Abstract
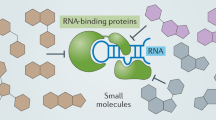
The development of innovative methodologies to identify RNA binders has attracted enormous attention in chemical biology and drug discovery. Although antibiotics targeting bacterial ribosomal RNA have been on the market for decades, the renewed interest in RNA targeting reflects the need to better understand complex intracellular processes involving RNA. In this context, small molecules are privileged tools used to explore the biological functions of RNA and to validate RNAs as therapeutic targets, and they eventually are to become new drugs. Despite recent progress, the rational design of specific RNA binders requires a better understanding of the interactions which occur with the RNA target to reach the desired biological response. In this Review, we discuss the challenges to approaching this underexplored chemical space, together with recent strategies to bind, interact and affect biologically relevant RNAs.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Childs-Disney, J. L. et al. Targeting RNA structures with small molecules. Nat. Rev. Drug Discov. 21, 736–762 (2022).
Crooke, S. T., Baker, B. F., Crooke, R. M. & Liang, X. H. Antisense technology: an overview and prospectus. Nat. Rev. Drug Discov. 20, 427–453 (2021).
Roberts, T. C., Langer, R. & Wood, M. J. A. Advances in oligonucleotide drug delivery. Nat. Rev. Drug Discov. 19, 673–694 (2020).
Falese, J. P., Donlic, A. & Hargrove, A. E. Targeting RNA with small molecules: from fundamental principles towards the clinic. Chem. Soc. Rev. 50, 2224–2243 (2021).
Warner, K. D., Hajdin, C. E. & Weeks, K. M. Principles for targeting RNA with drug-like small molecules. Nat. Rev. Drug Discov. 17, 547–558 (2018).
Zamani, F. & Suzuki, T. Synthetic RNA modulators in drug discovery. J. Med. Chem. 64, 7110–7155 (2021).
Magnet, S. & Blanchard, J. S. Molecular insights into aminoglycoside action and resistance. Chem. Rev. 105, 477–498 (2005).
Wilson, D. N. Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nat. Rev. Microbiol. 12, 35–48 (2014).
Thomas, J. R. & Hergenrother, P. J. Targeting RNA with small molecules. Chem. Rev. 108, 1171–1224 (2008).
Ratni, H., Scalco, R. S. & Stephan, A. H. Risdiplam, the first approved small molecule splicing modifier drug as a blueprint for future transformative medicines. ACS Med. Chem. Lett. 12, 874–877 (2021).
Di Giorgio, A. & Duca, M. Synthetic small-molecule RNA ligands: future prospects as therapeutic agents. MedChemComm 10, 1242–1255 (2019).
Ganser, L. R. et al. High-performance virtual screening by targeting a high-resolution RNA dynamic ensemble. Nat. Struct. Mol. Biol. 25, 425–434 (2018).
Padroni, G., Patwardhan, N. N., Schapira, M. & Hargrove, A. E. Systematic analysis of the interactions driving small molecule-RNA recognition. RSC Med. Chem. 11, 802–813 (2020).
Wang, S., Huber, P. W., Cui, M., Czarnik, A. W. & Mei, H. Y. Binding of neomycin to the TAR element of HIV-1 RNA induces dissociation of Tat protein by an allosteric mechanism. Biochemistry 37, 5549–5557 (1998).
Golkar, T. et al. Structural basis for plazomicin antibiotic action and resistance. Commun. Biol. 4, 729 (2021).
Aradi, K., Di Giorgio, A. & Duca, M. Aminoglycoside conjugation for RNA targeting: antimicrobials and beyond. Chem. Eur. J. 26, 12273–12309 (2020).
Bera, S., Mondal, D., Palit, S. & Schweizer, F. Structural modifications of the neomycin class of aminoglycosides. MedChemComm 7, 1499–1534 (2016).
Ennifar, E. et al. Structure-guided discovery of a novel aminoglycoside conjugate targeting HIV-1 RNA viral genome. ACS Chem. Biol. 8, 2509–2517 (2013).
Blount, K. F. & Tor, Y. A tale of two targets: differential RNA selectivity of nucleobase-aminoglycoside conjugates. ChemBioChem 7, 1612–1621 (2006).
Gebert, L. F. R. & MacRae, I. J. Regulation of microRNA function in animals. Nat. Rev. Mol. Cell. Biol. 20, 21–37 (2019).
