Molecular Immunology (2019)
-
The composition of the perinatal intestinal microbiota in cattle
Scientific Reports (2018)
-
Transcriptome analysis reveals regional and temporal differences in mucosal immune system development in the small intestine of neonatal calves
BMC Genomics (2016)