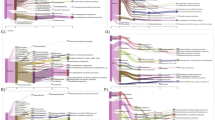
Abstract
Aerobic methane-oxidizing bacteria (MOB) use a restricted substrate range, yet >30 species-equivalent operational taxonomical units (OTUs) are found in one paddy soil. How these OTUs physically share their microhabitat is unknown. Here we highly resolved the vertical distribution of MOB and their activity. Using microcosms and cryosectioning, we sub-sampled the top 3-mm of a water-saturated soil at near in situ conditions in 100-μm steps. We assessed the community structure and activity using the particulate methane monooxygenase gene pmoA as a functional and phylogenetic marker by terminal restriction fragment length polymorphism (t-RFLP), a pmoA-specific diagnostic microarray, and cloning and sequencing. pmoA genes and transcripts were quantified using competitive reverse transcriptase PCR combined with t-RFLP. Only a subset of the methanotroph community was active. Oxygen microprofiles showed that 89% of total respiration was confined to a 0.67-mm-thick zone immediately above the oxic–anoxic interface, most probably driven by methane oxidation. In this zone, a Methylobacter-affiliated OTU was highly active with up to 18 pmoA transcripts per cell and seemed to be adapted to oxygen and methane concentrations in the micromolar range. Analysis of transcripts with a pmoA-specific microarray found a Methylosarcina-affiliated OTU associated with the surface zone. High oxygen but only nanomolar methane concentrations at the surface suggested an adaptation of this OTU to oligotrophic conditions. No transcripts of type II methanotrophs (Methylosinus, Methylocystis) were found, which indicated that this group was represented by resting stages only. Hence, different OTUs within a single guild shared the same microenvironment and exploited different niches.
Similar content being viewed by others
Introduction
Methane is, next to water vapor and carbon dioxide, the most important greenhouse gas (Intergovernmental Panel on Climate Change, 2007), with natural wetlands and irrigated rice fields together emitting about one-third of the total (Conrad, 2009). Their contribution would be even higher without the activity of aerobic methane-oxidizing bacteria (MOB), which act as a biofilter, mitigating emissions to the atmosphere (Reeburgh et al., 1993). MOB use methane as the sole source of carbon and energy, provided oxygen is available (Trotsenko and Murrell, 2008). Owing to this dual dependency, they thrive at oxic–anoxic interfaces, where both substrates are supplied (Brune et al., 2000). In flooded soils and sediments, these interfaces are located at the soil surface and in the rhizosphere of macrophytes when present (Bosse and Frenzel, 1997; Bodelier et al., 2006). Rhizospheric MOB in both paddy fields and natural wetlands have been often studied (Calhoun and King, 1997; Eller and Frenzel, 2001; Sorrell et al., 2002; Shrestha et al., 2008; Vishwakarma et al., 2009), but work at the soil surface has been mainly focused on process measurements (Conrad and Rothfuss, 1991; Frenzel et al., 1992; Bosse et al., 1993). The soil surface is characterized by sharp counter-gradients of oxidized and reduced species. Where these gradients overlap, ⩾90% of potentially emitted methane is oxidized (Frenzel et al., 1990; Conrad and Rothfuss, 1991).
MOB can be divided into two major groups, type I and type II, being equivalent to the families Methylococcaceae (γ-proteobacteria), and Methylocystaceae and Bejerinckiaceae (α-proteobacteria), respectively. The key enzyme of all MOB is methane monooxygenase (MMO), and the overwhelming majority of cultivated MOB possess a membrane-bound MMO (particulate MMO). Only the genera Methylocella and Methyloferula lack this enzyme and instead have a soluble MMO (sMMO) (Dedysh et al., 2000; Dedysh, 2009; Vorobev et al., 2011). The pmoA gene, which encodes the β-subunit of particulate MMO, is an excellent functional marker for studying MOB in most environments (McDonald and Murrell, 1997; Dumont and Murrell, 2005; McDonald et al., 2008). Its phylogeny reflects very well that of the 16S rRNA gene (Kolb et al., 2003; Degelmann et al., 2010). Type I MOB can be further divided into type Ia (for example, Methylomonas, Methylobacter, Methylosarcina and Methylomicrobium), and type Ib (for example, Methylococcus and Methylocaldum). Recently, MOB belonging to the phylum Verrucomicrobia have been isolated, but these seem to be restricted to extreme environments (Dunfield et al., 2007; Pol et al., 2007; Op den Camp et al., 2009).
Sanger- and pyro-sequencing have provided a large database of high-quality pmoA sequences (Lüke et al., 2010; Lüke and Frenzel, 2011). Based on pmoA phylogeny, type II MOB form a coherent cluster well represented by cultivated strains. Many distinct groups lacking cultivated representatives have been allocated to type I MOB. Numerous sequences are located at an intermediate position between the pmoA gene of MOB and the amoA gene of ammonia oxidizers. The substrate of the enzymes encoded by these sequences remains uncertain, with the exception of a few proven methane (Stoecker et al., 2006; Dunfield et al., 2007) and alkane oxidizers (Sayavedra-Soto et al., 2011; Coleman et al., 2012).
More than 30 operational taxonomical units (OTUs) corresponding to the species level have been found in a single paddy soil (Lüke et al., 2010). This raises the question whether and how their niches are separated, and whether all these individual OTUs really contribute to overall methane oxidation. However, not all of these 30 OTUs need to be active at the same time as methanotrophs form resting stages (Whittenbury et al., 1970a, 1970b). Indeed, the development and activity of methanotroph communities has been suggested to vary depending on methane flow (Krause et al., 2012), nitrogen supply (Rudd et al., 1976; Graham et al., 1993; Bodelier et al., 2000a, 2000b; Noll et al., 2008), disturbance (Ho et al., 2011) and grazing (Murase and Frenzel, 2008). At the macroscale, MOB community structure within a paddy field varies randomly, probably as ploughing prevents the development of explicit spatial patterns (Krause et al., 2009). Rice roots select for specific communities, thereby favouring the growth of certain OTUs (Lüke et al., 2011). Another aspect of spatial organization, however, has not yet been addressed — the community structure at the microscale. We focused on the soil surface and hypothesized that activity would be highest right at the oxic–anoxic interface, which would potentially separate MOB according to substrate availability, for example, high methane/low oxygen in deeper layers and low methane/high oxygen in shallower layers.
The study of gradient organisms requires a physical model that mimics naturally occurring gradients. With MOB, these are primarily the counter-gradients of oxygen and methane (Gilbert and Frenzel, 1998). We constructed microcosms that allow incubation of the top 3-mm of a water-saturated soil at near in situ conditions (Murase and Frenzel, 2007). When methane was supplied from below and air was supplied from above, a functioning methanotrophic community developed within a few days, oxidizing virtually all the methane that otherwise would have passed through this soil layer. We used cryosectioning (Murase et al., 2006) to subsample the soil from top to bottom in 100-μm steps. Focusing on pmoA as a functional and phylogenetic marker, we analyzed genes and transcripts along this depth profile, using pmoA transcripts as a proxy for species-specific activity. The community structure was assessed by terminal restriction fragment length polymorphism (t-RFLP), by a pmoA-specific diagnostic microarray (Bodrossy et al., 2003), and by cloning and sequencing. Oxygen microprofiles were used to model the depth-dependent oxygen consumption rate (Berg et al., 1998). Rates were correlated to copy numbers of pmoA genes and transcripts. Transcripts were quantified using a combination of competitive reverse transcriptase PCR (RT-PCR) and t-RFLP.
