Abstract
The importance of dysregulation of microRNA (miRNA) expression in nonalcoholic steatohepatitis (NASH) has been increasingly recognized; however, the association between altered expression of miRNAs and pathophysiological features of NASH and whether there is a connection between susceptibility to NASH and altered expression of miRNAs are largely unknown. In this study, male inbred C57BL/6J and DBA/2J mice were fed a lipogenic methyl-deficient diet that causes liver injury similar to human NASH, and the expression of miRNAs and the level of proteins targeted by these miRNAs in the livers were determined. Administration of the methyl-deficient diet triggered NASH-specific changes in the livers of C57BL/6J and DBA/2J mice, with the magnitude being more severe in DBA/2J mice. This was evidenced by a greater extent of expression of fibrosis-related genes in the livers of methyl-deficient DBA/2J mice. The development of NASH was accompanied by prominent changes in the expression of miRNAs, including miR-29c, miR-34a, miR-155, and miR-200b. Interestingly, changes in the expression of these miRNAs and protein levels of their targets, including Cebp-β, Socs 1, Zeb-1, and E-cadherin, in the livers of DBA/2J mice fed a methyl-deficient diet were more pronounced as compared with those in C57BL/6J mice. These results show that alterations in the expression of miRNAs are a prominent event during development of NASH induced by methyl deficiency and strongly suggest that severity of NASH and susceptibility to NASH may be determined by variations in miRNA expression response. More important, our data provide a mechanistic link between alterations in miRNA expression and pathophysiological and pathomorphological features of NASH.
Similar content being viewed by others
Main
Nonalcoholic fatty liver disease (NAFLD), a hepatic manifestation of the metabolic syndrome, and NASH, a progressive form of NAFLD, are two of the most common chronic liver diseases in the United States and worldwide.1, 2, 3 These two pathological states have been shown to progress to cirrhosis and hepatocellular carcinoma.3 The successful prevention and treatment of NASH relies on the ability to identify vulnerable sub-populations and detect the disease early, and greatly depends on a better understanding of the underlying molecular mechanisms involved in NASH pathogenesis. It is generally accepted that dysregulation of several physiological processes, including lipid metabolism, insulin resistance, immune response, inflamation, and oxidative stress, is associated with NASH pathogenesis.1, 2, 4 In addition, the results of recent studies have established a critical role of miRNAs in NASH.5, 6, 7, 8 Specifically, expression profiling studies have identified several differentially expressed miRNAs, including miR-21, miR-34a, miR-122, and miR-155, in human5 and mouse NASH,6, 7, 8 indicating their role in the pathogenesis of NASH. However, despite the current efforts and accomplishments in NASH experimental and clinical research and NASH clinical management, major gaps remain in understanding the etiology and progression of the disease at the basic level,2, 3 particularly, whether the different susceptibility to NASH is associated with an altered expression of miRNAs.
The differential susceptibility of mice to NASH has been attributed to strain variations in the expression of genes involved in hepatic lipid metabolism,9 triglyceride levels,10 oxidative stress,11 and c-Myc and NFκB signaling networks.8 Recently, we demonstrated that strain differences in the liver epigenomic status are associated with individual susceptibility to hepatic steatosis.12 Specifically, feeding DBA/2J mice a lipogenic methyl-deficient diet resulted in more prominent epigenetic and pathomorphological changes in the livers as compared with C57BL/6J mice.12
The present study was undertaken to (1) determine the association between altered expression of miRNAs and pathophysiological features of NASH in mice and (2) to determine whether strain-specific susceptibility of mice to NASH is associated with differences in miRNA expression.
MATERIALS AND METHODS
Animals, Diets, and Experimental Design
Male C57BL/6J and DBA/2J mice (Jackson Laboratory, Bar Harbor, ME, USA) were housed in sterilized cages in a temperature-controlled (24°C) room, with a 12 h light/dark cycle, and given ad libitum access to purified water and NIH-31-pelleted diet (Purina Mills, Richmond, IN, USA). At 8 weeks of age, the mice from each strain were allocated randomly into two groups, one control and one experimental. The mice from the experimental group were maintained on a low-methionine (0.18%) diet, lacking in choline and folic acid (Dyets, Inc, Bethlehem, PA, USA), for 12 weeks. The mice from the control group received diet supplemented with 0.4% methionine, 0.3% choline bitartrate, and 2 mg/kg folic acid. Diets were stored at 4°C and given ad libitum, with twice a week replacement. Five experimental and five control mice were killed at 6 and 12 weeks after diet initiation. Their livers were excised, frozen immediately in liquid nitrogen, and stored at −80°C for subsequent analyses. All animal experimental procedures were performed in accordance with the animal study protocol approved by the National Center for Toxicological Research Animal Care and Use Committee.