Maucort, C. et al. Design and implementation of synthetic RNA binders for the inhibition of miR-21 biogenesis. ACS Med. Chem. Lett. 12, 899–906 (2021).
Vo, D. D. et al. Building of neomycin-nucleobase-amino acid conjugates for the inhibition of oncogenic miRNAs biogenesis. Org. Biomol. Chem. 16, 6262–6274 (2018).
Vo, D. D. et al. Targeting the production of oncogenic microRNAs with multimodal synthetic small molecules. ACS Chem. Biol. 9, 711–721 (2014).
Vo, D. D. et al. Oncogenic MicroRNAs biogenesis as a drug target: structure-activity relationship studies on new aminoglycoside conjugates. Chem. Eur. J. 22, 5350–5362 (2016).
Malnuit, V., Duca, M. & Benhida, R. Targeting DNA base pair mismatch with artificial nucleobases. Advances and perspectives in triple helix strategy. Org. Biomol. Chem. 9, 326–336 (2011).
Costales, M. G. et al. Small molecule inhibition of microRNA-210 reprograms an oncogenic hypoxic circuit. J. Am. Chem. Soc. 139, 3446–3455 (2017).
Disney, M. D. et al. Two-dimensional combinatorial screening identifies specific aminoglycoside-RNA internal loop partners. J. Am. Chem. Soc. 130, 11185–11194 (2008).
Velagapudi, S. P. & Disney, M. D. Two-dimensional combinatorial screening enables the bottom-up design of a microRNA-10b inhibitor. Chem. Commun. 50, 3027–3029 (2014).
Davis, B. et al. Rational design of inhibitors of HIV-1 TAR RNA through the stabilisation of electrostatic “hot spots”. J. Mol. Biol. 336, 343–356 (2004).
Patwardhan, N. N. et al. Amiloride as a new RNA-binding scaffold with activity against HIV-1 TAR. MedChemComm 8, 1022–1036 (2017).
Davila-Calderon, J. et al. IRES-targeting small molecule inhibits enterovirus 71 replication via allosteric stabilization of a ternary complex. Nat. Commun. 11, 4775 (2020).
Patwardhan, N. N., Cai, Z., Newson, C. N. & Hargrove, A. E. Fluorescent peptide displacement as a general assay for screening small molecule libraries against RNA. Org. Biomol. Chem. 17, 1778–1786 (2019).
Zafferani, M. et al. Amilorides inhibit SARS-CoV-2 replication in vitro by targeting RNA structures. Sci. Adv. 7, eabl6096 (2021).
Wilson, L., Gage, P. & Ewart, G. Hexamethylene amiloride blocks E protein ion channels and inhibits coronavirus replication. Virology 353, 294–306 (2006).
Hagler, L. D. et al. Versatile target-guided screen for discovering bidirectional transcription inhibitors of a trinucleotide repeat disease. ACS Med. Chem. Lett. 12, 935–940 (2021).
Hagler, L. D. et al. Expanded DNA and RNA trinucleotide repeats in myotonic dystrophy type 1 select their own multitarget, sequence-selective inhibitors. Biochemistry 59, 3463–3472 (2020).
Mirkin, S. M. Expandable DNA repeats and human disease. Nature 447, 932–940 (2007).
Wong, C. H. et al. Targeting toxic RNAs that cause myotonic dystrophy type 1 (DM1) with a bisamidinium inhibitor. J. Am. Chem. Soc. 136, 6355–6361 (2014).
Krueger, S. B., Lanzendorf, A. N., Jeon, H. H. & Zimmerman, S. C. Selective and reversible ligand assembly on the DNA and RNA repeat sequences in myotonic dystrophy. ChemBioChem 23, e202200260 (2022).
Pushechnikov, A. et al. Rational design of ligands targeting triplet repeating transcripts that cause RNA dominant disease: application to myotonic muscular dystrophy type 1 and spinocerebellar ataxia type 3. J. Am. Chem. Soc. 131, 9767–9779 (2009).
Jahromi, A. H. et al. A novel CUG(exp).MBNL1 inhibitor with therapeutic potential for myotonic dystrophy type 1. ACS Chem. Biol. 8, 1037–1043 (2013).