In addition to pmoA, we used mmoX encoding for a subunit of the sMMO to search for genes and transcripts of aerobic MOB that lack a pmoA. Three different primer sets were applied to DNA and RNA extracted from microcosms incubated for 2, 4 and 6 weeks, respectively. While this design allows to cover potential successional changes, the chosen primer sets are expected to target a wide range of mmoX diversity.
Materials and methods
Soil microcosm incubation and sampling
The construction and setup of the microcosms have been described previously (Murase and Frenzel, 2007). Briefly, 14 g dry rice field soil from Vercelli (Italy) was saturated with 7 ml demineralized water and incubated on a polytetrafluoroethylene membrane, which divided the microcosm into an upper and a lower compartment. The upper compartment contained oxygen at atmospheric concentrations, while the lower chamber was connected to an external reservoir (volume 1 l) with nitrogen gas supplemented with methane (15%).
We set up four microscosms (1–4). During incubation, methane and oxygen concentrations were monitored by gas chromatography. Methane was added regularly to the reservoir keeping concentration stable (s.e. 0.16% CH4, n=18). The microcosms were incubated in the dark for 14 days at 25 °C. Present and active methanotroph populations show some succession, but most changes occur from 25 days onwards (Krause et al., 2010). Prior to sampling, vertical oxygen profiles were determined using an oxygen microelectrode (OX50, Unisense, Aarhus, Denmark). The microcosms were then shock frozen in liquid nitrogen and stored at −80 °C until further analysis. For sub-sampling, the frozen soil was attached to a pre-cooled stage with Tissue-Tek OCT Compound (Sakura Finetek, Staufen, Germany). A cryotome (cryostat HM500M, MICROM, Walldorf, Germany) was used to prepare subsamples 100 μm thick (30 subsamples per microcosm). The subsamples were stored in 500 μl RNAlater-ICE (Ambion, Austin, TX, USA) at −20 °C for subsequent nucleic acid extraction.
Nucleic acid extraction
DNA and RNA were extracted following the protocol of Lueders et al. (2004) with minor modifications (Krause et al., 2010). RNA was prepared by digestion of 1 mg total nucleic acid with RQ1 RNase-free DNase (Promega, Madison, WI, USA) and subsequent purification using the RNeasy Mini Kit (Qiagen, Hilden, Germany) according to manufacturer’s instructions. pmoA transcripts were enriched using the mRNA-only Prokaryotic mRNA Isolation Kit (Epicentre Biotechnologies, Madison, WI, USA) and again purified with the RNeasy Mini Kit (Qiagen).
Complementary DNA (cDNA) synthesis and pmoA amplification
cDNA was synthesized and pmoA was amplified using the One-step Access RT-PCR System (Promega) with the forward primer A189f (5′-GGNGACTGGGACTTCTGG-3′) (Holmes et al., 1995) and the reverse primer mb661r (5′-CCGGMGCAACGTCYTTACC-3′) (Costello and Lidstrom, 1999). For t-RFLP, the forward primer was FAM labeled. For microarray analysis, the reverse primer contained the T7 promoter site (Bodrossy et al., 2003). A total of 1 μl purified template RNA was mixed with 5 μl AMV/Tfl 5 × reaction buffer (Promega), 0.01 mg bovine serum albumin (Roche, Basel, Switzerland), 2.5 nmol of each dNTP (Promega), 8 pmol of each primer, 25 nmol MgSO4 (Promega), 5% (v/v) DMSO, 20 U RNasin Plus (Promega), 2.5 U Tfl DNA polymerase (Promega), 2.5 U AMV reverse transcriptase (Promega) and molecular-grade water (Sigma-Aldrich, Munich, Germany) in a total volume of 25 μl. Reactions without AMV reverse transcriptase were used to check for DNA contamination. The first strand of cDNA was synthesized at 45 °C in 45 min, followed by 2 min at 94 °C to inactivate the AMV reverse transcriptase. The template was amplified in 35 cycles (30 s at 94 °C, 1 min at 55 °C, 1 min at 68 °C, final elongation 7 min at 68 °C). PCR products were checked on a 1% agarose gel and extracted from the gel using the QIAquick Gel Extraction Kit (Qiagen). Genomic copies of the pmoA gene were amplified following the same protocol, but without the initial cDNA synthesis step.
Cloning and sequencing
Cloning and sequencing was done as described before (Lüke et al., 2010).
t-RFLP analysis
The purified PCR product (100 ng) was digested with FastDigest MspI enzyme (Fermentas, St Leon-Rot, Germany) at 37 °C for 6 min. Digested samples were purified with Post-Reaction Clean-Up Spin Columns (Sigma-Aldrich) according to the manufacturer’s instructions. A total of 2 μl of each purified sample was mixed with 11 μl Hi-Di Formamide (Applied Biosystems, Foster City, CA, USA) and 0.2 μl of an internal DNA fragment length standard (MapMarker 1000, 50–1000 bp, x-rhodamine, Eurogentec, Ougree, Belgium) and denatured for 2 min at 94 °C. The terminal restriction fragments (tRFs) were separated and detected with capillary electrophoresis and an automatic sequencer (3130 Genetic Analyzer, Applied Biosystems; 30 min at 15 kV and 9 μA). The tRF patterns were analyzed with GeneMapper Version 4.0 (Applied Biosystems).
Microarray analysis
In vitro transcription, fragmentation, hybridization, scanning and data analysis were performed as described elsewhere (Stralis-Pavese et al., 2004; Stralis-Pavese et al., 2011).
Competitive t-RFLP
A competitive PCR (cPCR) assay (Han and Semrau, 2004) was adapted to quantify copy numbers of the pmoA gene and its transcripts. The assay was optimized for the most abundant and active OTU. A vector-born copy of an environmental Methylobacter-related pmoA gene (accession number) was used for standard preparation. Primers were A189f_T7 (5′-TAATACGACTCACTATAGGGGGNGACTGGGACTTCTGG-3′) and Inner-rev-661 (5′-CCGGMGCAACGTCYTTACCACTCAGGAGTACCAGTTCTT-3′). Concentrations of DNA and RNA standards were determined using RiboGreen and PicoGreen, respectively (Molecular Probes Inc., Eugene, OR, USA). For each sample, a minimum of three PCR or RT-PCR amplifications was performed as described above. Each reaction contained equal amounts of the environmental template, but varying standard concentrations. After amplification, PCR products were processed and analyzed by t-RFLP. We regressed the logarithms of standard-to-sample ratios to the logarithms of standard added; the amount of standard at the equivalence point equaled the unknown copy number (Freeman et al., 1999). For further details and an example see Supplement 1.
Statistical analysis
t-RFLP data were analyzed as described before (Krause et al., 2010). Briefly, the tRFs were binned to OTUs based on an in silico analysis of about 500 sequences from field and greenhouse experiments of Vercelli soil (Lüke et al., 2010). tRF profiles were standardized (Dunbar et al., 2001) and expressed as fractions. For microarray analysis, signals were standardized (i) against the mean of the overall array intensities (Lüke et al., 2011) and (ii) against an experimentally determined reference value for positive detection (Bodrossy et al., 2003). Statistical analysis and graphics were done in R (R Development Core Team, 2011). Analysis of similarity and non-metric multidimensional scaling (using the function metaMDS) were done with the vegan package, version 2.1-0 (Oksanen et al., 2011).