Necropsy, Tissue Processing, and Histopathology
The liver tissue samples were processed, analyzed, and scored for steatosis, hepatocellular degeneration, inflammation, hepatocellular karyocytomegaly, and oval cell proliferation as previously described.12
RNA Extraction and Microarray Expression Analysis
Total RNA was extracted from liver tissue using miRNAeasy Mini kits (Qiagen, Valencia, CA, USA) according to the manufacturer's instructions. The gene expression profile was determined using Agilent whole genome 4 × 44K mouse microarrays (Agilent Technologies, Santa Clara, CA, USA). Sample labeling and microarray processing were performed as detailed in the ‘One-Color Microarray-Based Gene Expression Analysis’ version 1.0 (Agilent Technologies) protocol. The miRNA microarray analysis was performed by LC Sciences (Houston, TX, USA), as reported previously in detail.13, 14
miRNA Expression Analysis by Quantitative Reverse Transcription Real-Time PCR (qRT-PCR)
Total RNA (200 ng) was used for qRT-PCRs of miR-29c, miR-34a, miR-122, miR-155, miR-192, miR-200b, miR-203, and miR-221 using TaqMan miRNA assays (Applied Biosystems, Foster City, CA, USA), according to the manufacturer's instructions. snoRNA202 was used as an endogenous control. The relative amount of each miRNA was measured using the 2−ΔΔCt method.15 All qRT-PCR reactions were conducted in triplicate and repeated twice.13, 14
Western Blot Analysis of Protein Expression
The levels of DNA methyltransferase 1 (Dnmt1), Dnmt3a, Dnmt3b, CCAAT enhancer-binding protein β (C/ebp-β), C/ebp-α, suppressor of cytokine signaling-1 (Socs1), E-cadherin (Cdh1), and β-actin proteins in the livers of control mice and mice fed the methyl-deficient diet were determined as described previously.12
Transfection of Mouse Primary Hepatocytes with pre-miR-155 and pre-miR-200b
Mouse (CD-1) primary hepatocytes (Invitrogen, Carlsbad, CA, USA) were transfected with 20 nM of either pre-miR-155 or pre-miR-200b (20 nM each; Applied Biosystems), in three independent replicates, using Lipofectamin 2000 transfection reagent (Invitrogen), according to the manufacturer's instructions. Primary hepatocytes transfected with scrambled RNA oligonucleotide served as a control. At 48 h after transfection, the medium was changed and transfection was repeated. At 48 h after the second transfection, adherent cells were harvested by mild trypsinization, washed in phosphate-buffered saline, and the viability of cells was determined. The cells were then immediately frozen at −80°C for subsequent analyses.
Isolation of Hepatic DNA
DNA was extracted by a slight modification of the method reported previously by Nakamura et al.16
Liquid Chromatography Combined with Electrospray Tandem Mass Spectrometry (LC-MS/MS) Analyses of 8-oxodeoxyguanosine (8-oxodG)
The measurement of 8-oxodG by LC-MS/MS was adapted from the method described by Powel et al.17 DNA samples (30–50 μg) were dissolved in 80 mM Tris-HCl buffer/20 mM MgCl2 (pH 7.0) containing 2.75 pmol [15N5]8-OH-dG as internal standard and enzymatically hydrolyzed to individual nucleosides by incubation with 40 U of DNase I, 2.7 mU of phosphodiesterase I, and 2 U of alkaline phosphatase. The released 8-oxodG was purified by reverse-phase high-performance liquid chromatography using a Beckman Ultrasphere ODS C18 column (5 μm, 4.6 × 250 mm, Beckman, Fullerton, CA, USA). The isocratic mobile phase was 7% MeOH in 10 mM ammonium formate (pH 4.3) with a flow rate of 1 ml/min. Fractions were collected at 2 min intervals before and after the elution of 8-oxodG. The quantitative analysis of 8-oxodG by LC-MS/MS was performed with a 1100 capillary high-performance liquid chromatograph (Agilent, Wilmington, DE, USA) coupled to a TSQ-Quantum triple quad mass analyzer (Thermo Finnigan, San Jose, CA, USA). A 3.5-μm Zorbax XDB-C18 column (0.3 × 150 mm; Agilent) was operated with a binary mobile phase of 2% 10 mM ammonium formate (pH 4.3) and 98% methanol, followed by a linear gradient increase in methanol from 2 to 30% from 0 to 5 min, holding at 30% for 10 min, and immediately returned to initial conditions that were held for 15 min. Both 8-oxodG and [15N5]8-OH-dG internal standard were detected by single reaction monitoring of the transition of nucleoside to base adduct at m/z 284.2–168.2 and m/z 289.2–173.2, respectively. MS conditions were as follows: spray voltage, 2200 V; and heated capillary temperature, 350°C. All reagents were purchased from Sigma-Aldrich (St Louis, MO, USA) and were of ACS grade or higher.
