Abstract
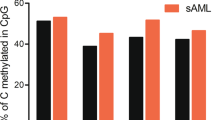
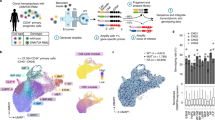
Cytogenetically normal myelodysplastic syndrome (CN-MDS) can pose diagnostic challenges and its pathogenetic mechanism remains elusive. By focusing on cytogenetically normal refractory cytopenia with multilineage dysplasia (CN-RCMD), a subtype of MDS, our genome-wide profiling showed ∼4600 annotated gene promoters with increased Histone H3 lysine 27 trimethylation (H3K27me3) in CN-RCMD, when compared with normal controls. Computational analysis revealed a statistically significant enrichment of the PU.1-binding DNA motif (PU-box) in the regions with increased H3K27me3. An inverse relationship between the levels of H3K27me3 and the levels of PU.1 binding and its downstream myeloid gene expressions was observed. Whole-exome sequencing analysis and Sanger sequencing analysis revealed some recurrent mutations, but no mutations in the PU.1 regulatory regions or in the EZH1/2, H3K27 methytransferase encoding genes. Using an MDS-derived erythroid/myeloid line and primary MDS bone marrow cells, we demonstrated that H3K27me3 inhibitors can increase the expression of PU.1 and its downstream genes and also promote cell differentiation via reducing H3K27me3 at the PU.1 gene locus. Finally, ectopic expression of PU.1 significantly inhibited proliferation of the MDS-derived cell line. Based on these data, we propose a hypothetical model of epigenetic inactivation of the PU.1 pathway due to increased H3K27me3 in some cases of CN-RCMD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Tefferi A, Vardiman JW . Myelodysplastic syndromes. N Engl J Med. 2009; 361: 1872–1885.
Nolte F, Hofmann WK . Myelodysplastic syndromes: molecular pathogenesis and genomic changes. Ann Hematol 2008; 87: 777–795.
Vardiman JW, Thiele J, Arber DA, Brunning RD, Borowitz MJ, Porwit A et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood 2009; 114: 937–951.
Haase D . Cytogenetic features in myelodysplastic syndromes. Ann Hematol 2008; 87: 515–526.
Orkin SH . Diversification of haematopoietic stem cells to specific lineages. Nat Rev Genet 2000; 1: 57–64.
Orkin SH, Zon LI . Hematopoiesis: an evolving paradigm for stem cell biology. Cell 2008; 132: 631–644.
Scott EW, Simon MC, Anastasi J, Singh H . Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science 1994; 265: 1573–1577.
Fisher RC, Scott EW . Role of PU.1 in hematopoiesis. Stem Cells 1998; 16: 25–37.
DeKoter RP, Walsh JC, Singh H . PU.1 regulates both cytokine-dependent proliferation and differentiation of granulocyte/macrophage progenitors. EMBO J 1998; 17: 4456–4468.
Hu CJ, Rao S, Ramirez-Bergeron DL, Garrett-Sinha LA, Gerondakis S, Clark MR et al. PU.1/Spi-B regulation of c-rel is essential for mature B cell survival. Immunity 2001; 15: 545–555.
Friedman AD . Transcriptional control of granulocyte and monocyte development. Oncogene 2007; 26: 6816–6828.
Spooner CJ, Cheng JX, Pujadas E, Laslo P, Singh H . A recurrent network involving the transcription factors PU.1 and Gfi1 orchestrates innate and adaptive immune cell fates. Immunity 2009; 31: 576–586.
Arinobu Y, Mizuno S, Chong Y, Shigematsu H, Iino T, Iwasaki H et al. Reciprocal activation of GATA-1 and PU.1 marks initial specification of hematopoietic stem cells into myeloerythroid and myelolymphoid lineages. Cell Stem Cell 2007; 1: 416–427.
Cao R, Zhang Y . The functions of E(Z)/EZH2-mediated methylation of lysine 27 in histone H3. Curr Opin Genet Dev 2004; 14: 155–164.
Shen X, Liu Y, Hsu YJ, Fujiwara Y, Kim J, Mao X et al. EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol Cell 2008; 32: 491–502.
Lee TI, Jenner RG, Boyer LA, Guenther MG, Levine SS, Kumar RM et al. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 2006; 125: 301–313.
Papayannopoulou T, Nakamoto B, Kurachi S, Tweeddale M, Messner H . Surface antigenic profile and globin phenotype of two new human erythroleukemia lines: characterization and interpretations. Blood 1988; 72: 1029–1038.
Lanotte M, Martin-Thouvenin V, Najman S, Balerini P, Valensi F, Berger R . NB4, a maturation inducible cell line with t(15;17) marker isolated from a human acute promyelocytic leukemia (M3). Blood 1991; 77: 1080–1086.
Ren B, Robert F, Wyrick JJ, Aparicio O, Jennings EG, Simon I et al. Genome-wide location and function of DNA binding proteins. Science 2000; 290: 2306–2309.
Pio F, Kodandapani R, Ni CZ, Shepard W, Klemsz M, McKercher SR et al. New insights on DNA recognition by ets proteins from the crystal structure of the PU.1 ETS domain-DNA complex. J Biol Chem 1996; 271: 23329–23337.
Kodandapani R, Pio F, Ni CZ, Piccialli G, Klemsz M, McKercher S et al. A new pattern for helix-turn-helix recognition revealed by the PU.1 ETS-domain-DNA complex. Nature 1996; 380: 456–460.