Soil microcosms for mmoX analyses
To evaluate the potential role of sMMO, we used another 8 microscosms that were installed and incubated as described above. Two microcosms were killed after 2 weeks, and three microcosms each after 4 and 6 weeks, respectively. Soil was sampled in 0.5 g aliquots, shock frozen in liquid nitrogen, and stored at −80 °C till further analysis. DNA and RNA were simultaneously extracted and prepared as described above. RNA samples were reverse transcribed with random hexamer primers and SuperScript III reverse transcriptase (Invitrogen, Darmstadt, Germany). Amplification of mmoX gene and transcript sequences was done using (i) primer set mmoX206f/886r (Hutchens et al., 2004, ii) primer set mmoXf92/r1430 (Islam et al., 2008), and (iii) primer set mmoXLF/LR (Rahman et al., 2011). Primer set mmoX206f/886r covers a fairly wide range of mmoX diversity, whereas mmoXf92/r1430 includes verrrucomicrobial sequences. Primer set mmoXLF/LR is specific for Methylocella. To check for cDNA quality, we amplified both rRNA and pmoA with primer sets 8F/1392R (Amann et al., 1995) and A189f/682r (Holmes et al., 1995), respectively. We got products from all samples. Amplicons generated with mmoX206f/886r from DNA extracted from microcosms after 2 and 6 weeks of incubation were cloned and sequenced as described above. Sequencing was carried out by GATC (GATC Biotech AG, Konstanz, Germany). Phylogenetic trees were constructed from sequence data using the ARB software package (Ludwig et al., 2004).
Sequences
pmoA sequence data have been submitted to EMBL under accession numbers HE805099–HE805112. mmoX sequence data have been submitted to GenBank under accession numbers JQ889714–JQ889792.
Results
Methane oxidation and t-RFLP analysis
We followed methane concentrations over time in both the lower and upper compartments of the microcosms. The lower compartment with the methane source simulated the methanogenic soil layer. The upper compartment, or headspace, was replenished with air every 2 days. Initially, in all microcosms, up to 3% methane accumulated in the headspace, but after 5 days of incubation, accumulation ceased resulting in an average headspace concentration of 228 ppmv CH4 which indicated the presence of an active and efficient methanotrophic community.
After 2 weeks of incubation, the oxic–anoxic interface in all microcosms stabilized between 1.5 and 1.7 mm (Figure 1a). After 14 days, the soil of four microcosms was shock frozen with liquid nitrogen, removed intact, mounted in a cryotome and sub-sampled in 100-μm layers by sectioning. A total of 30 subsamples per microcosm were analyzed (total n=120).
Vertical profiles of oxygen concentration and of tRFs derived from pmoA genes and their transcripts. Data are from microcosm 1. (a) Average oxygen profile (± s.e., n=4) measured with a microelectrode. Relative abundance of pmoA tRFs derived from (b) DNA and (c) RNA. The phylogenetic affiliation of tRFs is given together with their size in brackets. rRNA: combined relative abundances of different tRFs derived from unspecific reverse and amplification of 16S and 23S rRNA sequences; RPCs: rice paddy clusters (Lüke et al., 2010).
For an initial overview, we analyzed the genes and transcripts by pmoA t-RFLP. We assigned OTUs based on an in silico analysis of 500 pmoA clone sequences plus another 3500 sequences retrieved by pyrosequencing (Lüke et al., 2011; Lüke and Frenzel, 2011). The assignments were supported by 15 sequences generated from cloned mRNA (another 80 clones were derived from rRNA, see below). Figure 1 shows a synopsis of the average oxygen microprofile and the DNA and mRNA-based t-RFLP profiles from microcosm 1. The DNA-based pattern was diverse, with dominating fragments identified as type Ia (Methylobacter) and Ib, and as type II (Methylocystis and Methylosinus). The type-II-specific fragment was dominant below 2.2 mm depth, whereas fragments assigned to type Ib MOB had their highest relative abundance around the oxic–anoxic interface between 1.0 and 2.0 mm depth (Figure 1b). This increasing dominance of type-I-specific fragments was even more pronounced in the pmoA transcripts (Figure 1c). Methylobacter-related tRFs dominated around the oxic–anoxic interface, but also in the upper 0.5 mm. Based on t-RFLP, type II MOB did not transcribe the pmoA gene. This was consistent with cloning and sequencing of pmoA transcripts: type-II-specific sequences were lacking. A considerable fraction of tRFs was not derived from pmoA transcripts but rather from rRNA, as observed previously with pmoA RT-PCR (Krause et al., 2010). These false-positive tRFs became most obvious below the oxic–anoxic interface, which suggested an extremely low number of target molecules in the anoxic soil.
The pmoA gene diversity along the depth profiles was consistent between all four microcosms. On average, the dominating methanotroph groups (type II and type Ia) showed an alternating pattern with a predominance of type Ia around the oxic–anoxic interface (Figure 2). Also the transcript analysis was consistent with the results described for microcosm 1: a lack of type II, a pronounced dominance of type Ia around the oxic–anoxic interface, and a high fraction of false-positive products at depths where no aerobic methane oxidation was expected.
Vertical differentiation of the DNA-based population structure in all four microcosms. Mean relative abundances of type II and type Ia MOB (± s.e., n=4). Type II corresponds to tRF 244; type Ia is the sum of all other fragments. Unlike RT-PCR, the DNA-based analysis did not suffer from unspecific amplification of rRNA genes.
Quantification of transcripts by competitive t-RFLP
Neither the pmoA microarray nor the classic t-RFLP (Horz et al., 2001) allow quantification sensu stricto, even if they are often regarded as semi-quantitative. To quantify copy numbers of the pmoA gene and its transcripts, we combined cPCR or RT-PCR with t-RFLP, respectively. We focused on a fragment affiliated to Methylobacter that showed high relative abundances at the oxic–anoxic interface (tRF 508, Figure 1b). This OTU reached copy numbers up to 3.73 × 108 pmoA genes per gram fresh weight of soil, and a maximum of 9.5 transcripts per genomic pmoA copy at the oxic–anoxic interface (Figure 3). Based on this distribution and further supported by microarray data (Figure 4, see below), we defined three depth zones: surface zone, oxic–anoxic interface and anoxic zone (Table 1). The average number of pmoA transcripts in the highly active oxic–anoxic interface were one order of magnitude higher than in the surface and anoxic zones (Table 1). The highest transcript:gene copy ratios coincided with the area of highest oxygen consumption, as calculated from the oxygen microprofiles (Figure 3).
Competitive t-RFLP analysis of a Methylobacter-related fragment (tRF 508). Copy numbers of the pmoA gene and the pmoA RNA/DNA ratio are shown with Sawitzky–Golay smoothers (n=5). The oxygen consumption rate was calculated from the microprofile shown in Figure 1a using Berg’s program PROFILE ver. 1.0 (Berg et al., 1998), considering the top 2 mm.
Non-metric multidimensional scaling (NMDS; Bray–Curtis dissimilarities; stress=0.0689; R2=0.997; linear fit, R2=0.989) of transcript patterns derived from hybridization to a pmoA microarray; average values from four microcosms. Layers are shown as black symbols. The green, red and blue hulls visualize the three different zones (surface, 0–0.8 mm; oxic–anoxic interface, 0.8–2 mm and anoxic, 2–3 mm), respectively. The differences between the zones are highly significant (analysis of similarity, Bray–Curtis dissimilarity, P<0.001). Red symbols mark the different probes. Probe O_Mmb562, which separated the surface zone from the other two zones, is specific for Methylosarcina. The original data from each microcosm are documented in Supplement 3, and probes considered for the ordination together with the rationale for selecting them are given in Supplement 2.