Determination of Genomic and Mitochondrial DNA Damage
The extent of genomic DNA damage was determined by measuring the levels of histone H2AX phosphorylation and histone H4 lysine 20 dimethylation by western immunoblotting.12, 18 The extent of mitochondrial DNA was determined by a quantitative PCR technique (qPCR).19, 20
Statistical Analyses
Results are presented as mean±s.e.m. Statistical analyses were conducted by two-way ANOVA, using diet and strain as fixed factors. Pairwise comparisons were conducted by the Student–Newman–Keuls test. P-values <0.05 were considered significant.
RESULTS
Expression of Fibrosis-relevant Genes in the Livers of C57BL/6J and DBA/2J Mice Fed a Methyl-deficient Diet
The results of our previous study showed that DBA/2J mice are more susceptible to NASH induced by methyl deficiency than are C57BL/6J mice.12 To further confirm that finding, we analyzed the histomorphological changes and the expression level of several critical fibrosis-associated genes,21 including tumor necrosis factor α (Tnf), procollagen (Col) genes, platelet-derived growth factor (Pdgf) C and D, platelet-derived growth factor receptor (Pdgfr) α, and matrix metallopeptidase (Mmp) 2 and 3, in the livers of C57BL/6J and DBA/2J mice fed the methyl-deficient diet for 12 weeks. Feeding a methyl-deficient diet resulted in more pronounced pathological changes in the livers of DBA/2J mice as compared with C57BL/6J mice (Supplementary Figure 1) and a greater upregulation of Col1a1, Col1a2, Col3a1, Col4a6, Col5a2, Col6a1, Col16a1, Pdgfc, Pdgfd, Pdgfrα, Mmp2, and Mmp3 genes in livers of methyl-deficient DBA/2J mice than in C57BL/6J mice (Supplementary Table 1).
Effects of Methyl-deficient Diet on the Extent of Genomic and Mitochondrial DNA Damage in the Livers of C57BL/6J and DBA/2J Mice
Another well-documented fundamental event in the development of NASH is mitochondrial dysfunction and induction of oxidative stress.2, 4, 22 In view of this, we studied the extent of genomic and mitochondrial DNA damage in the livers of C57BL/6J and DBA/2J mice fed a methyl-deficient diet. Figure 1a shows that, in livers of control mice, the level of 8-oxodG did not change over the 12-week period. Administration of the methyl-deficient diet to DBA/2J mice resulted in progressive accumulation of 8-oxodG in hepatic DNA, with a difference being significant after 6 and 12 weeks of deficiency (Figure 1a). In contrast, in the livers of C57BL/6J mice fed a methyl-deficient diet, the levels of 8-oxodG slightly increased after 12 weeks only.
Genomic and mitochondrial DNA damage in the livers of C57BL/6J and DBA/2J mice fed control and methyl-deficient diet for 12 weeks. (a) Levels of 8-oxodG in DNA in the livers of C57BL/6J and DBA/2J mice fed control or a methyl-deficient diet for 12 weeks. (b) Western blot analysis of histone H2AX phosphorylation. (c) Level of mitochondrial DNA damage as detected by qPCR. Data are presented as mean±s.d. (n=5). *Significantly different from age-matched control mice. **Significantly different from methyl-deficient C57BL/6J mice.
In addition to the increased levels of 8-oxodG in DNA, the livers from methyl-deficient C57BL/6J and DBA/2J mice were characterized by an increased level of histone H2AX phosphorylation (Figure 1b) and histone H4 lysine 20 dimethylation (data not shown), dependable markers for DNA damage. However, the magnitude of DNA damage in the livers of DBA/2J mice was more pronounced compared with that in the C57BL/6J strain, with the difference being significant after 12 weeks on a diet (Figure 1b). Feeding mice a methyl-deficient diet also resulted in mitochondrial DNA damage, with the extent being greater in DBA/2J (Figure 1c).
