Abstract
Der is an essential and widely conserved GTPase that assists assembly of a large ribosomal subunit in bacteria. Der associates specifically with the 50S subunit in a GTP-dependent manner and the cells depleted of Der accumulate the structurally unstable 50S subunit, which dissociates into an aberrant subunit at a lower Mg2+ concentration. As Der is an essential and ubiquitous protein in bacteria, it may prove to be an ideal cellular target against which new antibiotics can be developed. In the present study, we describe our attempts to identify novel antibiotics specifically targeting Der GTPase. We performed the structure-based design of Der inhibitors using the X-ray crystal structure of Thermotoga maritima Der (TmDer). Virtual screening of commercially available chemical library retrieved 257 small molecules that potentially inhibit Der GTPase activity. These 257 chemicals were tested for their in vitro effects on TmDer GTPase and in vivo antibacterial activities. We identified three structurally diverse compounds, SBI-34462, -34566 and -34612, that are both biologically active against bacterial cells and putative enzymatic inhibitors of Der GTPase homologs. We also presented the possible interactions of each compound with the Der GTP-binding site to understand the mechanism of inhibition. Therefore, our lead compounds inhibiting Der GTPase provide scaffolds for the development of novel antibiotics against antibiotic-resistant pathogenic bacteria.
Similar content being viewed by others
Introduction
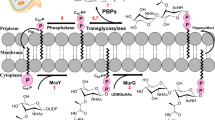
The emergence of antibiotics-resistant bacterial strains is a major threat to the clinical treatment of pathogenic bacterial infections, which limits the use of currently available antibiotics. Vancomycin-resistant pathogenic bacteria are now quite widely found.1 As vancomycin is considered to be the last resort in the present treatment of drug-resistant bacterial infections, development of new potent antibiotics has become extremely demanding. Antibiotics such as penicillin target bacterial cell wall biosynthesis, and some antibiotics, such as kanamycin, streptomycin and tetracycline, inhibit protein synthesis. Cellular processes such as RNA and DNA synthesis are interrupted by rifampicin2 and novomycin,3 respectively. These currently available conventional antibiotics target various macromolecular biosynthetic systems in the bacteria described above, as they are unique in bacterial cells or distinct from those in eukaryotic cells. While the screening of new antibiotics against macromolecular biosynthesis targets is ongoing, the alarming pace of the development of drug resistance has created the urgent need for the search for non-conventional antibacterial targets.
In the search for novel antibacterial targets, the GTPase superfamily offers an attractive set of targets for intervention into various critical functions of bacterial cells. GTPases have critical roles in regulation of cellular functions such as signal transduction, translation, translocation of polypeptides and cell division by modulating conformational change between GTP- and GDP-bound forms.4, 5, 6 Even though there is low primary sequence similarity among different GTPase families, most GTPases have an impressively common structural topology consisting of six β-strands surrounded by five α-helices.4, 5, 7
Previously we characterized an essential GTPase, Der (double Era-like protein), from Escherichia coli (EcDer)8, 9, 10, 11, 12 and from a hyperthermophilic bacterium Thermotoga maritima (TmDer).8, 13 The 3D structure of TmDer was solved by X-ray crystallography.13 Der homologs among other eubacteria, including E. coli, show highly conserved sequence homology from the N-terminus to the C-terminus.8 Der has a C-terminal KH-like (RNA-binding module-like) domain flanked by two tandemly repeated GTP-binding domains at the N-terminus.13, 14 In case of T. maritima Der, each of the two Der GTP-binding domains expressed separately has an intrinsic GTPase activity, though the activity of the N-terminal GD1 domain is stronger than that of the GD2 domain.13 It has been demonstrated that Der has a critical role in 50S subunit biogenesis, as it associates with 50S subunit in the presence of a GTP analog9, 15 and its depletion caused conformational instability of 50S subunit at a low Mg2+ concentration.9 GTP binds to Der in a cooperative manner and the interruption of cooperative nucleotide association disrupts the interaction of Der with the 50S subunit.11, 12, 15, 16 The essential role of Der in 50S subunit assembly was further supported by the isolation of Der as a suppressor for the rrmJ null phenotype.17 RrmJ is a 23S ribosomal RNA methyltransferase and its deletion resulted in reduction of the functional 70S ribosomes and accumulation of 30S and 50S subunits. The accumulated 50S subunits in rrmJ deletion cells were also disintegrated into abnormal subunits at a lower Mg2+ concentration similar to that observed in the Der-depleted cells.9, 11, 17 Furthermore, the primary sequence of Der is highly ubiquitous in most eubacteria. Importantly, this GTPase subfamily does not exist in eukaryotes. Therefore, the properties of Der GTPase that make it an ideal cellular target against which new antibiotics can be developed are as follows: (i) it is uniquely specific for bacteria, so that antibiotics developed against it will not be harmful to the human host; (ii) it is highly conserved in bacteria, so that antibiotics will be effective against a wide variety of pathogens; and (iii) it has an essential function in bacteria, so that antibiotics targeting its cellular activity will cause severe growth inhibition or accelerated death of bacterial cells.