Microarray analysis of pmoA transcripts
We analyzed the transcript patterns of the four microcosms using non-metric multidimensional scaling (Figure 4). Each of the three depth zones formed a distinct cluster with significant differences to each other (analysis of similarity, Bray–Curtis dissimilarity, P<0.001). Probes used for ordination and a few others yielding obvious spatial patterns are listed in Supplement 1.
The original microarray data from all four microcosms (1–4) provided additional information (Supplement 2). Hybridization signals for type-II-specific probes were observed mostly in the oxic–anoxic interface, but the signals were faint. The only type II probe that gave a stronger signal (P_MM_MsT343) is unspecific, that is, it binds also to many type Ib sequences. This signal coincided with that of the general probes for type Ib (Ib453 and Ib559).
The general probes for type Ia (O_Ia193, O_Ia575) gave strong signals not only at the oxic–anoxic interface but also in the anoxic zone. Furthermore, probe Mb271 gave signals in all three depth zones, but most signals in microcosms 1 and 2 were in the oxic–anoxic interface. In three out of the four microcosms, the upper 0.4 mm was characterized by a strong signal of probe O_Mmb562, which is indicative for Methylosarcina. Also the largely redundant probes Mmb303 and Mmb304 (Supplement 2) gave consistently high signals, which suggested that the activity of Methylosarcina was indeed high in the surface zone (Supplement 3, microcosms 2–4).
To evaluate the potential role of sMMO, we sampled two to three replicate microcosms after 2, 4 and 6 weeks, respectively. All mmoX sequences retrieved could be affiliated with type II MOB of the genera Methylosinus and Methylocystis (Supplement 4). No transcripts were found, whereas a Methycella-specific assay even failed to produce products from DNA suggesting that this genus is missing (Supplement 4).
Discussion
Previous experiments with gradient microcosms have already shown that focusing on the active layer allows processes and interactions to be analyzed in unparalleled detail (Murase and Frenzel, 2007; Krause et al., 2010). While only 3 mm thick, the soil layer in the microcosm was considerably thinner and allowed a stronger focus on the organisms of interest than in many other experiments (Dumont et al., 2011, Siljanen et al., 2011). Even working at a resolution of 1 cm dilutes the active layer with the microbial seed bank in the bulk soil and limits interpretability, regardless if major soil compartments are sampled separately (Eller et al., 2005). Dividing the soil further into 100-μm-thick layers brought an unprecedented resolution that was sufficient not only to analyze the vertical position of different OTUs but also physiological differences within one OTU. While applied here to a laboratory system, this technique can also be adapted to retrieve real-time snapshots from sediment surfaces using freeze cores (Macumber et al., 2011).
Using the pmoA gene as a functional and phylogenetic marker for MOB, we analyzed genes and transcripts at the sub-mm scale along a depth profile from the soil surface down into the anoxic zone. As predicted from oxygen microprofiles, methanotrophic activity was indeed located at the oxic–anoxic interface. Using pmoA transcripts as a proxy for species-specific activity, we correlated the transcript-to-gene ratio of specific methanotroph taxa to methane-driven respiration.
Transcripts of mmoX, a gene encoding for a subunit of sMMO found as a second monooxygenase in some MOB, could not be detected, even if the gene was found (Supplement 4). MOB of the genus Methylocella lack pmoA, but has mmoX (Dedysh, 2009). This genus was previously thought to be acidophilic, but could recently be detected in circum-neutral environments, too (Rahman et al., 2011). However, it was undetectable in the paddy soil under study (Supplement 4). Hence, pmoA is a reliable functional and phylogenetic marker for this microcosm experiment.
Concentration profiles and activity
At the oxic–anoxic interface, oxygen is not only used for the direct mineralization of organic matter but also for the re-oxidation of end products from anaerobic processes (Brune et al., 2000). In freshwater environments, methane is most important and may be the dominant oxidation substrate at the interface. Our microcosms were designed to model this situation, but can in principle be re-configured to focus on other redox processes or to study the interaction between different competing processes.
The 16 oxygen microprofiles measured (4 per microcosm) showed the same characteristics: a near-linear decrease in oxygen concentration down to a depth of about 1.3 mm, followed by a pronounced curvature before the concentration reached zero at about 1.8 mm depth (Figure 1a). The concomitant methane concentrations at the oxic–anoxic interface were most probably in the lower micromolar range (Gilbert and Frenzel, 1998). The methane sensors built for previous work (Rothfuss et al., 1994; Gilbert and Frenzel 1998) do not achieve the sub-mm resolution required for this experiment. While further miniaturization is feasible, the detection limit would decrease proportionally, making a sensitive analysis impossible. Membrane-inlet mass spectrometry (Lloyd et al., 1986; Lloyd et al., 1996; Beckmann et al., 2004) suffers from the same constraints, and the development of a microscale biosensor for methane (Damgaard and Revsbech, 1997) has been discontinued. Hence, interpretation has to rely on oxygen microprofiles alone.
We calculated oxygen consumption using Berg’s program PROFILE ver. 1.0 (Berg et al., 1998) and a sediment diffusion coefficient measured in this paddy soil (Rothfuss and Conrad, 1994; Noll et al., 2005). The algorithm calculates the zone-specific respiration rate using Fick’s second law (Figure 3). Assuming constant porosity, the respiration rate at the surface was modest, followed by an area where no significant respiration took place. High values contributing 89% of the total oxygen respiration were calculated for the zone above the interface (Figure 3). Assuming a stoichiometry of CH4:O2=1:2, the total respiration of 36.6 nmol O2 cm−2 h−1 corresponded to a methane oxidation rate of 18.3 nmol CH4 cm−2h−1.
t-RFLP patterns and quantification
In community profiling studies of MOB, t-RFLP analysis targeting the pmoA gene has a long tradition (Horz et al., 2001; Hoffmann et al., 2002; Mohanty et al., 2007). The results, however, depend on the choice of primer sets. Compared with the 682r reverse primer (Holmes et al., 1995), the reverse primer mb661r (Costello and Lidstrom, 1999; Bourne et al., 2001) covers methanotroph diversity, but not the homologous amoA gene encoding for a subunit of ammonium monooxygenase. Furthermore, primer mb661r seems to be superior for resolving type I diversity (Bourne et al., 2001; Lüke et al., 2010).
When we compared DNA- and RNA-based community profiles, we found striking differences between the extant (DNA-based) and active (RNA-based) populations (Figure 1). The extant population changed consistently with depth in all four microcosms studied (Figure 2). Virtually no transcripts of type II MOB were found, but a Methylobacter-like OTU (tRF 508) was most prominent around the oxic–anoxic interface (Figures 1b and c). Taking transcription as a proxy for activity, this dominance suggested a corresponding localization of Methylobacter-specific methane oxidation. This conclusion, however, depends largely on the high fraction of unspecific tRFs derived from ribosomal RNA. While this large fraction of false-positive tRFs suggested an extremely low content of pmoA mRNA compared with rRNA, it is only a tentative measure. We therefore adapted a cPCR assay (Han and Semrau, 2004) and used it to quantify the most prominent Methylobacter-like OTU (tRF 508; Figure 3).
cPCR has rarely been applied in microbial ecology (Han and Semrau, 2004), but it has certain advantages, in particular if combined with t-RFLP. t-RFLP alone gives only relative values. Provided adequate standards can be constructed, t-RFLP can easily be modified to retrieve truly quantitative data. In addition, RT-cPCR helps in overcoming the variability inherent to the RT step (Freeman et al., 1999). In the particular case of pmoA, RT-cPCR helped in circumventing the problem with unspecific RT-PCR products; the latter had restriction sites that were different from those of the OTU of interest. Some limitations of t-RFLP still apply, for example, a limited phylogenetic resolution and the co-occurrence of particular tRFs in more than one phylotype. Hence, a decent sequence database is a must. However, cPCR combined with t-RFLP has the major advantage that not only quantification but also other tasks such as community analysis (t-RFLP, microarray) and sequencing can be based on the same assays, whereas covering different MOB types by qPCR requires different primer sets (Kolb et al., 2003).