Dysregulation of miRNAs in the Livers of C57BL/6J and DBA/2J Mice Fed a Methyl-Deficient Diet
miRNA microarrays were used to analyze miRNA expression profiles in the livers of control C57BL/6J mice and C57BL/6J mice fed a methyl-deficient diet that causes a pathological state similar to human NASH.6, 23, 24 It is believed that this model is the best-established experimental model to study NASH.25 We identified 74 miRNAs (40 upregulated and 34 downregulated) that were differentially expressed (P<0.05) after 12 weeks of methyl deficiency (Supplementary Figure 2). The expression of the most altered miRNAs (miR-29c, miR-34a, miR-122, miR-155, miR-192, miR-200b, miR-203, and miR-221) in the livers of C57BL/6J mice fed the methyl-deficient diet was confirmed by qRT-PCR. In addition, we analyzed the expression of these miRNAs in the livers of control DBA/2J mice and DBA/2J mice fed the methyl-deficient diet. Interestingly, we detected an increased expression of miR-34a, miR-155, and miR-200b in the livers of both C57BL/6J and DBA/2J mice fed the methyl-deficient diet for 12 weeks, with the magnitude being significantly greater in DBA/2J mice (Table 1). Specifically, the expression of miR-34a, miR-155, and miR-200b in the livers of DBA/2J mice fed a methyl-deficient diet was 4.9, 5,9, and 2.9 times greater than that in C57BL/6J mice. In contrast, the livers of C57BL/6J mice fed a methyl-deficient diet were characterized by pronounced downregulation of miR-29c, whereas its expression in the livers of DBA/2J mice did not change.
Protein Level of miRNA Targets in the Livers of C57BL/6J and DBA/2J Mice Fed a Methyl-deficient Diet
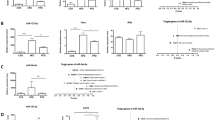
To establish the significance of miRNA dysregulation in the pathogenesis of NASH, specifically, to determine whether the aberrant expression of miRNAs is associated with pathophysiological features of NASH-induced liver injury, we determined protein levels of the targets of the most differentially expressed miRNAs. The results of recent studies have shown that the targets for miR-155 are C/ebp-β,6, 26 a liver-enriched transcription factor, and Socs1 tumor suppressor.27 Figure 2 shows that the increased expression of miR-155 in the livers of C57BL/6J and DBA/2J methyl-deficient mice (Figure 2a) was associated with the decreased levels of C/ebp-β and Socs1 proteins (Figure 2b). Furthermore, the increase in miR-155 and decrease in C/ebp-β and Socs1 were more pronounced in the livers of DBA/2J mice (Figure 2).
Expression changes of miR-155 and C/ebp-β and Socs1 proteins in the livers of C57BL/6J and DBA/2J mice fed control and methyl-deficient diet (MDD) for 12 weeks. (a) qRT-PCR analysis of miR-155 expression. The miRNA expression data are presented as average fold change normalized to snoRNA-202 in the livers of methyl-deficient mice compared with control mice. The level of miRNA expression was measured using the 2−ΔΔCt method.15 These results were reproduced in two independent experiments. (b) Western blot analysis of C/ebp-β and Socs1 proteins. Liver tissue lysates were separated by SDS-PAGE and subjected to western immunoblotting using specific antibodies against C/ebp-β and Socs1 proteins. Equal sample loading was confirmed by immunostaining against β-actin. These results were reproduced in two independent experiments. Representative western immunoblot images are shown. Chemiluminescence detection was performed with the Immobilon Western Chemiluminescent HRP Substrate (Millipore Corporation) and measured directly by a BioSpectrum Imaging System (UVP). Signal intensity was analyzed by ImageQuant software (Molecular Dynamics). Data are presented as mean±s.d. (n=5). *Significantly different from age-matched control mice. **Significantly different from methyl-deficient C57BL/6J mice.
The experimentally confirmed targets for miR-29 family are de novo DNA methyltransferases, namely, Dnmt3a and Dnmt3b.28 Figure 3 shows that downregulation of miR-29c (Figure 3a) was associated with increased protein levels of Dnmt3a and Dnmt3b in the livers of C57BL/6J mice fed the methyl-deficient diet (Figure 3b) as compared with age-matched control C57BL/6J mice. In contrast, the expression of miR-29 and the level of Dnmt3a and Dnmt3b proteins in the livers of DBA/2J mice fed the methyl-deficient diet did not change (Figure 3).
Expression changes of miR-29c and Dnmt1, Dnmt3a, and Dnmt3b proteins in the livers of C57BL/6J and DBA/2J mice fed control and methyl-deficient diet for 12 weeks. (a) qRT-PCR analysis of miR-29c expression. (b) Western blot analysis of Dnmt1, Dnmt3a, and Dnmt3b proteins. Data are presented as mean±s.d. (n=5). *Significantly different from age-matched control mice. **Significantly different from methyl-deficient C57BL/6J mice.
Feeding C57BL/6J and DBA/2J mice a methyl-deficient diet for 12 weeks resulted in substantial upregulation of miR-200b (Figure 4a) and downregulation of its Zeb1 target29 (Figure 4b), a well-known transcriptional repressor of E-cadherin, by 26 and 52%, respectively. The decreased Zeb1 levels in the livers of C57BL/6J and DBA/2J mice exposed to methyl deficiency were consequently accompanied by 2.2 and 3.3 times upregulation of E-cadherin (Cdh1) protein level, respectively, as compared with age-matched control mice (Figure 4b). Interestingly, a greater increase of E-cadherin protein in the livers of methyl-deficient DBA/2J mice corresponded to a greater expression of miR-200b. In contrast, the protein level of C/ebp-α, another predicted target for mouse miR-200b, did not change in the livers of C57BL/6J mice fed the methyl-deficient diet, and slightly decreased in DBA/2J mice (Figure 4b).