Chien, C. M. et al. Structural basis for targeting T:T mismatch with triaminotriazine-acridine conjugate induces a U-shaped head-to-head four-way junction in CTG repeat DNA. J. Am. Chem. Soc. 142, 11165–11172 (2020).
Murata, A., Otabe, T., Zhang, J. & Nakatani, K. BzDANP, a small-molecule modulator of pre-miR-29a maturation by dicer. ACS Chem. Biol. 11, 2790–2796 (2016).
Lombes, T. et al. Investigation of RNA-ligand interactions by 19F NMR spectroscopy using fluorinated probes. Angew. Chem. Int. Ed. Engl. 51, 9530–9534 (2012).
Murata, A., Nakamori, M. & Nakatani, K. Modulating RNA secondary and tertiary structures by mismatch binding ligands. Methods 167, 78–91 (2019).
Murata, A. et al. Small molecule-induced dimerization of hairpin RNA interfered with the Dicer cleavage reaction. Biochemistry 60, 245–249 (2021).
Konieczny, P. et al. Cyclic mismatch binding ligands interact with disease-associated CGG trinucleotide repeats in RNA and suppress their translation. Nucleic Acids Res. 49, 9479–9495 (2021).
Mukherjee, S. et al. HT-SELEX-based identification of binding pre-miRNA hairpin-motif for small molecules. Mol. Ther. Nucleic Acids 27, 165–174 (2022).
Howe, J. A. et al. Selective small-molecule inhibition of an RNA structural element. Nature 526, 672–677 (2015).
Howe, J. A. et al. Atomic resolution mechanistic studies of ribocil: a highly selective unnatural ligand mimic of the E. coli FMN riboswitch. RNA Biol. 13, 946–954 (2016).
Aguilar, R. et al. Targeting Xist with compounds that disrupt RNA structure and X inactivation. Nature 604, 160–166 (2022).
Carrette, L. L. G. et al. A mixed modality approach towards Xi reactivation for Rett syndrome and other X-linked disorders. Proc. Natl Acad. Sci. USA 115, E668–E675 (2018).
Ursu, A. et al. Design of small molecules targeting RNA structure from sequence. Chem. Soc. Rev. 49, 7252–7270 (2020).
Velagapudi, S. P., Seedhouse, S. J. & Disney, M. D. Structure-activity relationships through sequencing (StARTS) defines optimal and suboptimal RNA motif targets for small molecules. Angew. Chem. Int. Ed. Engl. 49, 3816–3818 (2010).
Disney, M. D. et al. Inforna 2.0: a platform for the sequence-based design of small molecules targeting structured RNAs. ACS Chem. Biol. 11, 1720–1728 (2016).
Velagapudi, S. P., Gallo, S. M. & Disney, M. D. Sequence-based design of bioactive small molecules that target precursor microRNAs. Nat. Chem. Biol. 10, 291–297 (2014).
Haniff, H. S. et al. A structure-specific small molecule inhibits a miRNA-200 family member precursor and reverses a type 2 diabetes phenotype. Cell Chem. Biol. 29, 300–311.e10 (2022).
Chen, J. L. et al. Design, optimization, and study of small molecules that target tau pre-mRNA and affect splicing. J. Am. Chem. Soc. 142, 8706–8727 (2020).
Dibrov, S. M. et al. Structure of a hepatitis C virus RNA domain in complex with a translation inhibitor reveals a binding mode reminiscent of riboswitches. Proc. Natl Acad. Sci. USA 109, 5223–5228 (2012).
Shibata, T. et al. Small molecule targeting r(UGGAA)n disrupts RNA foci and alleviates disease phenotype in Drosophila model. Nat. Commun. 12, 236 (2021).
Boehr, D. D., Nussinov, R. & Wright, P. E. The role of dynamic conformational ensembles in biomolecular recognition. Nat. Chem. Biol. 5, 789–796 (2009).
Nakatani, K. Possibilities and challenges of small molecule organic compounds for the treatment of repeat diseases. Proc. Jpn Acad. Ser. B Phys. Biol. Sci. 98, 30–48 (2022).
Vogt, A. D. & Di Cera, E. Conformational selection or induced fit? A critical appraisal of the kinetic mechanism. Biochemistry 51, 5894–5902 (2012).