In the anoxic zone below 2 mm, the pmoA copy number was low and may correspond to the initial numbers present in the dry soil when the experiment was started. The slightly higher copy number in the top zone suggested that growth had taken place there at least for some time (Table 1). The maximum transcript:gene ratio coincided with the highest respiration rate, but was localized slightly deeper in the soil than the maximum pmoA copy number (Figure 3). If we consider two pmoCAB operons per cell (Semrau et al., 1995), the Methylobacter-affiliated OTU had up to 18 transcripts per cell (Figure 3). The half-life of pmoA mRNA is unknown. The half-life of other mRNAs may be as short as 30 s, but could also be much longer depending on the environment and the growth state (Arraiano et al., 2010; Steglich et al., 2010). However, we are confident that we preserved the in situ mRNA content by shock freezing the soil with liquid nitrogen when still in the microcosm.
The soil surface in the microcosms was characterized by high oxygen (Figure 1) and low methane concentrations. Before we sampled the soil for molecular analysis, 318 ppmv methane had accumulated in the headspace, which corresponds to 400 nM methane in the pore water near the soil surface. Hence, MOB living in the top layers of the microcosms were exposed to this or a slightly higher methane concentration immediately before shock freezing. However, the Methylobacter-affiliated OTU had less than one transcript per cell (Table 1), which suggested that this situation was rather unfavorable. In the anoxic zone, however, the Methylobacter-affiliated OTU had nearly four transcripts per cell (Table 1). Our microcosm design aimed at producing a planar system varying only with depth. Microelectrode measurements have a high spatial resolution (<0.1 mm, corresponding to ca. 0.53 nl), whereas the molecular data refer to the entire area of the microcosm (28.3 cm2, 0.28 ml per layer). However, the 16 oxygen profiles were so reproducible that we can rule out spatial heterogeneities. Hence, factors other than locally deviating oxygen penetration depth must be responsible for the high transcript:gene ratio below the oxic–anoxic interface. The nitrite-reducing methanotroph, ‘Candidatus Methylomirabilis oxyfera’ is able to generate O2 from NO via dismutase (Ettwig et al., 2010; Strous, 2011). However, we have no indication that Methylobacter possesses this trait, and nitrite was not detectable in another microcosm experiment with the same soil (Krause et al., 2010). Recently developed microelectrodes (Revsbech et al., 2009; Revsbech et al., 2011) have demonstrated nanomolar oxygen concentrations in areas that have been considered anoxic so far. However, the design of our microcosms includes a trap to remove any oxygen that might have diffused into the lower compartment (Murase and Frenzel, 2007). Hence, oxygen may have been present in trace amounts in the methane-rich ‘anoxic’ zone below the oxic–anoxic interface, but diffusive transport of oxygen to MOB must have been negligible, if it occurred at all. Therefore, the mRNA:DNA ratio points to a differentiated physiological status of the cells depending on depth and a surprisingly high number of transcripts in the anoxic zone.
Microarray analyses
Although the microarray analysis leads to essentially the same conclusions as t-RFLP analysis, the higher phylogenetic resolution of the microarrays may allow differentiation down to the species level (Stralis-Pavese et al., 2011). First developed for DNA-based analyses, the microarray has been successfully applied to study pmoA transcripts (Bodrossy et al., 2006; Chen et al., 2007; Krause et al., 2010). Here we used it to refine the transcript analysis (Figure 4, Supplement 3). The microarray design follows a multiple-probes approach. This has significant advantages and may help in detecting groups not yet covered by a specific probe by another, more conserved and general probe. A certain degree of redundancy also helps to exclude false-positive hybridization signals, but may introduce a bias in ordination analyses if a particular phylotype hybridizes with more than one probe. Hence, we used only mutually exclusive probes with a well-defined phylogenetic coverage for ordination analysis. These probes and a few others showing obvious spatial patterns are listed in Supplement 2. The ordination (non-metric multidimensional scaling, Figure 4) gave a clear separation between the surface, oxic–anoxic interface and bottom zones.
The only signal that could be associated with type II MOB was most probably false positive, as the respective probe hybridizes also with different rice paddy clusters (Lüke et al., 2010) belonging to type I (Supplement 3); the general probes for type II gave no signal. Type II MOB have been described as more abundant than type I but as contributing to methane oxidation mainly under high-methane concentrations, whereas type I may thrive if the methane source strength decreases (Henckel et al., 2000). This, however, may be a misconception: even at a high source strength, that is, high-methane production rates, methanotrophic activity shapes counter-gradients, resulting in a steady state with concomitantly low oxygen and methane concentrations at the oxic–anoxic interface. Thus, not methane concentration but rather the energy flow through a population may be the decisive factor (Krause et al., 2012). Type II MOB may occasionally become active in such a situation, but the controlling factor(s) are unknown (Krause et al., 2010). However, extinguishing 97.5% of all microbiota stimulates the exponential growth of type II MOB, which eventually become by far the dominant group (Ho et al., 2011). Hence, thanks to their robust resting stages (Whittenbury et al., 1970a, 1970b), type II MOB may apply a sit-and-wait strategy, taking advantage of changing situations that may be a catastrophe for others.
Type I MOB may have benefited from the rather constant conditions in the microcosms. Probes indicative for Methylobacter gave positive signals in the oxic–anoxic interface and the anoxic zone (Supplement 3), but much less so in the surface zone. This corresponds to the quantification with cPCR/t-RFLP (Figure 3), which revealed a higher mRNA:DNA ratio in the anoxic zone than in the surface zone. The role of type II MOB and Methylobacter is surprisingly similar to that found in a seasonal study on an alpine meadow (Abell et al., 2009), in which type II MOB remained largely unaffected by season and environment but nevertheless represented the dominant MOB. Methylobacter-related MOB, however, were found to be responsible for the majority of methane oxidation.
Probes indicative for Methylosarcina gave significant signals in the surface zone (Supplement 3), which suggested activity at high oxygen concentrations (245–132 μM) and low methane concentrations (ca. 400 nM). This is much higher than the atmospheric methane concentration (1.8 ppmv, corresponding to 2.3 nM), but cultures of and sequences affiliated to Methylosarcina have so far only been retrieved from high-methane environments, such as lake sediments, rice paddies and landfills (Wise et al., 2001; Kalyuzhnaya et al., 2005; Henneberger et al., 2011; Lüke and Frenzel 2011). Apparent Km constants in environments with high source strength are usually >1 μM CH4 (Conrad, 1996), but some cultivated MOB, in particular Methylocystis strains, may grow for an extended period at 120 nM CH4 and less (Knief and Dunfield, 2005). For Methylosarcina-like MOB, however, activity at low methane concentrations has not yet been reported.