Expression changes of miR-200b and Cdh1 and C/ebp-α proteins in the livers of C57BL/6J and DBA/2J mice fed control and methyl-deficient diet for 12 weeks. (a) qRT-PCR analysis of miR-200b expression. (b) Western blot analysis of Cdh1 and C/ebp-α proteins. Data are presented as mean±s.d. (n=5). * Significantly different from age-matched control mice. **Significantly different from methyl-deficient C57BL/6J mice.
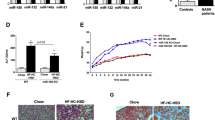
Ectopic Upregulation of miR-155 and miR-200b in Mouse Primary Hepatocytes
To confirm that altered levels of C/ebp-β, Socs1, and E-cadherin proteins in the livers of mice fed a methyl-deficient diet are associated with aberrant expression of miR-155 and miR-200b, we transfected mouse primary hepatocytes with these miRNAs. Figure 5 shows that ectopic upregulation of miR-155 in mouse primary hepatocytes was accompanied by a decreased level of C/ebp-β and Socs1 proteins, whereas the level of these proteins remained unchanged in hepatocytes transfected with miR-200b (Figure 5). Similarly, transfection of hepatocytes with miR-200b upregulated the level of E-cadherin (Cdh1) only.
Western blot analysis of C/ebp-β, Socs1, and Cdh1 proteins in mouse primary hepatocytes transfected with pre-mir-155 or pre-miR-200b. Mouse primary hepatocytes were transfected with 20 nM of either pre-miR-155 or pre-miR-200b. Mouse primary hepatocytes transfected with scrambled RNA oligonucleotide (mock) served as a control.
DISCUSSION
The current view on the pathogenesis of NASH favors a model in which NASH is associated with altered hepatic lipid metabolism, inflammation, and oxidative stress; however, the underlying mechanisms of these pathophysiological processes are not fully understood. The results of this study show that feeding C57BL/6J and DBA/2J mice a methyl-deficient diet, which is known to induce NASH in mice,6, 23, 24 results in aberrant expression of miRNAs, specifically miR-29c, miR-34a, miR-122, miR-155, miR-192, miR-200b, miR-203, and miR-221. Similar findings have been reported recently by Wang et al6 during NASH-induced hepatocarcinogenesis in mice fed a choline-deficient and amino acid-defined diet. In addition, this study shows the association of aberrant expression of miRNAs with fundamental pathophysiological features of NASH. These data indicate that dysregulation of miRNA expression is a critical event in the pathogenesis of NASH. More important, the results of our study show that strain-specific differences in susceptibility of mice to NASH are associated with differences in miRNA expression. Specifically, DBA/2J mice, a strain that develops more prominent NASH-specific changes, as evidenced by more severe pathomorphological changes12 (Supplementary Figure 1) and greater upregulation of fibrosis-related genes (Supplementary Table 1) compared with C57BL/6J mice, were characterized by a greater magnitude of miRNA expression changes in the livers.
The key pathophysiological features of NASH are an altered lipid metabolism and hepatocyte apoptosis.30, 31 The results of our study show a profound downregulation of miR-122, a liver-specific miRNA, which is important for normal lipid metabolism,32, 33 and dramatic upregulation of miR-34a, a critical regulator of apoptosis,34 in the livers of mice fed a methyl-deficient diet (Table 1). These changes correspond to more pronounced triglyceride accumulation and more severe cell death in DBA/2J mice fed a methyl-deficient diet as detected in our previous study12 and in this study. Several recent reports have shown that miRNAs miR-122 and miR-34a are two of the most frequently dysregulated miRNAs in NASH.5, 35 Specifically, increased expression of miR-34a and decreased expression of miR-122 have been demonstrated recently in the livers of ob/ob mice, a well-established hyperglycemic model of NAFLD, and in streptozotocin-induced type 1 diabetic mice.35 These are two main risk factors for the development of NAFLD and NASH in humans. Indeed, a similar dysregulation in expression of miR-34a and miR-122 has been reported in human NASH.5
The results of a recent study have shown the importance of miR-155 upregulation in NASH-induced hepatocarcinogenesis.6 miR-155 targets two critical tumor suppressors, C/ebp-β and Socs1,26, 27 which are frequently downregulated in hepatocellular carcinomas and hepatoblastomas.36, 37, 38 Moreover, in our previous study, we showed that inhibition of Socs1 expression is a critical event in hepatocarcinogenesis.39 In addition to the crucial role of Socs1 as a negative regulator of cytokine-induced signal transduction pathway,40 Socs1 functions as an important mediator of cellular oxidative stress.41 It is well known that several hepatic genes, including methionine adenosyltransferase 1a (Mat1a) and catalase, involved in regulation of oxidative stress and frequently downregulated in NASH as well,42, 43, 44 are regulated by C/ebp-β.45, 46 This corresponds to the results of our study that show the induction of oxidative stress (Figure 5), another key event in the pathogenesis of NASH,2, 4, 22 in the livers of mice fed the methyl-deficient diet.
