Abstract
Hypertension (HTN) is one of the most common emerging disease in developing countries. It alters endothelial cell structure and function, resulting in several diseases, such as cardiovascular disease, peripheral vasculopathy, cerebrovascular disease and nephropathy. Although much progress has been made in researching HTN in recent years, early diagnosis and treatment of HTN are not yet satisfactory, and progression control/treatment is still poor. MicroRNAs are well-known regulators of the physiological and developmental processes of HTN. Our results revealed that miR-510 was upregulated in blood samples from HTN patients, whereas no significant differences were observed in the control samples. Methylation analyses corroborated the miR-510 upregulation in patient samples. These results suggested that miR-510 can be used as a novel biomarker for diagnosis and as a new therapeutic target for HTN.
Similar content being viewed by others
Introduction
Hypertension (HTN) is one of the most common noncommunicable diseases worldwide and is becoming increasingly common in developing countries, such as India. It results in a substantial public health burden on cardiovascular health and health-care systems in most countries.1 HTN is responsible for 57% of all stroke deaths and 24% of all coronary heart disease deaths in India.2 Interestingly, the World Health Organization stated that HTN is one of the most significant causes of premature death worldwide.3 HTN has caused ~7.5 million deaths, which accounts for 12.8% of the global mortality, and 57 million disability-adjusted life years or 3.7% of total disability-adjusted life years have been attributed to HTN.4 HTN can be inhibited by lifestyle modification and medical involvement. Moreover, screening and early management of HTN through periodic surveillance will decrease disease progression.5
The molecular mechanisms underlying the genetic predisposition to HTN have been extensively researched. The protein-coding regions of the genome account for <2% of the entire human DNA, suggesting that genetic mechanisms besides protein-coding genes are likely to contribute to HTN regulation.6
MicroRNAs (miRNAs) or circulating miRNAs are important post-transcriptional regulators of gene expression in many types of diseases and they are associated with various diseases.7, 8 Recent studies have shown that miR-9 and miR-126 are strongly related to essential HTN in humans, suggesting that they may be potential biomarkers and candidate therapeutic targets for essential HTN.9 A recent study showed that miR-510 is involved in cell proliferation, migration and apoptosis,10 and it is a major regulatory circuit in cells.11 Interestingly, another study revealed that miR-510 directly binds to the 3′UTR of peroxiredoxin 1 and blocks its protein expression, thereby suppressing migration of human breast cancer cells.12
Several studies have examined miR-510 in other diseases, such as cancer, but there is no evidence showing that miR-510 is involved in HTN. MiRNA silencing via methylation of CpG islands may be an important mechanism for disease progression.13 Although DNA methylation is an important mechanism for miRNA upregulation, this has not been explored in HTN. Thus, our study predominantly focused on epigenetic regulation and expression of circulating miR-510 in HTN, which could be useful for elucidating the molecular mechanism of miR-510 in HTN and its role in disease progression. Circulating miRNAs are known to be an important factor involved in the initiation and progression of HTN. On the basis of the above information, in the present study, we studied the mechanism of circulating miR-510 in blood samples from HTN patients, and we found that miR-510 was a diagnostic and prognostic marker for HTN.
Materials and Methods
Clinical samples
A total of 428 genetically unrelated patients were selected and divided into two groups. Group I consisted of 208 essential hypertensive patients and Group II consisted of 220 healthy volunteers (age- and sex-matched controls) who were randomly selected. Data collected from each subject included age, height, weight, body mass index and family history. HTN was defined as a systolic blood pressure >140 mm Hg and sustained diastolic blood pressure >90 mm Hg. Blood pressure was measured in subjects who were seated and resting for 5 min, and was taken twice to calculate the mean systolic blood pressure and diastolic blood pressure. No patients were receiving antihypertensive drugs, and patients with secondary HTN, renal, liver and cardiac abnormalities were excluded from the present study.