Bailor, M. H., Mustoe, A. M., Brooks, C. L. III & Al-Hashimi, H. M. Topological constraints: using RNA secondary structure to model 3D conformation, folding pathways, and dynamic adaptation. Curr. Opin. Struct. Biol. 21, 296–305 (2011).
Marusic, M., Toplishek, M. & Plavec, J. NMR of RNA — structure and interactions. Curr. Opin. Struct. Biol. 79, 102532 (2023).
Shortridge, M. D. et al. Drug-like small molecules that inhibit expression of the oncogenic microRNA-21. ACS Chem. Biol. 18, 237–250 (2023).
Campagne, S. et al. Structural basis of a small molecule targeting RNA for a specific splicing correction. Nat. Chem. Biol. 15, 1191–1198 (2019).
Moumne, R. et al. Fluorinated diaminocyclopentanes as chiral sensitive NMR probes of RNA. Struct. J. Am. Chem. Soc. 132, 13111–13113 (2010).
Bevilacqua, P. C., Ritchey, L. E., Su, Z. & Assmann, S. M. Genome-wide analysis of RNA secondary structure. Annu. Rev. Genet. 50, 235–266 (2016).
Deigan, K. E., Li, T. W., Mathews, D. H. & Weeks, K. M. Accurate SHAPE-directed RNA structure determination. Proc. Natl Acad. Sci. USA 106, 97–102 (2009).
Siegfried, N. A., Busan, S., Rice, G. M., Nelson, J. A. & Weeks, K. M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods 11, 959–965 (2014).
Weeks, K. M. SHAPE directed discovery of new functions in large RNAs. Acc. Chem. Res. 54, 2502–2517 (2021).
Zeller, M. J. et al. SHAPE-enabled fragment-based ligand discovery for RNA. Proc. Natl Acad. Sci. USA 119, e2122660119 (2022).
Wang, Y., Parmar, S., Schneekloth, J. S. & Tiwary, P. Interrogating RNA-small molecule interactions with structure probing and artificial intelligence-augmented molecular simulations. ACS Cent. Sci. 8, 741–748 (2022).
Velagapudi, S. P., Li, Y. & Disney, M. D. A cross-linking approach to map small molecule-RNA binding sites in cells. Bioorg. Med. Chem. Lett. 29, 1532–1536 (2019).
Suresh, B. M. et al. A general fragment-based approach to identify and optimize bioactive ligands targeting RNA. Proc. Natl Acad. Sci. USA 117, 33197–33203 (2020).
Balaratnam, S. et al. A chemical probe based on the PreQ(1) metabolite enables transcriptome-wide mapping of binding sites. Nat. Commun. 12, 5856 (2021).
Tong, Y. et al. Transcriptome-wide mapping of small-molecule RNA-binding sites in cells informs an isoform-specific degrader of QSOX1 mRNA. J. Am. Chem. Soc. 144, 11620–11625 (2022).
Vlassov, V. V., Zuber, G., Felden, B., Behr, J. P. & Giege, R. Cleavage of tRNA with imidazole and spermine imidazole constructs: a new approach for probing RNA structure. Nucleic Acids Res. 23, 3161–3167 (1995).
Tamkovich, N. et al. Design, RNA cleavage and antiviral activity of new artificial ribonucleases derived from mono-, di- and tripeptides connected by linkers of different hydrophobicity. Bioorg. Med. Chem. 24, 1346–1355 (2016).
Martin, C. et al. Design, synthesis, and evaluation of neomycin-imidazole conjugates for RNA cleavage. ChemPlusChem 87, e202200250 (2022).
Hecht, S. M. Bleomycin: new perspectives on the mechanism of action. J. Nat. Prod. 63, 158–168 (2000).
Angelbello, A. J. & Disney, M. D. Bleomycin can cleave an oncogenic noncoding RNA. ChemBioChem 19, 43–47 (2018).
Li, Y. & Disney, M. D. Precise small molecule degradation of a noncoding RNA identifies cellular binding sites and modulates an oncogenic phenotype. ACS Chem. Biol. 13, 3065–3071 (2018).
Angelbello, A. J. et al. Precise small-molecule cleavage of an r(CUG) repeat expansion in a myotonic dystrophy mouse model. Proc. Natl Acad. Sci. USA 116, 7799–7804 (2019).