In a previous study,12 we showed that feeding DBA/2J mice a lipogenic methyl-deficient diet also resulted in more prominent epigenetic changes in the livers of DBA/2J mice as compared with C57BL/6J mice. Specifically, administration of a methyl-deficient diet to C57BL/6J and DBA/2J mice resulted in a pronounced demethylation of genomic DNA and repetitive elements, with the magnitude being greater in DBA/2J mice. The results of this study indicate that the greater degree of demethylation of genomic DNA and, particularly, repetitive elements in the livers of methyl-deficient DBA2J mice compared with methyl-deficient C57BL/6J mice may be associated with the miR-29-driven differences in DNMTs expression (Figure 3). It has been shown that cooperation between Dnmt1 and de novo DNA methyltransferases, namely, Dnmt3a and Dnmt3b, is absolutely required for the maintenance of the methylation of repetitive elements and that Dnmt1 by itself is not capable of maintaining methylation of repetitive elements.47 Therefore, the decreased miR-29 expression and consequent upregulation of Dnmt3a and Dnmt3b may be a mechanism that protects the genome from excessive demethylation.
Differential expression of the miR-200 family, including miR-200b, miR-200c, and miR-429, in the livers of methyl-deficient C57BL/6J and DBA/2J mice is an important finding of the study. These miRNAs target transcriptional repressors of E-cadherin, ZEB1, and ZEB2,29, 48 resulting in an increased expression of E-cadherin. More important, increased E-cadherin expression starting at preneoplastic stages has been associated with a more aggressive hepatocarcinogenic process in mice.49
The results of our in vitro experiments confirmed the mechanistic link between altered expression of miR-155 and miR-200b and their targets. Similarly, in our previous study, we showed that transfection of rat hepatocytes with miR-34a induced apoptosis.50
In conclusion, these findings show that alterations in the expression of miRNAs are a prominent event in NASH induced by methyl deficiency in mice and strongly suggest that differences in the susceptibility to NASH and the severity of NASH may be determined by variations in miRNA expression response. More important, our data provide a mechanistic link between alterations in miRNA expression and pathophysiological and pathomorphological features of NASH. Additional studies will be necessary to determine whether altered expression of miRNAs, by itself, may induce the development of NASH and whether miRNA correction weakens NASH symptoms and decreases NASH progression.
References
Farrell GC, Larter CZ . Nonalcoholic fatty liver disease: from steatosis to cirrhosis. Hepatology 2006;43:S99–S112.
Marra F, Gastaldelli A, Svegliati Baroni G, et al. Molecular basis and mechanisms of progression of non-alcoholic steatohepatitis. Trends Mol Med 2008;14:72–81.
Siegel AB, Zhu AX . Metabolic syndrome and hepatocellular carcinoma: two growing epidemics with a potential link. Cancer 2009;115:5651–5661.
Matsuzawa N, Takamura T, Kurita S, et al. Lipid-induced oxidative stress causes steatohepatitis in mice fed an atherogenic diet. Hepatology 2007;46:1392–1403.
Cheung O, Puri P, Eicken C, et al. Nonalcoholic steatohepatitis is associated with altered hepatic microRNA expression. Hepatology 2008;48:1810–1820.
Wang B, Majumder S, Nuovo G, et al. Role of microRNA-155 at early stages of hepatocarcinogenesis induced by choline-deficient and amino-defined diet in C57BL/6 mice. Hepatology 2009;50:1152–1161.
Dolganiuc A, Petrasek J, Kodys K, et al. MicroRNA expression profile in Lieber-DeCarli diet-induced alcoholic and methionine choline deficient diet-induced nonalcoholic steatohepatitis models in mice. Alcohol Clin Exp Res 2009;33:1704–1710.
Hill-Baskin AE, Markiewski MM, Buchner DA, et al. Diet-induced hepatocellular carcinoma in genetically predisposed mice. Hum Mol Genet 2009;18:2975–2988.