In this study, we applied computer-aided pharmacophore modeling to retrieve putative small-molecule inhibitors of the bacterial Der GTPase, based on the crystal structure of T. maritima Der. By screening the computationally retrieved chemical library, we identified three chemicals, SBI-34462, -34566 and -34612, that are both bioactive to cells and inhibitory to the GTPase activity. Thus, our work potentially accelerates the discovery of a novel class of therapeutic agents.
Materials and methods
Molecular modeling of Der structure and virtual screening for lead compounds
The structure of Der with the GD1 domain in the empty state was taken from Protein Data Bank co-ordinates (PDB ID: 1MKY).13 The 3D structure of Der has been regularized with the ECEPP/3 force field18 in order to adjust X-ray co-ordinates to standard polypeptide geometry. GD1 was superimposed with auxiliary templates of active and inactive states; that is, crystal structures of Ras in complex with GTP (PDB ID: 121P)19 and GDP (PDB ID: 4Q21),20 respectively, and the ligand molecules were transferred from auxiliary templates into GD1 (Figure 1). We used a procedure of Monte Carlo simulations with flexible ligand docking protocols developed at Sapient Discovery, LLC.21 Monte Carlo search with minimization was used to optimize the positions and conformations of the ligand in the binding site, positions of Mg2+ ion and water molecules involved in the ligand binding and conformations of Der side chains located in a 5.0–7.0Å shell around the ligand. The GD1 switch I loop residues 24–37 that are absent in the X-ray structure were built into preliminary models of complexes. Conformations of GD1 switch I loop were optimized by Sapient Discovery, LLC's proprietary loop optimization algorithm22 and by the loop search procedure available in the ICM software package (San Diego, CA, USA).23 The crystal structure was used as an initial model of Der in the empty state (no ligand bound to GD1). Der 3D structures with GTP and GDP bound to the GD1 domain were utilized as initial models of the active and inactive states, respectively. Independent molecular dynamics simulations were performed starting from the three initial structures. The AMBER software package24 was used for molecular dynamics simulations. Initial models were solvated, that is, embedded in a box of water molecules and energy minimized with solvent. An equilibration period of 40–50 ps was followed by an acquisition period of 50 ps, during which the dynamic trajectories were collected and molecular snapshots of the system were saved every 10 ps (Figure 2). The force field and software used in this study have been described in an earlier work.25 Conformations of the ligand visited during dynamic simulations were used to derive DynaPharm templates. As the potential inhibitor of the Der GTPase activity is supposed to block the GTP-binding site while fixing an inactive state of the protein, a dynamic model of the inactive GD1–GDP complex was used as the primary target. The dynamic model of the active GD1–GTP complex was considered as an alternative target that might result in discovery of both inhibitors and activators of Der GTPase.