Venous blood samples (5 ml) were collected from each subject for the analyses. A portion of each blood sample was stored in an EDTA tube (Becton Dickson, New Jersy, NJ, USA) for genomic DNA extraction, and the remaining sample was left to clot to obtain serum and stored at −20 °C for further analysis. Serum samples were analyzed by a HUMASTAR-300 autoanalyser (HUMAN Gesellschaft für Biochemica und Diagnostica mbH, Wiesbaden, Germany) using kits supplied by Human Diagnostics GmbH (HUMAN Gesellschaft für Biochemica und Diagnostica mbH) to determine the levels of fasting blood glucose, serum urea, creatinine, total cholesterol, high-density lipoprotein cholesterol and triglycerides. Low-density lipoprotein cholesterol was calculated by the Friedewald formula. The Third Report of the National Cholesterol Education Program guidelines (NCEP report 2001) were followed for the classification of lipid profiles. Serum sodium, potassium and chloride levels were determined using a HUMLYTE electrolyte analyzer (ion-selective electrode method, HUMAN Gesellschaft für Biochemica und Diagnostica mbH).
RNA isolation and quantification of miRNAs
Total RNA was extracted using a RNeasy Mini Kit (Qiagen, New Delhi, India), and the expression pattern of miR-510 was analyzed as described by Balcells et al.,14 where poly (A) tailing of the miRNA was followed by reverse transcription with a tagged poly (T) primer. Quantification of miRNAs was performed by PCR using the following primers: forward: 5′-AGTATGGCCCGGCCGTGA-3′; reverse: 5′-AGGTCCATTTTTTTTTTTTTTTCCT-3′. miR-510 levels were normalized to 18s RNA levels using the 2ΔΔCt model.
Bisulfite conversion and methylation analysis
DNA and bisulfite-treated DNA were extracted from HTN blood samples and normal blood samples using a QIAamp Mini Kit (Qiagen) as described previously.15 Briefly, DNA was treated with bisulfite using the EZ DNA Methylation Gold Kit (Zymo Research, Chennai, India) following the manufacturer’s instructions.
DNA methylation analysis
Methylation status of the miR-510 promoter was detected using the methods described in the study by Vogelsang et al.,15 which consisted of a two-stage methylation-specific PCR (MSP) or nested MSP that involved a first-round amplification using primers unbiased toward the methylation status of the analyzed region followed by conventional MSP. During the first round, the amplification primers MSP-miR-510-fw (5′-ACA ATCACT CTT TCC ATA TAC-3′) and MSP-miR-510-rv (5′-GCC CAG TCA CAT CTA GTT-3′) were used to amplify a 562 bp fragment including a portion of CpG-rich promoter region from bisulfite-treated DNA templates. We used the amplification protocol and MSP analysis described in the study by Vogelsang et al.15 Specific primer sets to selectively amplify unmethylated and methylated alleles were designed according to previous references as follows:16 miR-510-U-fw 5′-ATC TCG TTT TTA TTT AAA CGT-3′, miR-510-U-rv 5′-AGT GAC TCA CCCTAT TCT CTG ATG-3′ (unmethylated amplicon size: 144 bp) and miR-510-M-fw 5′-ACA AAA GCC GCC TAT CCC TGT CCC CTA-3′, miR-510-M-rv 5′-TTA GAA TGG ACC TCT AAACTC AAA-3′ (methylated amplicon size: 147 bp).
Statistical methods
All clinical data are expressed as the mean±s.d. All statistical analyses were carried out using SPSS software version 15.0 for Microsoft Windows (SPSS software, IBM, India). Continuous variables were compared between the cases and control groups using two-tailed Student’s t-tests. P<0.05 was considered statistically significant.
Results and Discussion
In general, miRNAs control a wide range of biological functions, such as proliferation, differentiation, apoptosis and other gene functions.17, 18, 19 Recent reports have shown that miRNAs can act as a repressor or inducer, and have key roles in the initiation and progression of HTN.9 MiRNAs have been detected in body fluids, such as serum, plasma and urine, and can be readily used as noninvasive biomarkers.20 Our study group included 20 essential HTN patients and 20 controls. Table 1 shows the baseline demographic, clinical and laboratory data of the study population. Among the patients, 12 (58.2%) were men and 8 (41.8%) were women. There was no significant difference between patients and controls with respect to age and body mass index (P>0.005). Salt-sensitive-hydroxylase (SBH) and dopamine beta-hydroxylase (DBH) were higher in patients compared with controls (P=<0.005).