Li Y, Okuno Y, Zhang P, Radomska HS, Chen H, Iwasaki H et al. Regulation of the PU.1 gene by distal elements. Blood 2001; 98: 2958–2965.
Leddin M, Perrod C, Hoogenkamp M, Ghani S, Assi S, Heinz S et al. Two distinct auto-regulatory loops operate at the PU.1 locus in B cells and myeloid cells. Blood 2011; 117: 2827–2838.
Chen H, Zhang P, Radomska HS, Hetherington CJ, Zhang DE, Tenen DG . Octamer binding factors and their coactivator can activate the murine PU.1 (spi-1) promoter. J Biol Chem 1996; 271: 15743–15752.
Kotake Y, Cao R, Viatour P, Sage J, Zhang Y, Xiong Y . pRB family proteins are required for H3K27 trimethylation and Polycomb repression complexes binding to and silencing p16INK4alpha tumor suppressor gene. Genes Dev 2007; 21: 49–54.
Fiskus W, Wang Y, Sreekumar A, Buckley KM, Shi H, Jillella A et al. Combined epigenetic therapy with the histone methyltransferase EZH2 inhibitor 3-deazaneplanocin A and the histone deacetylase inhibitor panobinostat against human AML cells. Blood 2009; 114: 2733–2743.
Xu F, Li X, Wu L, Zhang Q, Yang R, Yang Y et al. Overexpression of the EZH2, RING1 and BMI1 genes is common in myelodysplastic syndromes: relation to adverse epigenetic alteration and poor prognostic scoring. Ann Hematol 2011; 90: 643–653.
Issa JP . Epigenetic changes in the myelodysplastic syndrome. Hematol Oncol Clin North Am 2010; 24: 317–330.
Tan J, Yang X, Zhuang L, Jiang X, Chen W, Lee PL et al. Pharmacologic disruption of Polycomb-repressive complex 2-mediated gene repression selectively induces apoptosis in cancer cells. Genes Dev 2007; 21: 1050–1063.
Tseng CK, Marquez VE, Fuller RW, Goldstein BM, Haines DR, McPherson H et al. Synthesis of 3-deazaneplanocin A, a powerful inhibitor of S-adenosylhomocysteine hydrolase with potent and selective in vitro and in vivo antiviral activities. J Med Chem 1989; 32: 1442–1446.
Fiskus W, Pranpat M, Balasis M, Herger B, Rao R, Chinnaiyan A et al. Histone deacetylase inhibitors deplete enhancer of zeste 2 and associated polycomb repressive complex 2 proteins in human acute leukemia cells. Mol Cancer Ther 2006; 5: 3096–3104.
Kubicek S, O'Sullivan RJ, August EM, Hickey ER, Zhang Q, Teodoro ML et al. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol Cell 2007; 25: 473–481.
Mueller BU, Pabst T, Fos J, Petkovic V, Fey MF, Asou N et al. ATRA resolves the differentiation block in t(15;17) acute myeloid leukemia by restoring PU.1 expression. Blood 2006; 107: 3330–3338.
Adolfsson J, Mansson R, Buza-Vidas N, Hultquist A, Liuba K, Jensen CT et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential a revised road map for adult blood lineage commitment. Cell 2005; 121: 295–306.
Goardon N, Marchi E, Atzberger A, Quek L, Schuh A, Soneji S et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 2011; 19: 138–152.
Cheng JX, Anastasi J, Vardiman JW . Analysis of DNA methylation and Histone H3 trimethylation at lysine 27 (H3K27me3) in myelodysplastic syndrome (MDS) with a novel sequential methylated DNA immunoprecipitation (SMeDIP) and sequential chromatin-immunoprecipitation (SChIP) technology: a pilot study of refractory cytopenia with multilineage dysplasia (RCMD). Modern Pathology 2008; 21: 249A (Abstract no. 1140).
Kondo Y, Shen L, Cheng AS, Ahmed S, Boumber Y, Charo C et al. Gene silencing in cancer by histone H3 lysine 27 trimethylation independent of promoter DNA methylation. Nat Genet 2008; 40: 741–750.
Nikoloski G, Langemeijer SM, Kuiper RP, Knops R, Massop M, Tonnissen ER et al. Somatic mutations of the histone methyltransferase gene EZH2 in myelodysplastic syndromes. Nat Genet 2010; 42: 665–667.
Ernst T, Chase AJ, Score J, Hidalgo-Curtis CE, Bryant C, Jones AV et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet 2010; 42: 722–726.
Bonadies N, Pabst T, Mueller BU . Heterozygous deletion of the PU.1 locus in human AML. Blood 2010; 115: 331–334.
Huang G, Zhang P, Hirai H, Elf S, Yan X, Chen Z et al. PU.1 is a major downstream target of AML1 (RUNX1) in adult mouse hematopoiesis. Nat Genet 2008; 40: 51–60.
Acknowledgements
We wish to thank Ms Ana Rodrigues in Bioinformatics for helping with computational analyses, and Ms Mandy Roteman, Ms Kelly Doston and Ms Christine A Feak for their assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Cheng, J., Anastasi, J., Watanabe, K. et al. Genome-wide profiling reveals epigenetic inactivation of the PU.1 pathway by histone H3 lysine 27 trimethylation in cytogenetically normal myelodysplastic syndrome. Leukemia 27, 1291–1300 (2013). https://doi.org/10.1038/leu.2013.45
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2013.45