Structural overlap of the GD1 domain of Der (PDB ID: 1MKY, yellow) and the Ras-GTP complex (PDB ID: 121P, blue). GTP is shown in a space-filling models colored by atom type; Mg2+ is shown as a green ball. Ras switch I and II loops are colored red. The terminal residues of undefined GD1 switch I loop are labeled in green.
Der GD1 domain models with open and closed conformations of the Switch I loop. The model of GD1–GTP complex is displayed as a white ribbon with the Switch I loop colored blue and the GTP molecule colored by atom type. The model of GD1–GDP complex is displayed as a yellow ribbon with the loop colored red and the GDP molecule colored magenta.
The first set of lead chemicals based on the DynaPharm templates was generated by geometric matching of the queries against our database of commercially available compounds containing more than five million molecules. The pharmacophore template was translated into a three-dimensional query. The distances and angles, as well as their tolerances, were derived from the molecular dynamics trajectories. A 3D search based on these structural queries results in compounds that satisfy the distance and angle criteria contained in the query. The first list of hits was derived using the geometric criteria only. These compounds were further filtered based on calculated drug-likeness multilevel chemical compatibility (MLCC) score, logPo/w and solubility. Highly unstable, toxic, mutagenic or carcinogenic compounds were eliminated.
Purification of GTPase protein
For the expression of TmDer and EcDer, pETTmDer8, 13 or pETEcDer12 was introduced into BL21(DE3). The expression and purification of TmDer was carried out as described previously.8 EcDer was overexpressed and purified with a similar protocol except for a heat treatment. The open reading frames of der in Deinococcus radiodurans and Staphylococcus aureus were cloned in NdeI–EcoRI sites of pET28a (N-terminal His•tag fusion, Novagen, Gibbstown, NJ, USA), yielding pET28DrDer and pET28SaDer. pET28DrDer, pET28SaDer or pET28Era12 was transformed in BL21(DE3). The His-tagged proteins were overexpressed and purified using Ni-NTA column as described in the manufacturer's protocol of QIAGEN (Valencia, CA, USA).
In vitro GTPase assay
To quantitate the GTPase activty of Der in the presence of compound, [γ-32P]GTP was used for the GTPase reaction. The reaction was carried out in a 50-μl reaction mixture containing 50 mM Tris-HCl (pH 7.5), 400 mM KCl, 5 mM MgCl2, 1 mM DTT, 10 μM [γ-32P]GTP, 100 μM of compound and 5 μM of purified TmDer (or other GTPases) protein at 70 °C (or 37 °C for other GTPases) for 20 min. The reaction was terminated by the addition of 200 μl of 6% active charcoal and the measurement of the free inorganic phosphate with a liquid scintillation counter was carried out as described previously.8 All 257 compounds were dissolved in 100% dimethyl sulfoxide and tested for assays in duplicate reactions.
Bacterial strains and in vivo growth inhibition
Two bacterial strains of E. coli (W3110, Gram-negative) and S. aureus (Gram-positive) were used to test the inhibitory effects of compounds on cell growth. Overnight cell cultures of E. coli or S. aureus grown in LB medium were evenly spread on LB plates. Autoclaved filter paper disks (∅, 8 mm) were placed on each plate. Five microliters of each solution (100 mM) was spotted on the filter paper disk. Plates were incubated at 37 °C overnight. As control, 5 μl of 100% dimethyl sulfoxide (negative control), 50 μg ml−1 ampicillin (positive control) or 50 μg ml−1 kanamycin (positive control) was spotted on the filter paper disks. The antibacterial effect of the compounds was visualized by a halo formation around the filter paper disks. The MICs were determined by spotting approximately 5 × 103 bacterial cells to mini plates containing various concentrations of each compound.