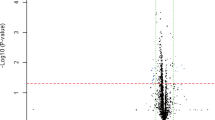
Serum total cholesterol, HDL, LDL, very low density lipoprotein (VLDL) and triglyceride levels were significantly higher in patients than the controls (P=<0.005). Blood glucose and serum sodium levels also showed differences between patients and controls (P=<0.005). There were no significant differences between patients and controls with respect to serum creatinine, urea, potassium and chloride (P=>0.005). To elucidate the mechanisms underlying abnormal miRNA expression in HTN, an increasing number of studies have investigated how miRNAs are regulated, and it is now widely accepted that miRNAs undergo the same regulatory mechanisms as any other classical protein-coding genes, including epigenetic regulation. We hypothesized that epigenetic modification of genes encoding the miRNAs is a key factor in inducing HTN, and we assessed this hypothesis in detail. To test this hypothesis, we examined the expression of mature miR-510 in blood samples from HTN patients and control samples. Our results suggested that the miR-510 level in blood samples from HTN patients was significantly higher than that in the normal control samples (Figure 1).
The expression of miR-510 in patient blood samples suggested that the upregulation of miR-510 is relevant to the initiation and development of HTN. On the basis of the results from the gene expression study, we assessed the molecular mechanisms underlying the upregulation of miR-510. A recent study by Formosa et al.13 reported the role of the methylation status of CpG islands upstream of a subset of miRNAs using MSP. Additional reports also have demonstrated that the epigenetic regulation of miRNAs is a widespread phenomenon, as shown, for example, for miR-9-1, miR-107, miR-127, miR-193a, miR-137, miR-342, miR-203, miR-34b/c and miR-1.21, 22
We used MSP or nested MSP to investigate the status of miR-510 promoter methylation in HTN blood and normal blood samples. Our hypermethylation study revealed that the miR-510 promoter was unmethylated in HTN blood samples, but in the control sample, it was found to be methylated. Representative examples of MSP analyses are shown in Figure 2. The methylation frequency for the miR-510 promoter was compared between the HTN blood and normal blood samples (Figure 3). Overall, we found that the methylation frequency was significantly different between these samples (81.0 vs. 65.0%, P=0.005). Our data suggested that hypermethylation is corroborated with miR-510 expression in blood samples. In this study, we found that miR-510 was unmethylated in hypertensive patient blood samples, whereas in control samples, their expression levels significantly correlated with methylation status of the miR-510 promoter. Moreover, this is the first report to show the methylation of the promoter of circulating miR-510, and further investigations are needed to confirm that miR-510 is a potential circulatory miRNA for HTN.
References
Srinath Reddy K, Shah B, Varghese C, Ramadoss A . Responding to the threat of chronic diseases in India. Lancet 2005; 366: 1744–1749.
Gupta R . Trends in hypertension epidemiology in India. J Hum Hypertens 2004; 18: 73–78.
Anchala R, Pant H, Prabhakaran D, Franco OH . 'Decision support system (DSS) for prevention of cardiovascular disease (CVD) among hypertensive (HTN) patients in Andhra Pradesh, India'–-a cluster randomised community intervention trial. BMC Public Health 2012; 12: 393.
Papathanasiou G, Zerva E, Zacharis I, Papandreou M, Papageorgiou E, Tzima C, Georgakopoulos D, Evangelou A . Association of high blood pressure with body mass index, smoking and physical activity in healthy young adults. Open Cardiovasc Med J 2015; 9: 5–17.
Midha T, Krishna V, Shukla R, Katiyar P, Kaur S, Martolia DS, Pandey U, Rao YK . Correlation between hypertension and hyperglycemia among young adults in India. World J Clin Cases 2015; 3: 171–179.
Marques FZ, Booth SA, Charchar FJ . The emerging role of non-coding RNA in essential hypertension and blood pressure regulation. J Hum Hypertens 2015; 29: 459–467.
Sekar D, Saravanan S, Karikalan K, Thirugnanasambantham K, Lalitha P, Islam VI . Role of microRNA 21 in mesenchymal stem cell (MSC) differentiation: a powerful biomarker in MSCs derived cells. Curr Pharm Biotechnol 2015; 16: 43–48.