Conclusions
As shown before, only a restricted subset of a diverse methanotroph community was active, and most activity was confined to a zone 0.67-mm thick. However, our experiment showed how different OTUs within a single guild can share the same microenvironment, thereby exploiting different niches. We hypothesized that activity would be highest immediately at the oxic–anoxic interface separating MOB according to substrate availability. This was indeed the case, with a Methylobacter-affiliated OTU and dominating overall methane oxidation located at the oxic–anoxic interface. This OTU seems to be well adapted to the oxic–anoxic interface, where oxygen and, presumably, methane concentrations are in the micromolar range. This OTU was not active at the surface with its high oxygen but only nanomolar methane concentrations. In contrast, transcripts of a Methylosarcina-affiliated OTU were associated with this surface layer, which suggested an adaptation to oligotrophic conditions.
References
Abell GCJ, Stralis-Pavese N, Sessitsch A, Bodrossy L . (2009). Grazing affects methanotroph activity and diversity in an alpine meadow soil. Environ Microbiol Rep 1: 457–465.
Amann RI, Ludwig W, Schleifer KH . (1995). Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Mol Biol Rev 59: 143–169.
Arraiano CM, Andrade JM, Domingues S, Guinote IB, Malecki M, Matos RG et al (2010). The critical role of RNA processing and degradation in the control of gene expression. FEMS Microbiol Rev 34: 883–923.
Beckmann M, Sheppard SK, Lloyd D . (2004). Mass spectrometric monitoring of gas dynamics in peat monoliths: effects of temperature and diurnal cycles on emissions. Atmos Environ 38: 6907–6913.
Berg P, Risgaard-Petersen N, Rysgaard S . (1998). Interpretation of measured concentration profiles in sediment pore water. Limnol Oceanogr 43: 1500–1510.
Bodelier PLE, Hahn AP, Arth I, Frenzel P . (2000a). Effects of ammonium-based fertilisation on microbial processes involved in methane emission from soils planted with rice. Biogeochemistry 51: 225–257.
Bodelier PLE, Roslev P, Henckel T, Frenzel P . (2000b). Stimulation by ammonium-based fertilizers of methane oxidation in soil around rice roots. Nature 403: 421–424.
Bodelier PLE, Sorrell BK, Drake HL, Küsel K, Hurek T, Reinhold-Hurek B et al (2006). Ecological aspects of microbes and microbial communities inhabiting the rhizosphere of wetland plants. In: Verhoeven JTA, Beltman B, Bobbink R, Whigham DF (eds). Wetlands and Natural Resource Management (Ecological Studies) Vol. 190. Springer: Berlin-Heidelberg, pp 205–238.
Bodrossy L, Stralis-Pavese N, Konrad-Koszler M, Weilharter A, Reichenauer TG, Schofer D et al (2006). mRNA-based parallel detection of active methanotroph populations by use of a diagnostic microarray. Appl Environ Microbiol 72: 1672–1676.
Bodrossy L, Stralis-Pavese N, Murrell JC, Radajewski S, Weilharter A, Sessitsch A . (2003). Development and validation of a diagnostic microbial microarray for methanotrophs. Environ Microbiol 5: 566–582.
Bosse U, Frenzel P . (1997). Activity and distribution of methane-oxidizing bacteria in flooded rice soil microcosms and in rice plants (Oryza sativa). Appl Environ Microbiol 63: 1199–1207.
Bosse U, Frenzel P, Conrad R . (1993). Inhibition of methane oxidation by ammonium in the surface layer of a littoral sediment. FEMS Microbiol Ecol 13: 123–134.
Bourne DG, McDonald IR, Murrell JC . (2001). Comparison of pmoA PCR primer sets as tools for investigating methanotroph diversity in three Danish soils. Appl Environ Microbiol 67: 3802–3809.
Brune A, Frenzel P, Cypionka H . (2000). Life at the oxic-anoxic interface: microbial activities and adaptations. FEMS Microbiol Rev 24: 691–710.
Calhoun A, King GM . (1997). Regulation of root-associated methanotrophy by oxygen availability in the rhizosphere of two aquatic macrophytes. Appl Environ Microbiol 63: 3051–3058.
Chen Y, Dumont MG, Cebron A, Murrell JC . (2007). Identification of active methanotrophs in a landfill cover soil through detection of expression of 16S rRNA and functional genes. Environ Microbiol 9: 2855–2869.
Coleman NV, Le NB, Ly MA, Ogawa HE, McCarl V, Wilson NL et al (2012). Hydrocarbon monooxygenase in Mycobacterium: recombinant expression of a member of the ammonia monooxygenase superfamily. ISME J 6: 171–182.
Conrad R . (1996). Soil microorganisms as controllers of atmospheric trace gases (H2, CO, CH4, OCS, N2O, and NO). Microbiol Rev 60: 609–640.
Conrad R . (2009). The global methane cycle: Recent advances in understanding the microbial processes involved. Environ Microbiol Rep 1: 285–292.
Conrad R, Rothfuss F . (1991). Methane oxidation in the soil surface layer of a flooded rice field and the effect of ammonium. Biol Fert Soils 12: 28–32.
Costello AM, Lidstrom ME . (1999). Molecular characterization of functional and phylogenetic genes from natural populations of methanotrophs in lake sediments. Appl Environ Microbiol 65: 5066–5074.
Damgaard LA, Revsbech NP . (1997). A microscale biosensor for methane containing methanotrophic bacteria and an internal oxygen reservoir. Anal Chem 69: 2262–2267.
Dedysh SN . (2009). Exploring methanotroph diversity in acidic northern wetlands: molecular and cultivation-based studies. Microbiology 78: 655–669.
Dedysh SN, Liesack W, Khmelenina VN, Suzina NE, Trotsenko YA, Semrau JD et al (2000). Methylocella palustris gen. nov., sp. nov., a new methane-oxidizing acidophilic bacterium from peat bogs, representing a novel subtype of serine-pathway methanotrophs. Int J Syst Evol Microbiol 50: 955–969.
Degelmann DM, Borken W, Drake HL, Kolb S . (2010). Different atmospheric methane-oxidizing communities in European beech and Norway spruce soils. Appl Environ Microbiol 76: 3228–3235.
Dumont MG, Murrell JC . (2005). Community-level analysis: key genes of aerobic methane oxidation. Environ Microbiol 397: 413–427.
Dumont MG, Pommerenke B, Casper P, Conrad R . (2011). DNA-, rRNA- and mRNA-based stable isotope probing of aerobic methanotrophs in lake sediment. Environ Microbiol 13: 1153–1167.
Dunbar J, Ticknor LO, Kuske CR . (2001). Phylogenetic specificity and reproducibility and new method for analysis of terminal restriction fragment profiles of 16S rRNA genes from bacterial communities. Appl Environ Microbiol 67: 190–197.
Dunfield PF, Yuryev A, Senin P, Smirnova AV, Stott MB, Hou S et al (2007). Methane oxidation by an extremely acidophilic bacterium of the phylum. Verrucomicrobia. Nature 450: 879–882.
Eller G, Frenzel P . (2001). Changes in activity and community structure of methane oxidising bacteria over the growth period of rice. Appl Environ Microbiol 67: 2395–2403.
Eller G, Krüger M, Frenzel P . (2005). Comparing field and microcosm experiments: a case study on methano- and methylotrophic bacteria in paddy soil. FEMS Microbiol Ecol 51: 279–291.
Ettwig KF, Butler MK, Le Paslier D, Pelletier E, Mangenot S, Kuypers MMM et al (2010). Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464: 543–548.
Freeman WM, Walker SJ, Vrana KE . (1999). Quantitative RT-PCR: pitfalls and potential. Biotechniques 26: 112–125.
Frenzel P, Rothfuss F, Conrad R . (1992). Oxygen profiles and methane turnover in a flooded rice microcosm. Biol Fert Soils 14: 84–89.