Results and Discussion
The effects of compounds on TmDer GTPase activity
The virtual screening described in Materials and Methods yielded 257 compounds as potential candidates for TmDer GTPase inhibitors. First, in order to determine whether any of these compounds can act as antibiotics, they were examined for their abilities to suppress GTPase activity of TmDer. Each of 257 computationally screened compounds was assayed for its effect on TmDer GTPase. The GTPase assays were carried out using purified TmDer in the presence of 10 μM [γ-32P]GTP and 100 μM of compound at 70 °C for 20 min. Note that the optimal enzymatic reaction of TmDer occurred at 70 °C, as we previously demonstrated.8 The release of the inorganic phosphate of reaction was measured as described in Materials and Methods and compared with that of the control reaction without any compound. Of a total of 257 compounds, only 2 compounds (SBI-34527 and -34495) were identified that decreased TmDer GTPase activity by approximately 20–30% compared with the control reaction (Tables 1 and 2). As mentioned earlier, T. maritima is a hyperthermophilic bacterium, whose optimal growth is approximately 70 °C. Thus, it is very tempting to speculate that high temperature may reduce the affinity of compounds to TmDer.
In the following experiments, we decided to test the compounds for their effect on Der homologs and Era GTPase for the following reasons: (i) most pathogenic bacteria are mesophilic, optimally growing at the host's body temperature, under which compounds may behave differently and (ii) whether or not each compound has the same pattern against all Der homologs or Era GTPase.
In order to obtain candidate compounds that can act as inhibitors for the GTPases described above, first, they were examined for their abilities to inhibit bacterial growth. The growth-inhibition effect of compounds was examined by spotting the solution of each compound on paper disks that were placed on LB plates spread with E. coli (Gram-negative) or S. aureus (Gram-positive) cells. Among 257 compounds tested, a total of 12 different molecules (Tables 1 and 2) formed a halo around the respective paper disks, suggesting an antibacterial activity.
The effects of compounds on Der homologs and Era GTPase
Thus, using these 12 compounds, which showed antibacterial activity as described above, we further assessed the effect of these selected compounds on E. coli, D. radiodurans and S. aureus Der (EcDer, DrDer, and SaDer) and E. coli Era GTPases, and compared with the results obtained with TmDer. The enzymatic assays were carried out as for TmDer except that N-terminal His-tagged proteins were used for DrDer, SaDer and Era and that the reactions were carried out at 37 °C (Tables 1 and 2).
Interestingly, SBI-34462, -34527 and -34566 were generally inhibitory to EcDer, DrDer, SaDer and Era GTPases within a range of 30–90% activity reduction. The GTPase activities of DrDer, SaDer and Era were more specifically inhibited by SBI-34437 and -34612 (Table 1). SBI-34438 and -34439 substantially activated TmDer and EcDer GTPases, while the GTPase activities of DrDer, SaDer and Era were marginally inhibited by those compounds. The remaining five compounds, SBI-34495, -34522, -34559, -34565 and -34703, did not show noticeable inhibition effects for those GTPases (Table 2). Thus, our results indicate that SBI-34462, -34566, -34612, -34437 and -34527 appear to be potent inhibitors for the subfamily of Der GTPase.
In vivo antibacterial activities of compounds
Twelve compounds isolated above (Tables 1 and 2) were further tested to determine the MICs for their antibacterial inhibitions. MIC was determined by inoculating a diluted culture to mini plates containing various concentrations of compounds as described in Materials and methods. The MICs of SBI-34438, -34439, -34462, -34495, -34522, -34559, -34565, -34566, -34612 and -34703 for Gram-positive bacterium S. aureus were lower than those of SBI-34437 and -34527. Notably, the MICs of SBI-34439, -34462, -34495, -34559, -34565, -34566, -34612 and -34703 were in the range of >25–50 mg l−1, suggesting a stronger antibacterial activity than SBI-34438 and -34522. More interestingly, only SBI-34703 was highly active against both E. coli and S. aureus, with an MIC of >30–40 mg l−1 (Table 2). For those 10 compounds with an MIC of <200 mg l−1, cell growth was monitored in a liquid culture in the presence of compounds in order to investigate their mode of antibacterial action. Ten compounds were examined for their bioactivities against S. aureus (Figures 3a–c). For controls, dimethyl sulfoxide, ampicillin or chloramphenicol was added to the culture of S. aureus; addition of chloramphenicol to the culture reduced the growth rate gradually. Notably, ampicillin seemed to decrease the culture density after its addition, while in a longer incubation ampicillin-resistant S. aureus cells resumed to grow.26, 27 SBI-34462, -34566, -34612 and -34703 were the most effective growth inhibitors without rendering S. aureus resistant to them (Figures 3a and c). SBI-34565 was as inhibitory as chloramphenicol (Figure 3c). The remaining five compounds showed marginal or minimum inhibitory activities (Figures 3b and c). For E. coli, SBI-34703 was tested for its inhibitory effect on cell growth. The addition of SBI-34703 to the exponentially growing E. coli cells induced dramatic reduction of culture turbidity compared with the control culture added with dimethyl sulfoxide. In E. coli culture, ampicillin or chloramphenicol induced cell lysis or growth arrest, respectively, immediately after its addition (Figure 3d).