Sekar D, Hairul Islam VI, Thirugnanasambantham K, Saravanan S . Relevance of miR-21 in HIV and non-HIV-related lymphomas. Tumour Biol 2014; 35: 8387–8393.
Kontaraki JE, Marketou ME, Zacharis EA, Parthenakis FI, Vardas PE . MicroRNA-9 and microRNA-126 expression levels in patients with essential hypertension: potential markers of target-organ damage. J Am Soc Hypertens 2014; 8: 368–375.
Chen D, Li Y, Yu Z, Li Y, Su Z, Ni L, Yang S, Gui Y, Lai Y . Downregulated microRNA-510-5p acts as a tumor suppressor in renal cell carcinoma. Mol Med Rep 2015; 12: 3061–3066.
Gaj P, Zagozdzon R . In silico analysis of microRNA-510 as a potential oncomir in human breast cancer. Breast Cancer Res 2014; 16: 403.
Guo QJ, Mills JN, Bandurraga SG, Nogueira LM, Mason NJ, Camp ER, Larue AC, Turner DP, Findlay VJ . MicroRNA-510 promotes cell and tumor growth by targeting peroxiredoxin1 in breast cancer. Breast Cancer Res 2013; 15: R70.
Formosa A, Lena AM, Markert EK, Cortelli S, Miano R, Mauriello A, Croce N, Vandesompele J, Mestdagh P, Finazzi-Agrò E, Levine AJ, Melino G, Bernardini S, Candi E . DNA methylation silences miR-132 in prostate cancer. Oncogene 2013; 32: 127–134.
Balcells I, Cirera S, Busk PK . Specific and sensitive quantitative RT-PCR of miRNAs with DNA primers. BMC Biotechnol 2011; 11: 70.
Vogelsang M, Paccez JD, Schafer G, Dzobo K, Zerbini LF, Parker MI . Aberrant methylation of the MSH3 promoter and distal enhancer in esophageal cancer patients exposed to first-hand tobacco smoke. J Cancer Res Clin Oncol 2014; 140: 1825–1833.
Kim HG, Lee S, Kim DY, Ryu SY, Joo JK, Kim JC, Lee KH, Lee JH . Aberrant methylation of DNA mismatch repair genes in elderly patients with sporadic gastric carcinoma: a comparison with younger patients. J Surg Oncol 2010; 101: 28–35.
Chan JA, Krichevsky AM, Kosik KS . MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 2005; 65: 6029–6033.
Feng B, Cao Y, Chen S, Ruiz M, Chakrabarti S . Reprint of: miRNA-1 regulates endothelin-1 in diabetes. Life Sci 2014; 118: 275–280.
Filios SR, Xu G, Chen J, Hong K, Jing G, Shalev A . MicroRNA-200 is induced by thioredoxin-interacting protein and regulates Zeb1 protein signaling and beta cell apoptosis. J Biol Chem 2014; 289: 36275–36283.
Srivastava A, Suy S, Collins SP, Kumar D . Circulating microRNA as biomarkers: an update in prostate cancer. Mol Cell Pharmacol 2011; 3: 115–124.
Lujambio A, Esteller M . How epigenetics can explain human metastasis: a new role for microRNAs. Cell Cycle 2009; 8: 377–382.
Valeri N, Vannini I, Fanini F, Calore F, Adair B, Fabbri M . Epigenetics, miRNAs, and human cancer: a new chapter in human gene regulation. Mamm Genome 2009; 20: 573–580.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Krishnan, R., Mani, P., Sivakumar, P. et al. Expression and methylation of circulating microRNA-510 in essential hypertension. Hypertens Res 40, 361–363 (2017). https://doi.org/10.1038/hr.2016.147
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hr.2016.147
Keywords
This article is cited by
-
miRNA profile and disease severity in patients with sickle cell anemia
Annals of Hematology (2022)
-
Impact of Nutritional Epigenetics in Essential Hypertension: Targeting microRNAs in the Gut-Liver Axis
Current Hypertension Reports (2021)
-
Decoding the functional role of long noncoding RNAs (lncRNAs) in hypertension progression
Hypertension Research (2020)
-
Epigenetic modification: a regulatory mechanism in essential hypertension
Hypertension Research (2019)
-
Methylation-dependent circulating microRNA 510 in preeclampsia patients
Hypertension Research (2019)