Frenzel P, Thebrath B, Conrad R . (1990). Oxidation of methane in the oxic surface layer of a deep lake sediment (Lake Constance). FEMS Microbiol Ecol 73: 149–158.
Gilbert B, Frenzel P . (1998). Rice roots and CH4 oxidation: the activity of bacteria, their distribution and the microenvironment. Soil Biol Biochem 30: 1903–1916.
Graham DW, Chaudhary JA, Hanson RS, Arnold RG . (1993). Factors affecting competition between type-I and type-II methanotrophs in two-organism, continuous-flow reactors. Microbiol Ecol 25: 1–17.
Han JI, Semrau JD . (2004). Quantification of gene expression in methanotrophs by competitive reverse transcription-polymerase chain reaction. Environ Microbiol 6: 388–399.
Henckel T, Roslev P, Conrad R . (2000). Effects of O2 and CH4 on presence and activity of the indigenous methanotrophic community in rice field soil. Environ Microbiol 2: 666–679.
Henneberger R, Lüke C, Mosberger L, Schroth MH . (2011). Structure and function of methanotrophic communities in a landfill-cover soil. FEMS Microbiol Ecol e-pub ahead of print 15 December 2011; doi:10.1111/j.1574-6941.2011.01278.x.
Ho A, Lüke C, Frenzel P . (2011). Recovery of methanotrophs from disturbance: population dynamics, evenness, and functioning. ISME J 5: 750–758.
Hoffmann T, Horz HP, Kemnitz D, Conrad R . (2002). Diversity of the particulate methane monooxygenase gene in methanotrophic samples from different rice field soils in China and the Philippines. Syst Appl Microbiol 25: 267–274.
Holmes AJ, Costello A, Lidstrom ME, Murrell JC . (1995). Evidence that particulate methane monooxygenase and ammonia monooxygenase may be evolutionarily related. FEMS Microbiol Lett 132: 203–208.
Horz HP, Yimga MT, Liesack W . (2001). Detection of methanotroph diversity on roots of submerged rice plants by molecular retrieval of pmoA, mmoX, mxaF, and 16S rRNA and ribosomal DNA, including pmoA-based terminal restriction fragment length polymorphism profiling. Appl Environ Microbiol 67: 4177–4185.
Hutchens E, Radajewski S, Dumont MG, McDonald IR, Murrell JC . (2004). Analysis of methanotrophic bacteria in Movile Cave by stable isotope probing. Environ Microbiol 6: 111–120.
Intergovernmental Panel on Climate Change. (2007) Climate Change 2007: The Physical Science Basis. Summary for Policymakers. Contribution of Working Group I to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change. Summary for Policymakers formally approved at the 10th Session of Working Group I of the IPCC. IPCC Secretariat: Geneva.
Islam T, Jensen S, Reigstad LJ, Larsen O, Birkeland NK . (2008). Methane oxidation at 55 °C and pH 2 by a thermoacidophilic bacterium belonging to the Verrucomicrobia phylum. Proc Natl Acad Sci USA 105: 300–304.
Kalyuzhnaya MG, Stolyar SM, Auman AJ, Lara JC, Lidstrom ME, Chistoserdova L . (2005). Methylosarcina lacus sp. nov., a methanotroph from Lake Washington, Seattle, USA, and emended description of the genus Methylosarcina. Int J Syst Evol Microbiol 55: 2345–2350.
Knief C, Dunfield PF . (2005). Response and adaptation of different methanotrophic bacteria to low methane mixing ratios. Environ Microbiol 7: 1307–1131.
Kolb S, Knief C, Stubner S, Conrad R . (2003). Quantitative detection of methanotrophs in soil by novel pmoA-targeted real-time PCR assays. Appl Environ Microbiol 69: 2423–2429.
Krause S, Lüke C, Frenzel P . (2009). Spatial heterogeneity of methanotrophs: a geostatistical analysis of pmoA-based T-RFLP patterns in a paddy soil. Environ Microbiol Rep 1: 393–397.
Krause S, Lüke C, Frenzel P . (2010). Succession of methanotrophs in oxygen-methane counter-gradients of flooded rice paddies. ISME J 4: 1603–1607.
Krause S, Lüke C, Frenzel P . (2012). Methane source strength and energy flow shape methanotrophic communities in oxygen-methane counter-gradients. Environ Microbiol Rep 4: 203–208.
Lloyd D, Davies KJP, Boddy L . (1986). Mass spectrometry as an ecological tool for in situ measurement of dissolved gases in sediment systems. FEMS Microbiol Ecol 38: 11–17.
Lloyd D, Thomas K, Price D, O’Neil B, Oliver K, Williams TN . (1996). A membrane-inlet mass spectrometer miniprobe for the direct simultaneous measurement of multiple gas species with spatial resolution of 1 mm. J Microbiol Meth 25: 145–151.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Lueders T, Manefield M, Friedrich MW . (2004). Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ Microbiol 6: 73–78.
Lüke C, Bodrossy L, Lupotto E, Frenzel P . (2011). Methanotrophic bacteria associated to rice roots: the cultivar effect assessed by T-RFLP and microarray analysis. Environ Microbiol Rep 3: 518–525.
Lüke C, Frenzel P . (2011). Potential of pmoA amplicon pyrosequencing for methanotroph diversity studies. Appl Environ Microbiol 77: 6305–6309.
Lüke C, Krause S, Cavigioli S, Greppi D, Lupotto E, Frenzel P . (2010). Biogeography of wetland rice methanotrophs. Environ Microbiol 12: 862–872.
Macumber AL, Patterson RT, Neville LA, Falck H . (2011). A sledge microtome for high resolution subsampling of freeze cores. J Paleolimnol 45: 307–310.
McDonald IR, Bodrossy L, Chen Y, Murrell JC . (2008). Molecular ecology techniques for the study of aerobic methanotrophs. Appl Environ Microbiol 74: 1305–1315.
McDonald IR, Murrell JC . (1997). The particulate methane monooxygenase gene pmoA and its use as a functional gene probe for methanotrophs. FEMS Microbiol Lett 156: 205–210.
Mohanty SR, Bodelier PLE, Conrad R . (2007). Effect of temperature on composition of the methanotrophic community in rice field and forest soil. FEMS Microbiol Ecol 62: 24–31.
Murase J, Frenzel P . (2007). A methane-driven microbial food web in a wetland rice soil. Environ Microbiol 9: 3025–3034.
Murase J, Frenzel P . (2008). Selective grazing of methanotrophs by protozoa in a rice field soil. FEMS Microbiol Ecol 65: 408–414.
Murase J, Noll M, Frenzel P . (2006). Impact of protists on activity and structure of the bacterial community in a rice field soil. Appl Environ Microbiol 72: 5436–5444.
Noll M, Frenzel P, Conrad R . (2008). Selective stimulation of type I methanotrophs in a rice paddy soil by urea fertilization revealed by RNA-based stable isotope probing. FEMS Microbiol Ecol 65: 125–132.
Noll M, Matthies D, Frenzel P, Derakshani M, Liesack W . (2005). Succession of bacterial community structure and diversity in a paddy soil oxygen gradient. Environ Microbiol 7: 382–395.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB et al (2011). vegan: Community Ecology Package 2.2-0http://vegan.r-forge.r-project.org/.
Op den Camp HJL, Islam TB, Stott MR, Harhangi H, Hynes A, Schouten S et al (2009). Environmental, genomic and taxonomic perspectives on methanotrophic Verrucomicrobia. Environ Microbiol Rep 1: 293–306.