The in vivo bioactivity of compounds on E. coli and S. aureus. Growth curves of S. aureus cells (a–c) and E. coli (W3110) (d) in the presence and in the absence of compounds. Overnight cultures of each bacterium were diluted in a fresh LB medium and incubated at 37 °C. At mid-exponential growth phase indicated by arrows, each compound was added at an MIC as determined in Tables 1 and 2. In a control culture, 0.15% (v/v) of dimethyl sulfoxide (DMSO), 50 mg l−1 of ampicillin or chloramphenicol at a final concentration was added. The OD was measured at 600 nm.
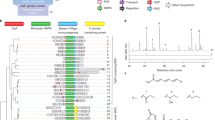
Twelve compounds shown in Figure 4 significantly inhibit at least three different GTPases (Figure 4b), have MICs less than 200 mg l−1 (Figure 4c), and both (Figure 4a). All compounds are structurally diverse forms, except for the pair of analogs SBI-34438 and -34439. Interestingly, all three compounds shown in Figure 4a and four of seven compounds shown in Figure 4c contain either benzoic acid or other aromatic group with carboxylic acid substitution. One compound, SBI-34703, has a dimerized indol scaffold. As we demonstrated in Table 1 and Figure 3a, SBI-34462, -34566 and -34612 are detrimental to S. aureus cells as well as to Der homologs. Therefore, they appear to be novel candidates for the next generation of antibiotics targeting Der GTPase.
The chemical structures and MWs of 12 compounds. The structures of 12 chemicals summarized in Tables 1 and 2 are presented. (a) The compounds with MICs of >25 or 50 mg l−1 that inhibit at least three GTPases. (b) The compounds with MICs of >200 mg l−1 that inhibit at least three GTPases. (c) The compounds with MICs of <200 mg l−1 that do not influence enzyme activity. Numbers indicate compound registration number by Sapient Discovery, LLC proprietary compound database. The MW of each compound is provided in the parentheses.