Yamazaki Y, Kakizaki S, Takizawa D, et al. Interstrain differences in susceptibility to non-alcoholic steatohepatitis. J Gastroenterol Hepatol 2008;23:276–282.
Lin X, Yue P, Chen Z, et al. Hepatic triglyceride contents are genetically determined in mice: results of a strain survey. Am J Physiol Gastrointest Liver Physiol 2005;288:G1179–G1189.
Syn WK, Yang L, Chiang D, et al. Genetic differences in oxidative stress and inflammatory responses to diet-induced obesity do not alter liver fibrosis in mice. Liver Int 2009;29:1262–1272.
Pogribny IP, Tryndyak VP, Bagnyukova TV, et al. Hepatic epigenetic phenotype predermines individual susceptibility to hepatic steatosis in mice fed a lipogenic methyl-deficient diet. J Hepatol 2009;51:176–186.
Pogribny IP, Tryndyak VP, Boyko A, et al. Induction of microRNAome deregulation in rat liver by long-term tamoxifen exposure. Mutat Res 2007;619:30–37.
Tryndyak VP, Beland FA, Pogribny IP . E-cadherin transcriptional down-regulation by epigenetic and microRNA-200 family alterations is related to mesenchymal and drug-resistant phenotypes in human breast cancer cells. Int J Cancer 2010;126:2575–2583.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001;25:402–408.
Nakamura J, La DK, Swenberg JA . 5′-nicked apurinic/apyrimidinic sites are resistant to beta-elimination by beta-polymerase and are persistent in human cultured cells after oxidative stress. J Biol Chem 2000;275:5323–5328.
Powell CL, Kosyk O, Ross PK, et al. Phenotypic anchoring of acetaminophen-induced oxidative stress with gene expression profiles in rat liver. Toxicol Sci 2006;93:213–222.
Hanasoge S, Ljungman M . H2AX phosphorylation after UV irradiation is trigerred by DNA repair intermediates and is mediated by the ATR kinase. Carcinogenesis 2007;28:2298–2304.
Yakes FM, Van Houten B . Mitochondrial DNA damage is more extensive and persists longer than nuclear DNA damage in human cells following oxidative stress. Proc Natl Acad Sci USA 1997;94:514–519.
Milano J, Day BJ . A catalytic antioxidant metalloporphyrin blocs hydrogen peroxide-induced mitochondrial DNA damage. Nucl Acids Res 2000;28:968–973.
Strnad P, Tao GZ, Zhou Q, et al. Keratin mutation predisposes to mouse liver fibrosis and unmasks differential effects of the carbon tetrachloride and thioacetamide models. Gastroenterology 2008;134:1169–1179.
Tessari P, Coracina A, Cosma A, et al. Hepatic lipid metabolism and non-alcoholic fatty liver disease. Nutr Metab Cardiovasc Dis 2009;19:291–302.
Anstee QM, Goldin RD . Mouse models in non-alcoholic fatty liver and steatohepatitis research. Int J Exp Pathol 2006;87:1–16.
Velayudham A, Dolganiuc A, Ellis M, et al. VSL#3 probiotic treatment attenuates fibrosis without changes in steatohepatitis in a diet-induced nonalcoholic steatohepatitis model in mice. Hepatology 2009;49:989–997.
Ariz U, Mato JM, Lu SC, et al. Nonalcoholic steatohepatitis, animal models, and biomarkers, what is new? Methods Mol Biol 2010;593:109–136.
Worm J, Stenvang J, Petri A, et al. Silencing of microRNA-155 in mice during acute inflammatory response leads to derepression of c/ebp Beta and down-regulation of G-CSF. Nucleic Acids Res 2009;37:5784–5792.
Iorio MV, Ferracin M, Liu CG, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 2005;65:7065–7070.
Fabbri M, Garzon R, Cimmino A, et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA 2007;104:15805–15810.
Hurteau GJ, Carlson JA, Spivack SD, et al. Overexpression of the microRNA has-miR-200c leads to reduced expression of transcription factor 8 and increased expression of E-cadherin. Cancer Res 2007;67:7972–7976.
Jou J, Choi SS, Diehl AM . Mechanisms of disease progression in nonalcoholic fatty liver disease. Semin Liver Dis 2008;28:370–379.
Malhi H, Gores GJ . Molecular mechanisms of lipotoxicity in nonalcoholic fatty liver disease. Semin Liver Dis 2008;28:360–369.
Lynn FC . Meta-regulation: microRNA regulation of glucose and lipid metabolism. Trends Endocrinol Metab 2009;20:452–459.
Girard M, Jacquemin E, Munnich A, et al. miR-122, a paradigm for the role of microRNAs in the liver. J Hepatol 2008;48:648–656.
Raver-Shapira N, Marciano E, Meiri E, et al. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol Cell 2007;26:731–743.