Liu, X. et al. Targeted degradation of the oncogenic microRNA 17-92 cluster by structure-targeting ligands. J. Am. Chem. Soc. 142, 6970–6982 (2020).
Haniff, H. S. et al. Targeting the SARS-CoV-2 RNA genome with small molecule binders and ribonuclease targeting chimera (RIBOTAC) degraders. ACS Cent. Sci. 6, 1713–1721 (2020).
Li, K. & Crews, C. M. PROTACs: past, present and future. Chem. Soc. Rev. 51, 5214–5236 (2022).
Lovering, F., Bikker, J. & Humblet, C. Escape from flatland: increasing saturation as an approach to improving clinical success. J. Med. Chem. 52, 6752–6756 (2009).
Morgan, B. S. & Hargrove, A. E. Synthetic Receptors for Biomolecules: Design Principles and Applications 253–325 (The Royal Society of Chemistry, 2015).
Kenyon, J., Prestwood, L. & Lever, A. Current perspectives on RNA secondary structure probing. Biochem. Soc. Trans. 42, 1251–1255 (2014).
Batey, R. T., Rambo, R. P. & Doudna, J. A. Tertiary motifs in RNA structure and folding. Angew. Chem. Int. Ed. Engl. 38, 2326–2343 (1999).
Butcher, S. E. & Pyle, A. M. The molecular interactions that stabilize RNA tertiary structure: RNA motifs, patterns, and networks. Acc. Chem. Res. 44, 1302–1311 (2011).
Jones, C. P. & Ferré-D’Amaré, A. R. RNA quaternary structure and global symmetry. Trends Biochem. Sci. 40, 211–220 (2015).
Donlic, A. et al. R-BIND 2.0: an updated database of bioactive RNA-targeting small molecules and associated RNA secondary structures. ACS Chem. Biol. 17, 1556–1566 (2022).
Morgan, B. S., Forte, J. E., Culver, R. N., Zhang, Y. & Hargrove, A. E. Discovery of key physicochemical, structural, and spatial properties of RNA-targeted bioactive ligands. Angew. Chem. Int. Ed. Engl. 56, 13498–13502 (2017).
Xie, J. & Frank, A. T. Mining for ligandable cavities in RNA. ACS Med. Chem. Lett. 12, 928–934 (2021).
Murray, C. W. & Rees, D. C. The rise of fragment-based drug discovery. Nat. Chem. 1, 187–192 (2009).
Umuhire Juru, A., Cai, Z., Jan, A. & Hargrove, A. E. Template-guided selection of RNA ligands using imine-based dynamic combinatorial chemistry. Chem. Commun. 56, 3555–3558 (2020).
Rzuczek, S. G., Park, H. & Disney, M. D. A toxic RNA catalyzes the in cellulo synthesis of its own inhibitor. Angew. Chem. Int. Ed. Engl. 53, 10956–10959 (2014).
Benhamou, R. I. et al. DNA-encoded library versus RNA-encoded library selection enables design of an oncogenic noncoding RNA inhibitor. Proc. Natl Acad. Sci. USA 119, e2114971119 (2022).
Rizvi, N. F. et al. Discovery of selective RNA-binding small molecules by affinity-selection mass spectrometry. ACS Chem. Biol. 13, 820–831 (2018).
Acknowledgements
We thank the Italian Institute of Technology (IIT) and Université Côte d’Azur for the financial support to M.P. as PhD fellowship. We also thank BoostUrCareer COFUND project with Région PACA and European Union’s Horizon 2020 research and innovation programme under grant agreement no. 847581 for PhD fellowship to S.K. and the National Center for Gene Therapy and Drugs based on RNA Technology project (CN00000041) funded by European Union’s NextGenerationEU PNRR MUR - M4C2 – programme (CUP J33C22001130001).
Author information
Authors and Affiliations
Contributions
All authors researched data for the article. M.D., S.A. and A.D.G. contributed substantially to the discussion of the content. S.K. and M.P. contributed substantially to the illustrations of the article. M.D. wrote the article. All authors reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Chemistry thanks Matthew Disney, Jessica Bush and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kovachka, S., Panosetti, M., Grimaldi, B. et al. Small molecule approaches to targeting RNA. Nat Rev Chem 8, 120–135 (2024). https://doi.org/10.1038/s41570-023-00569-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41570-023-00569-9