Models of Der GTPase complexes with bioactive compounds
Docking models of three bioactive compounds, SBI-34462, -34566 and -34612, with the GTP-binding site of GD1 domain with the lowest-energy closed conformation of Switch I loop are shown in Figs 5b–d, respectively. A model of GD1 in complex with GTP is displayed in Figure 5a for comparison. GTP-binding mode is very similar to that of Ras and other small GTPases.28 The guanine N1H and C2NH2 groups make two H-bonds with the Glu121 carboxyl, while guanine C6=O makes an H-bond with the backbone NH of Ala147. The ribose O2′H makes an H-bond with the carboxyl of Glu148 and the phosphate oxygens interact with backbone NH groups of residues 11–15. The loop residues 33–36 displayed at the top of Figures 5a–d contribute significantly to GTP binding. The Glu33 carboxyl and Gly34 NH groups make H-bonds with the ribose O3′H and O2′, respectively. The hydroxyl of Thr36 interacts with the γ-phosphate group. The threonine residue is highly conserved in Switch I loops of small GTPases. The common pharmacophore of the antibiotic compounds shown in Figs 5b–d is a carboxylic group attached to an aromatic ring, which makes multiple H-bonds with backbone NH groups of residues 12–15 and with ɛ-amino group of Lys14. Interactions of this carboxylic group mimic interactions of the β- and γ-phosphate groups of GTP. Aromatic rings of SBI-34462 (Figure 5b) and SBI-34612 (Figure 5d) are bound in the guanidine-binding site. These compounds, however, do not contain polar substituents that would make guanine-like H-bonds contributing to high binding affinity of GTP. Introduction of properly designed polar substituents to aromatic moieties of these compounds may significantly enhance their binding to Der GTP site. Only one compound, SBI-34612 (Figure 5d), makes a direct H-bond with Switch I loop. However, all three compounds make significant van der Waals interactions with the loop residues Gly34 and Val35 that may stabilize a closed conformation of the loop. The phenyl 3-CF3 group of SBI-34462 is located close to the carboxylic group of Glu33. These groups may interact through a water molecule. Introduction of a hydrogen bond donor at the position occupied by the CF3 group may enhance compound activity by direct H-binding with the Glu33 carboxyl.
Models of Der GD1 domain in complex with GTP (a) and three compounds with anti-bacterial activity, SBI-34462 (b), SBI-34566 (c) and SBI-34612 (d). The GTP-binding site of residues of Der GD1 domain is rendered as sticks colored by atom type with carbons colored gold. Ligands are shown as sticks with carbons colored yellow. Van der Waals surface of the protein-binding site is colored by atomic charges, with blue surface corresponding to positive charges and red surface corresponding to negative charges. Ligand–protein hydrogen bonds are shown as black dashed lines. The images were made using Accelrys DS Visualizer version 3.1 (Accelrys Software Inc., San Diego, CA, USA).
Analysis of Der binding modes and the structure–activity relationship of antibacterial activity found in this study suggest that future target compounds with high affinity to the active Der GTP-binding site in active state may contain the following pharmacophore groups: (a) benzoic acid or another aromatic group with carboxylic substitution that binds to the GTP phosphate-binding site, (b) an aromatic moiety mimicking guanine and (c) hydrogen bond donor interacting with the carbonyl of Glu33 residue of Switch I loop.
Among essential bacterial GTPases like Der, EF-Tu, IF-2 and FtsZ were previously studied as potential antibiotic targets because of their wide conservation in bacteria and critical functions such as translation and cell division within bacterial species, even though their eukaryotic homologs exist. Nevertheless, thiostrepton for IF-2, pulvomycin and GE2270A for EF-Tu, and PC190723, Berberine and Zantrins for FtsZ have been exploited and investigated for their inhibitory activity towards those GTPases.29, 30, 31, 32, 33, 34, 35, 36, 37, 38 The recent technological advances in computational modeling of protein structures have a crucial role in drug design and its retrieval. In our study, structure-based pharmacophore design screened a relatively small number of candidate molecules and we successfully identified three inhibitors (SBI-34462, -34566, and -34612) for an essential bacterial GTPase Der that has been unexploited for antibacterial target. Therefore, our lead compounds isolated in the present study provide scaffolds to improve and optimize those compounds through a further investigation of the molecular interactions between Der GTPase and putative antibiotics. This will enhance the development of novel antibiotics against antibiotic-resistant pathogenic bacteria.
References
Boneca, I. G. & Chiosis, G. Vancomycin resistance: occurrence, mechanisms and strategies to combat it. Expert Opin. Ther. Targets 7, 311–328 (2003).
Hartmann, G., Honikel, K. O., Knusel, F. & Nuesch, J. The specific inhibition of the DNA-directed RNA synthesis by rifamycin. Biochim. Biophys. Acta 145, 843–844 (1967).
Sugino, A., Higgins, N. P., Brown, P. O., Peebles, C. L. & Cozzarelli, N. R. Energy coupling in DNA gyrase and the mechanism of action of novobiocin. Proc. Natl Acad. Sci. USA 75, 4838–4842 (1978).