Li S, Chen X, Zhang H, et al. Diferential expression of microRNAs in mouse liver under aberrant energy metabolic status. J Lipid Res 2009;50:1756–1765.
Tomizawa M, Watanabe K, Saisho H, et al. Down-regulated expression of the CCAAT/enhancer binding protein alpha and beta genes in human hepatocellular carcinoma: a possible prognostic marker. Anticancer Res 2003;23:351–354.
Tomizawa M, Horie H, Yamamoto H, et al. Reciprocal expression of CCAAT/enhancer binding proteins alpha and beta in hepatoblastomas and its prognostic significance. Oncol Rep 2007;17:341–344.
Yoshikawa H, Matsubara K, Qian GS, et al. SOCS-1, a negative regulator of the JAK/STAT pathway. Is silenced by methylation in human hepatocellular carcinoma and shows growth-suppression activity. Nat Genet 2001;28:29–35.
Bagnyukova TV, Tryndyak VP, Muskhelishvili L, et al. Epigenetic downregulation of the suppressor of cytokine signaling 1 (Socs1) gene is associated with the STAT3 activation and development of hepatocellular carcinoma induced by methyl-deficiency in rats. Cell Cycle 2008;7:3202–3210.
Starr R, Willson TA, Viney EM, et al. A family of cytokine-inducible inhibitors of signaling. Nature 1997;387:917–921.
Oh J, Hur MW, Lee CE . SOCS1 protects protein tyrosine phosphatases upregulation and attenuates Jaks to suppress ROS-mediated apoptosis. Oncogene 2009;28:3145–3156.
Matínez-Chantar ML, Corrales FJ, et al. Spontaneous oxidative stress and liver tumors in mice lacking methionine adenosyltransferase 1A. FASEB J 2002;16:1291–1294.
Sreekumar R, Rosado B, Rasmussen D, et al. Hepatic gene expression in histologically progressive nonalcoholic steatohepatitis. Hepatology 2003;38:244–251.
Rubio A, Guruceaga E, Vázquez-Chantada M, et al. Identification of gene-pathway associated with non-alcoholic steatohepatitis. J Hepatol 2007;46:708–718.
Taniguchi M, Hashimoto M, Hori N, et al. CCAAT/enhancer binding protein-beta (C/EBP-beta), a pivotal regulator of the TATA-less promoter in the rat catalase gene. FEBS Lett 2005;579:5785–5790.
Ikeda R, Nishida T, Watanabe F, et al. Involvement of CCAAT/enhancer binding protein-beta (CEBPbeta) in epigenetic regulation of mouse methionine adenosyltransferase 1A gene expression. Int J Biochem Cell Biol 2008;40:1956–1969.
Liang G, Chan MF, Tomigahara Y, et al. Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol Cell Biol 2002;22:480–491.
Park SM, Gaur AB, Lengyel E, et al. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 2008;22:894–907.
Calvisi DF, Ladu S, Conner EA, et al. Disregulation of E-cadherin in transgenic mouse models of liver cancer. Lab Invest 2004;84:1137–1147.
Tryndyak VP, Ross SA, Beland FA, et al. Down-regulation of the microRNAs mir-34a, miR-127, and miR-200b in rat liver during hepatocarcinogenesis induced by a methyl-deficient diet. Mol Carcinogenesis 2009;48:479–487.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
DISCLAIMER
The views expressed in this paper do not necessarily represent those of the U.S. Food and Drug Administration.
Supplementary Information accompanies the paper on the Laboratory Investigation website
Rights and permissions
About this article
Cite this article
Pogribny, I., Starlard-Davenport, A., Tryndyak, V. et al. Difference in expression of hepatic microRNAs miR-29c, miR-34a, miR-155, and miR-200b is associated with strain-specific susceptibility to dietary nonalcoholic steatohepatitis in mice. Lab Invest 90, 1437–1446 (2010). https://doi.org/10.1038/labinvest.2010.113
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2010.113
Keywords
This article is cited by
-
Astaxanthin attenuates hepatic steatosis in high-fat diet-fed rats by suppressing microRNA-21 via transactivation of nuclear factor erythroid 2-related factor 2
Journal of Physiology and Biochemistry (2022)
-
LncRNA Gm12664–001 ameliorates nonalcoholic fatty liver through modulating miR-295-5p and CAV1 expression
Nutrition & Metabolism (2020)
-
Genetic Difference of Hypothyroidism-Induced Cognitive Dysfunction in C57BL/6j and 129/Sv Mice
Neurochemical Research (2019)
-
Association of plasma microRNA expression with age, genetic background and functional traits in dairy cattle
Scientific Reports (2018)
-
The roles of microRNA families in hepatic fibrosis
Cell & Bioscience (2017)