Pol A, Heijmans K, Harhangi HR, Tedesco D, Jetten MSM, Op den Camp HJM . (2007). Methanotrophy below pH1 by a new Verrucomicrobia species. Nature 450: 874–878.
R Development Core Team. (2011). R: A language and environment for statistical computing 2.13.1, http://cran.r-project.org/.
Rahman MT, Crombie A, Chen Y, Stralis-Pavese N, Bodrossy L, Meir P et al (2011). Environmental distribution and abundance of the facultative methanotroph Methylocella. ISME J 5: 1061–1066.
Reeburgh WS, Whalen SC, Alperin MJ . (1993). The role of methylotrophy in the global methane budget. In: Murrell JC, Kelly DP (eds). Microbial Growth on C1 Compounds. Intercept: Andover, pp 1–14.
Revsbech NP, Larsen LH, Gundersen J, Dalsgaard T, Ulloa O, Thamdrup B . (2009). Determination of ultra-low oxygen concentrations in oxygen minimum zones by the STOX sensor. Limnol Oceanogr Meth 7: 371–381.
Revsbech NP, Thamdrup B, Dalsgaard T, Canfield DE . (2011). Construction of STOX oxygen sensors and their tpplication for determination of O2 concentrations in oxygen minimum zones. Methods Enzymol 486: 325–341.
Rothfuss F, Conrad R . (1994). Development of a gas-diffusion probe for the determination of methane concentrations and diffusion characteristics in flooded paddy soil. FEMS Microbiol Ecol 14: 307–318.
Rothfuss F, Frenzel P, Conrad R . (1994). Gas diffusion probe for measurement of CH4 gradients. In: Stal LJ, Caumette P (eds). Microbial Mats: Structure, Development and Environmental Significance. Springer: Berlin, pp 167–172.
Rudd JWM, Furutani A, Flett RJ, Hamilton RD . (1976). Factors controlling methane oxidation in shield lakes: The role of nitrogen fixation and oxygen concentration. Limnol Oceanogr 21: 357–364.
Sayavedra-Soto LA, Hamamura N, Liu C-W, Kimbrel JA, Chang JH, Arp DJ . (2011). The membrane-associated monooxygenase in the butane-oxidizing Gram-positive bacterium Nocardioides sp. strain CF8 is a novel member of the AMO/PMO family. Environ Microbiol Rep 3: 390–396.
Semrau JD, Chistoserdov A, Lebron J, Costello A, Davagnino J, Kenna E et al (1995). Particulate methane monooxygenase genes in methanotrophs. J Bact 177: 3071–3079.
Shrestha M, Abraham WR, Shrestha PM, Noll M, Conrad R . (2008). Activity and composition of methanotrophic bacterial communities in planted rice soil studied by flux measurements, analyses of pmoA gene and stable isotope probing of phospholipid fatty acids. Environ Microbiol 10: 400–412.
Siljanen HMP, Saari A, Krause S, Lensu A, Abell GCJ, Bodrossy L et al (2011). Hydrology is reflected in the functioning and community composition of methanotrophs in the littoral wetland of a boreal lake. FEMS Microbiol Ecol 75: 430–445.
Sorrell BK, Downes MT, Stanger CL . (2002). Methanotrophic bacteria and their activity on submerged aquatic macrophytes. Aquat Bot 72: 107–119.
Steglich C, Lindell D, Futschik M, Rector T, Steen R, Chisholm SW . (2010). Short RNA half-lives in the slow-growing marine cyanobacterium Prochlorococcus. Genome Biol 11: R54.
Stoecker K, Bendinger B, Schoning B, Nielsen PH, Nielsen JL, Baranyi C et al (2006). Cohn’s Crenothrix is a filamentous methane oxidizer with an unusual methane monooxygenase. Proc Natl Acad Sci USA 103: 2363–2367.
Stralis-Pavese N, Abell GCJ, Sessitsch A, Bodrossy L . (2011). Analysis of methanotroph community composition using a pmoA-based microbial diagnostic microarray. Nat Protoc 6: 609–624.
Stralis-Pavese N, Sessitsch A, Weilharter A, Reichenauer T, Riesing J, Csontos J et al (2004). Optimization of diagnostic microarray for application in analysing landfill methanotroph communities under different plant covers. Environ Microbiol 6: 347–363.
Strous M . (2011). Beyond denitrification: alternative routes to dinitrogen. In: Moir JWB (ed.). Nitrogen Cycling in Bacteria: Molecular Analysis. Caister Academic Press: Norfolk, UK, pp 123–133.
Trotsenko YA, Murrell JC . (2008). Metabolic aspects of aerobic obligate methanotrophy. Adv Appl Microbiol 63: 183–229.
Vishwakarma P, Dumont MG, Bodrossy L, Stralis-Pavese N, Murrell JC, Dubey SK . (2009). Ecological and molecular analyses of the rhizospheric methanotroph community in tropical rice soil: effect of crop phenology and land-use history. Curr Sci 96: 1082–1089.
Vorobev AV, Baani M, Doronina NV, Brady AL, Liesack W, Dunfield PF et al (2011). Methyloferula stellata gen. nov., sp. nov., an acidophilic, obligately methanotrophic bacterium that possesses only a soluble methane monooxygenase. Int J Syst Evol Microbiol 61: 2456–2463.
Whittenbury R, Davies SL, Davey JF . (1970a). Exospores and cysts formed by methane-utilizing bacteria. J Gen Microbiol 61: 219–226.
Whittenbury R, Philips KC, Wilkinson JF . (1970b). Enrichment, isolation, and some properties of methane-utilizing bacteria. J Gen Microbiol 61: 205–218.
Wise MG, McArthur JV, Shimkets LJ . (2001). Methylosarcina fibrata gen. nov., sp. nov. and Methylosarcina quisquiliarum sp. nov., novel type I methanotrophs. Int J Syst Evol Microbiol 51: 611–621.
Acknowledgements
We thank Levente Bodrossy (Department of Bioresources, Austrian Research Centre Seibersdorf, Austria, and CSIRO Marine and Atmospheric Research, Hobart, Tasmania, Australia) for help and advice with the pmoA microarray. Elisabetta Lupotto, (CRA Agricultural Research Council, Rice Research Unit, Vercelli, Italy) helped us with soil sampling. Alexandra Hahn, MPI Marburg, helped with the microcosms for the mmoX experiment. This work, as part of the European Science Foundation EUROCORES Programme EuroEEFG, was supported from funds by the Deutsche Forschungsgemeinschaft to PF.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Reim, A., Lüke, C., Krause, S. et al. One millimetre makes the difference: high-resolution analysis of methane-oxidizing bacteria and their specific activity at the oxic–anoxic interface in a flooded paddy soil. ISME J 6, 2128–2139 (2012). https://doi.org/10.1038/ismej.2012.57
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.57
Keywords
This article is cited by
-
Interkingdom interaction: the soil isopod Porcellio scaber stimulates the methane-driven bacterial and fungal interaction
ISME Communications (2023)
-
The methane-driven interaction network in terrestrial methane hotspots
Environmental Microbiome (2022)
-
Microbial community structure is stratified at the millimeter-scale across the soil–water interface
ISME Communications (2022)
-
Aerobic Methanotrophy and Co-occurrence Networks of a Tropical Rainforest and Oil Palm Plantations in Malaysia
Microbial Ecology (2022)
-
Methane-dependent selenate reduction by a bacterial consortium
The ISME Journal (2021)