Bourne, H. R., Sanders, D. A. & Mccormick, F. The Gtpase superfamily - a conserved switch for diverse cell functions. Nature 348, 125–132 (1990).
Bourne, H. R., Sanders, D. A. & Mccormick, F. The Gtpase superfamily - conserved structure and molecular mechanism. Nature 349, 117–127 (1991).
Caldon, C. E. & March, P. E. Function of the universally conserved bacterial GTPases. Curr. Opin. Microbiol. 6, 135–139 (2003).
Sprang, S. R. G protein mechanisms: Insights from structural analysis. Annu. Rev. Biochem. 66, 639–678 (1997).
Hwang, J. & Inouye, M. An essential GTPase, Der, containing double GTP-binding domains from Escherichia coli and Thermotoga maritima. J. Biol. Chem. 276, 31415–31421 (2001).
Hwang, J. & Inouye, M. The tandem GTPase, Der, is essential for the biogenesis of 50S ribosomal subunits in Escherichia coli. Mol. Microbiol. 61, 1660–1672 (2006).
Hwang, J. & Inouye, M. RelA functionally suppresses the growth defect caused by a mutation in the G domain of the essential Der protein. J. Bacteriol. 190, 3236–3243 (2008).
Hwang, J. & Inouye, M. Interaction of an essential Escherichia coli GTPase, Der, with the 50S Ribosome via the KH-Like domain. J. Bacteriol. 192, 2277–2283 (2010).
Hwang, J. & Inouye, M. A bacterial GAP-like protein, Yihl, regulating the GTPase of Der, an essential GTP-binding protein in Escherichia coil. J. Mol. Biol. 399, 759–772 (2010).
Robinson, V. L., Hwang, J., Fox, E., Inouye, M. & Stock, A. M. Domain arrangement of Der, a switch protein containing two GTPase domains. Structure 10, 1649–1658 (2002).
Muench, S. P., Xu, L., Sedelnikova, S. E. & Rice, D. W. The essential GTPase YphC displays a major domain rearrangement associated with nucleotide binding. Proc. Natl Acad. Sci. USA 103, 12359–12364 (2006).
Tomar, S. K., Dhimole, N., Chatterjee, M. & Prakash, B. Distinct GDP/GTP bound states of the tandem G-domains of EngA regulate ribosome binding. Nucleic Acids Res. 37, 2359–2370 (2009).
Bharat, A., Jiang, M., Sullivan, S. M., Maddock, J. R. & Brown, E. D. Cooperative and critical roles for both G domains in the GTPase activity and cellular function of ribosome-associated Escherichia coli EngA. J. Bacteriol. 188, 7992–7996 (2006).
Tan, J., Jakob, U. & Bardwell, J. C. A. Overexpression of two different GTPases rescues a null mutation in a heat-induced rRNA methyltransferase. J. Bacteriol. 184, 2692–2698 (2002).
Nemethy, G. et al. Energy parameters in polypeptides.10. Improved geometrical parameters and nonbonded interactions for use in the Ecepp/3 algorithm, with application to proline-containing peptides. J. Phys. Chem. 96, 6472–6484 (1992).
Wittinghofer, F., Krengel, U., John, J., Kabsch, W. & Pai, E. F. 3-Dimensional structure of P21 in the active conformation and analysis of an oncogenic mutant. Environ. Health Perspect. 93, 11–15 (1991).
Milburn, M. V. et al. Molecular switch for signal transduction - structural differences between active and inactive forms of protooncogenic ras proteins. Science 247, 939–945 (1990).
Shenderovich, M. D., Kagan, R. M., Heseltine, P. N. R. & Ramnarayan, K. Structure-based phenotyping predicts HIV-1 protease inhibitor resistance. Protein Sci. 12, 1706–1718 (2003).
Dudek, M. J., Ramnarayan, K. & Ponder, J. W. Protein structure prediction using a combination of sequence homology and global energy minimization: II. Energy functions. J. Comput. Chem. 19, 548–573 (1998).
Wierenga, R. K. et al. Modeling, mutagenesis, and structural studies on the fully conserved phosphate-binding loop (loop 8) of triosephosphate isomerase: toward a new substrate specificity. Proteins Struct. Funct. Genet. 42, 383–389 (2001).
Pearlman, D. A. et al. Amber, a package of computer-programs for applying molecular mechanics, normal-mode analysis, molecular-dynamics and free-energy calculations to simulate the structural and energetic properties of molecules. Comput. Phys. Commun. 91, 1–41 (1995).
Ramnarayan, K., Rao, B. G. & Singh, U. C. The effect of polarization energy on the free-energy perturbation calculations. J. Chem. Phys. 92, 7057–7067 (1990).
Fuda, C., Suvorov, M., Vakulenko, S. B. & Mobashery, S. The basis for resistance to beta-lactam antibiotics by penicillin-binding protein 2a of methicillin-resistant Staphylococcus aureus. J. Biol. Chem. 279, 40802–40806 (2004).
Georgopapadakou, N. H. Penicillin-binding proteins and bacterial-resistance to beta-lactams. Antimicrob. Agents Chemother. 37, 2045–2053 (1993).
Wittinghofer, A. in GTPases: Frontiers in Molecular Biology (Oxford University Press, New York, 2000).
Anborgh, P. H., Okamura, S. & Parmeggiani, A. Effects of the antibiotic pulvomycin on the elongation factor Tu-dependent reactions. Comparison with other antibiotics. Biochemistry (Mosc.) 43, 15550–15556 (2004).
Andreu, J. M. et al. The antibacterial cell division inhibitor PC190723 is an FtsZ polymer-stabilizing agent that induces filament assembly and condensation. J. Biol. Chem. 285, 14239–14246 (2010).
Brandi, L. et al. The translation initiation functions of IF2: targets for thiostrepton inhibition. J. Mol. Biol. 335, 881–894 (2004).
Cetin, R. et al. Enacyloxin IIa, an inhibitor of protein biosynthesis that acts on elongation factor Tu and the ribosome. EMBO J. 15, 2604–2611 (1996).
Domadia, P. N., Bhunia, A., Sivaraman, J., Swarup, S. & Dasgupta, D. Berberine targets assembly of Escherichia coli cell division protein FtsZ. Biochemistry (Mosc.) 47, 3225–3234 (2008).
Mukherjee, S. et al. N-Benzyl-3-sulfonamidopyrrolidines as novel inhibitors of cell division in E-coli. Bioorg. Med. Chem. Lett. 17, 6651–6655 (2007).
Stokes, N. R. et al. Novel inhibitors of bacterial cytokinesis identified by a cell-based antibiotic screening assay. J. Biol. Chem. 280, 39709–39715 (2005).
Anborgh, P. H. & Parmeggiani, A. Probing the reactivity of the Gtp-bound and Gdp-bound conformations of elongation factor-Tu in complex with the antibiotic Ge2270-A. J. Biol. Chem. 268, 24622–24628 (1993).
Margalit, D. N. et al. Targeting cell division: small-molecule inhibitors of FtsZ GTPase perturb cytokinetic ring assembly and induce bacterial lethality. Proc. Natl Acad. Sci. USA 101, 11821–11826 (2004).
Haydon, D. J. et al. An inhibitor of FtsZ with potent and selective anti-staphylococcal activity. Science 321, 1673–1675 (2008).
Acknowledgements
We would like to thank Dr Sangita Phadtare for critical reading of this manuscript and suggestions. This work was supported by National Institutes of Health, Phase I SBIR Grant PHS 2003-2.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hwang, J., Tseitin, V., Ramnarayan, K. et al. Structure-based design and screening of inhibitors for an essential bacterial GTPase, Der. J Antibiot 65, 237–243 (2012). https://doi.org/10.1038/ja.2012.9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2012.9